
Essence of Linear Algebra (9): Singular Value Decomposition — The Crown Jewel of Linear Algebra
SVD decomposes any matrix -- not just square or symmetric ones. From image compression to Netflix recommendations, from face recognition to gene analysis, SVD is the most powerful and most universal decomposition in linear algebra.
Why SVD Earns the Crown#
The spectral theorem of Chapter 8 gave us $A = Q\Lambda Q^T$ — a beautifully clean factorisation, but only for symmetric matrices. Most matrices that show up in practice are not symmetric, and many are not even square:
- a photograph stored as a $1920 \times 1080$ pixel matrix,
- a Netflix-style user–movie rating matrix (millions of rows, thousands of columns),
- a document–term matrix in NLP (documents by vocabulary),
- a gene-expression matrix in bioinformatics.
This is the most powerful, most universally applicable decomposition in all of linear algebra.
A photography analogy#
Picture a photo as a matrix of pixel intensities. SVD says three things at once:
- Any photo decomposes into a sum of “basic layers.”
- The layers are ranked by importance — the first captures gross structure, the second secondary detail, the third still finer detail.
- Keeping just the first few layers recovers most of the image.
Think of a band recording: lead vocal, guitar, bass, drums. Drop a background harmony and the song still works; drop the lead vocal and it falls apart. SVD makes that intuition precise: the singular values measure exactly how much each layer contributes.
What you will learn#
- The definition of SVD and its three-step geometric meaning (rotate, stretch, rotate).
- How to compute singular values and singular vectors via $A^{\!\top}\!A$ and $AA^{\!\top}$ .
- The four fundamental subspaces, read directly off $U$ and $V$ .
- Low-rank approximation and the Eckart–Young theorem — the optimality result behind compression.
- The pseudoinverse: a universal “best inverse” for matrices that have no honest inverse.
- PCA as SVD in disguise.
- Applications: image compression, recommender systems, latent semantic analysis, denoising, eigenfaces.
Prerequisites#
- Eigenvalues and eigenvectors (Chapter 6 )
- Orthogonal matrices and projections (Chapter 7 )
- Symmetric matrices and the spectral theorem (Chapter 8 )
The Definition#
The fundamental theorem#
$$ A = U\,\Sigma\,V^{\!\top} $$where
- $U \in \mathbb{R}^{m\times m}$ is orthogonal (its columns are the left singular vectors $u_1, \ldots, u_m$ );
- $V \in \mathbb{R}^{n\times n}$ is orthogonal (its columns are the right singular vectors $v_1, \ldots, v_n$ );
- $\Sigma \in \mathbb{R}^{m\times n}$ has the singular values $\sigma_1 \ge \sigma_2 \ge \cdots \ge 0$ on its main diagonal and zeros elsewhere.
Three facts make the singular values special:
- They are non-negative real numbers — always. (Eigenvalues can be negative or complex.)
- They are arranged in descending order by convention.
- SVD exists for every matrix. That is exactly the property eigendecomposition lacks, and it is what makes SVD universal.
Geometric meaning: a transformation in three steps#
The factorisation $A = U\Sigma V^{\!\top}$ has a clean visual story. Reading right to left, applying $A$ to a vector means:
- Rotate by $V^{\!\top}$ : align the input with the “natural input directions” of $A$ .
- Stretch by $\Sigma$ : scale the $i$ -th coordinate by $\sigma_i$ .
- Rotate by $U$ : place the result into its final orientation in the output space.
The dough analogy: you knead with a rolling pin. First you turn the dough to a convenient angle ($V^{\!\top}$ ); then the pin flattens and stretches it ($\Sigma$ ); finally you rotate the flattened dough where you need it ($U$ ).
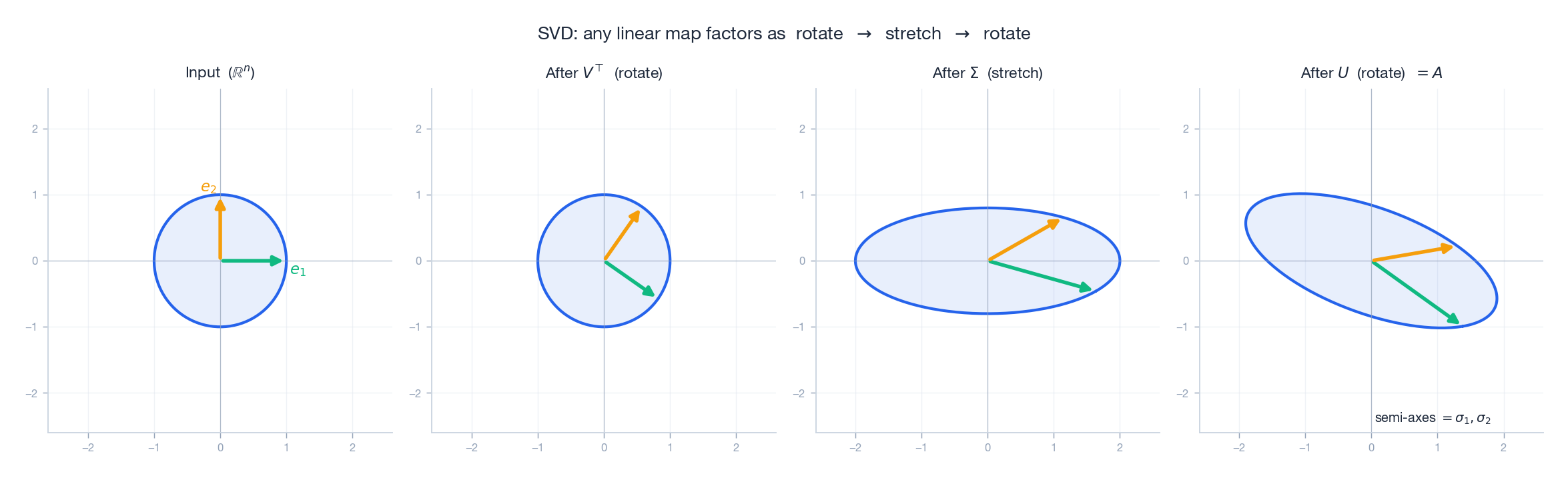
The figure tracks a unit circle through each stage. The orthogonal basis stays orthogonal under the two rotations; only the middle step (the diagonal $\Sigma$ ) actually changes shape, turning the circle into an ellipse with semi-axes $\sigma_1$ and $\sigma_2$ .
Unit circle to ellipse#
There is a complementary picture that compresses the same content into one input/output pair: the unit circle on the left, the image ellipse $A(\text{circle})$ on the right.
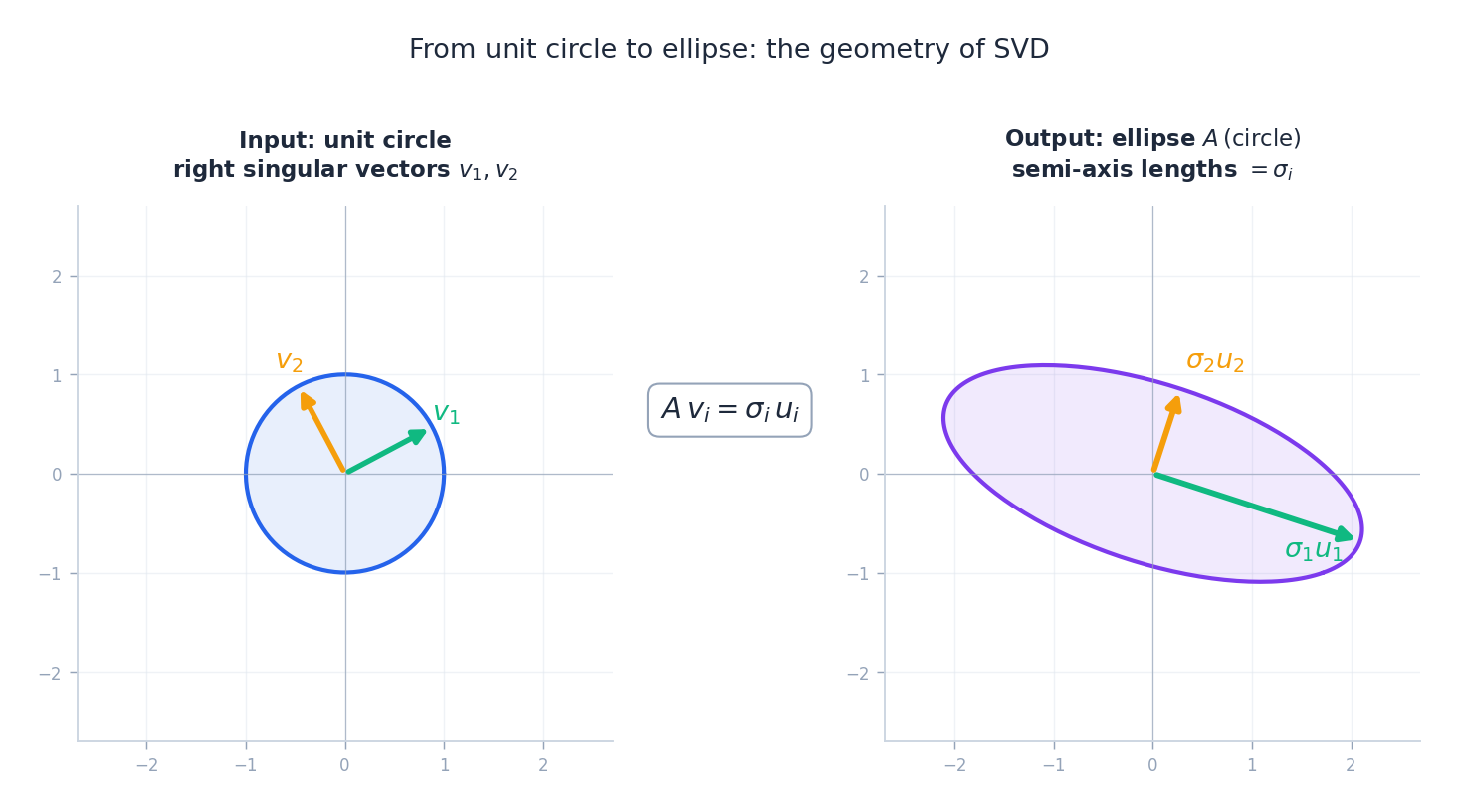
Reading the picture:
- The right singular vectors $v_1, v_2$ are the orthonormal directions on the input side that get mapped onto the principal axes of the output ellipse.
- The left singular vectors $u_1, u_2$ are the orthonormal directions of those axes.
- The singular values $\sigma_1, \sigma_2$ are the half-lengths of the axes.
This is the equation that makes everything else work.
Outer-product form#
$$ A = \sigma_1 u_1 v_1^{\!\top} + \sigma_2 u_2 v_2^{\!\top} + \cdots + \sigma_r u_r v_r^{\!\top}. $$Each $u_i v_i^{\!\top}$ is a rank-1 matrix, and the singular values are the weights. This perspective is the key to low-rank approximation: keep the largest weights, drop the rest.
Computing SVD#
The bridge to $A^{\!\top}\!A$ and $AA^{\!\top}$ #
$$ A^{\!\top}\!A = V\,\Sigma^{\!\top}\!\Sigma\,V^{\!\top}, \qquad AA^{\!\top} = U\,\Sigma\Sigma^{\!\top}\,U^{\!\top}. $$Both $A^{\!\top}\!A$ and $AA^{\!\top}$ are symmetric and positive semidefinite, so the spectral theorem applies. Reading these as spectral decompositions:
- columns of $V$ = orthonormal eigenvectors of $A^{\!\top}\!A$ (the right singular vectors),
- columns of $U$ = orthonormal eigenvectors of $AA^{\!\top}$ (the left singular vectors),
- $\sigma_i = \sqrt{\lambda_i}$ where $\lambda_i$ are the (nonnegative) eigenvalues — shared by both products.
Why $A^{\!\top}\!A$ ? Think of it as $A$ “acting twice”: go forward by $A$ , come back by $A^{\!\top}$ . The round trip amplifies a direction by exactly $\sigma^2$ , which is why eigenvalues of $A^{\!\top}\!A$ are squared singular values.
Step-by-step computation#
Given $m \times n$ matrix $A$ with $m \ge n$ :
- Form $A^{\!\top}\!A$ . Compute its (sorted) eigenvalues $\lambda_1 \ge \cdots \ge \lambda_n \ge 0$ and orthonormal eigenvectors $v_1, \ldots, v_n$ . These are the columns of $V$ .
- Set $\sigma_i = \sqrt{\lambda_i}$ .
- For each $\sigma_i > 0$ , define $u_i = A v_i / \sigma_i$ . These are the first $r$ columns of $U$ .
- If $r < m$ , extend $\{u_1, \ldots, u_r\}$ to an orthonormal basis of $\mathbb{R}^m$ (e.g. via Gram–Schmidt) to fill out $U$ .
In numerical practice nobody computes SVD this way: forming $A^{\!\top}\!A$ squares the condition number. Production code uses bidiagonalisation followed by the QR algorithm or divide-and-conquer methods. The above derivation is conceptual.
Worked example#
$$A^{\!\top}\!A = \begin{pmatrix} 1 & 1 \\ 1 & 2 \end{pmatrix}, \qquad \det(A^{\!\top}\!A - \lambda I) = \lambda^2 - 3\lambda + 1.$$ $$ \sigma_1 = \sqrt{\tfrac{3+\sqrt 5}{2}} \approx 1.618, \qquad \sigma_2 = \sqrt{\tfrac{3-\sqrt 5}{2}} \approx 0.618. $$Find the eigenvectors of $A^{\!\top}\!A$ to assemble $V$ , then $u_i = A v_i / \sigma_i$ gives $U$ . (The product $\sigma_1 \sigma_2 = 1 = |\det A|$ is a useful sanity check.)
Eigenvalues vs singular values: side by side#
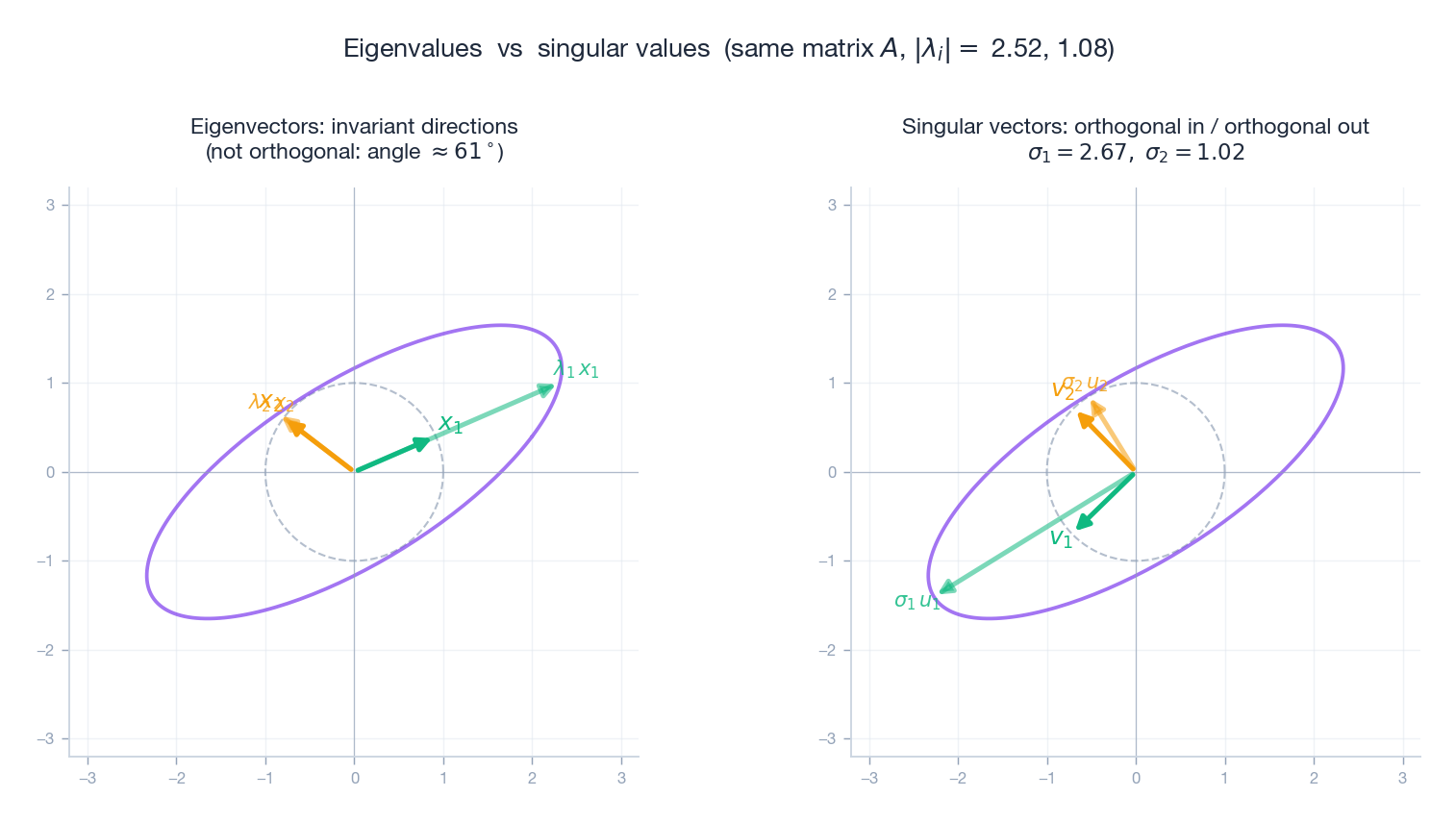
For symmetric $A$ , eigenvalues and singular values agree (up to signs). For general $A$ they do not, and the contrast is illuminating.
| Eigenvectors | Singular vectors | |
|---|---|---|
| Defined by | $A x = \lambda x$ | $A v = \sigma u$ with $u \perp$ each other |
| Orthogonality | not in general | always (both $V$ and $U$ are orthonormal) |
| Values | $\lambda_i \in \mathbb{C}$ | $\sigma_i \in \mathbb{R}_{\ge 0}$ |
| Geometric story | invariant directions, scaled by $\lambda$ | input axes mapped to output axes, scaled by $\sigma$ |
| Exists for | square matrices, often diagonalisable | every matrix |
The figure makes the difference visceral: the same non-symmetric $A$ has eigenvectors at an oblique angle, but the SVD picks a perpendicular pair on each side.
SVD and the Four Fundamental Subspaces#
SVD gives the cleanest possible picture of a matrix’s structure. For $A \in \mathbb{R}^{m \times n}$ with rank $r$ :
| Subspace | Dimension | Orthonormal basis from SVD | Lives in |
|---|---|---|---|
| Row space $\mathcal{C}(A^{\!\top})$ | $r$ | $v_1, \ldots, v_r$ | $\mathbb{R}^n$ |
| Null space $\mathcal{N}(A)$ | $n - r$ | $v_{r+1}, \ldots, v_n$ | $\mathbb{R}^n$ |
| Column space $\mathcal{C}(A)$ | $r$ | $u_1, \ldots, u_r$ | $\mathbb{R}^m$ |
| Left null space $\mathcal{N}(A^{\!\top})$ | $m - r$ | $u_{r+1}, \ldots, u_m$ | $\mathbb{R}^m$ |
The orthonormal basis $\{v_1, \ldots, v_r\}$ of the row space is mapped onto the orthonormal basis $\{u_1, \ldots, u_r\}$ of the column space, with each direction stretched by its $\sigma_i$ . Everything in the null space is sent to zero. That is the whole action of $A$ in one sentence.
Low-Rank Approximation: the Theorem Behind Compression#

The Eckart–Young theorem#
$$ A_k = \sigma_1 u_1 v_1^{\!\top} + \cdots + \sigma_k u_k v_k^{\!\top}. $$ $$ \|A - A_k\|_F \;=\; \min_{\operatorname{rank}(B) \le k} \|A - B\|_F \;=\; \sqrt{\sigma_{k+1}^{\,2} + \cdots + \sigma_r^{\,2}}. $$The same statement holds in the operator (2-)norm with $\|A - A_k\|_2 = \sigma_{k+1}$ .
So $A_k$ isn’t just a low-rank approximation — it is provably optimal. No clever rank-$k$ matrix can do better.
MP3 analogy. MP3 compression discards high-frequency components the human ear barely registers. SVD truncation does the same thing for matrices: discard the components carrying the least “energy,” keep the loud ones.
Layer by layer#
It helps to see the layers stack up. Each $\sigma_i u_i v_i^{\!\top}$ is a single rank-1 image; partial sums approach the original.
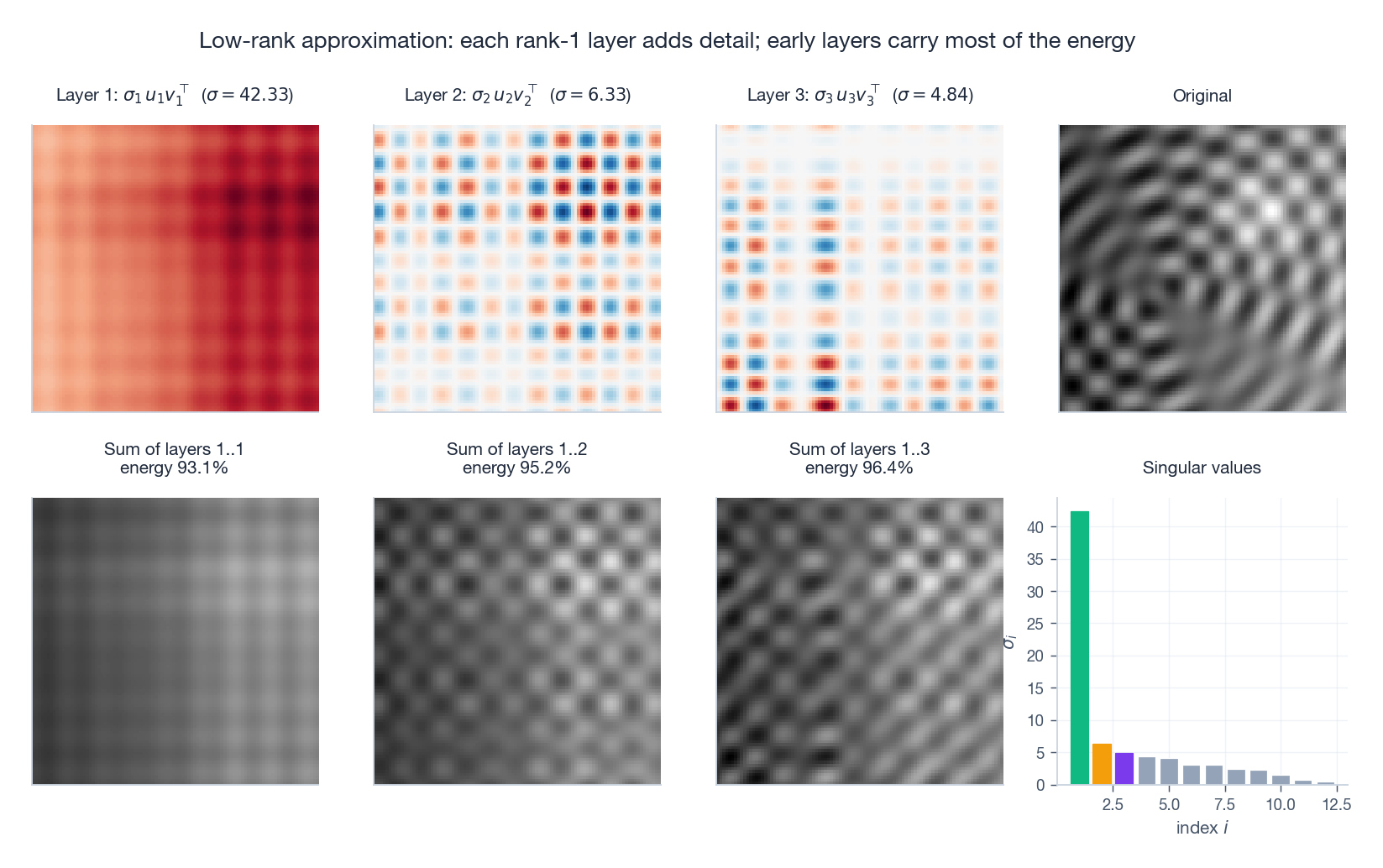
Top row: the first three rank-1 layers themselves (positive in red, negative in blue) plus the original. Bottom row: the cumulative sums after 1, 2, 3 layers, with the singular value bar chart on the right. Even three components capture a recognisable amount of the picture.
Energy#
$$ \|A\|_F^2 = \sigma_1^2 + \sigma_2^2 + \cdots + \sigma_r^2. $$ $$ \text{energy retained} = \frac{\sigma_1^2 + \cdots + \sigma_k^2}{\sigma_1^2 + \cdots + \sigma_r^2}. $$For most natural data the spectrum decays quickly: a $1000 \times 1000$ photograph might capture 95% of its energy in the first 50 singular values.
Image compression in numbers#
$$ \text{numbers stored} = k\,(m + n + 1). $$For a $500 \times 500$ image with $k = 50$ : the original needs $250{,}000$ numbers, the rank-50 approximation needs $50{,}050$ — a 5x reduction with usually imperceptible loss.
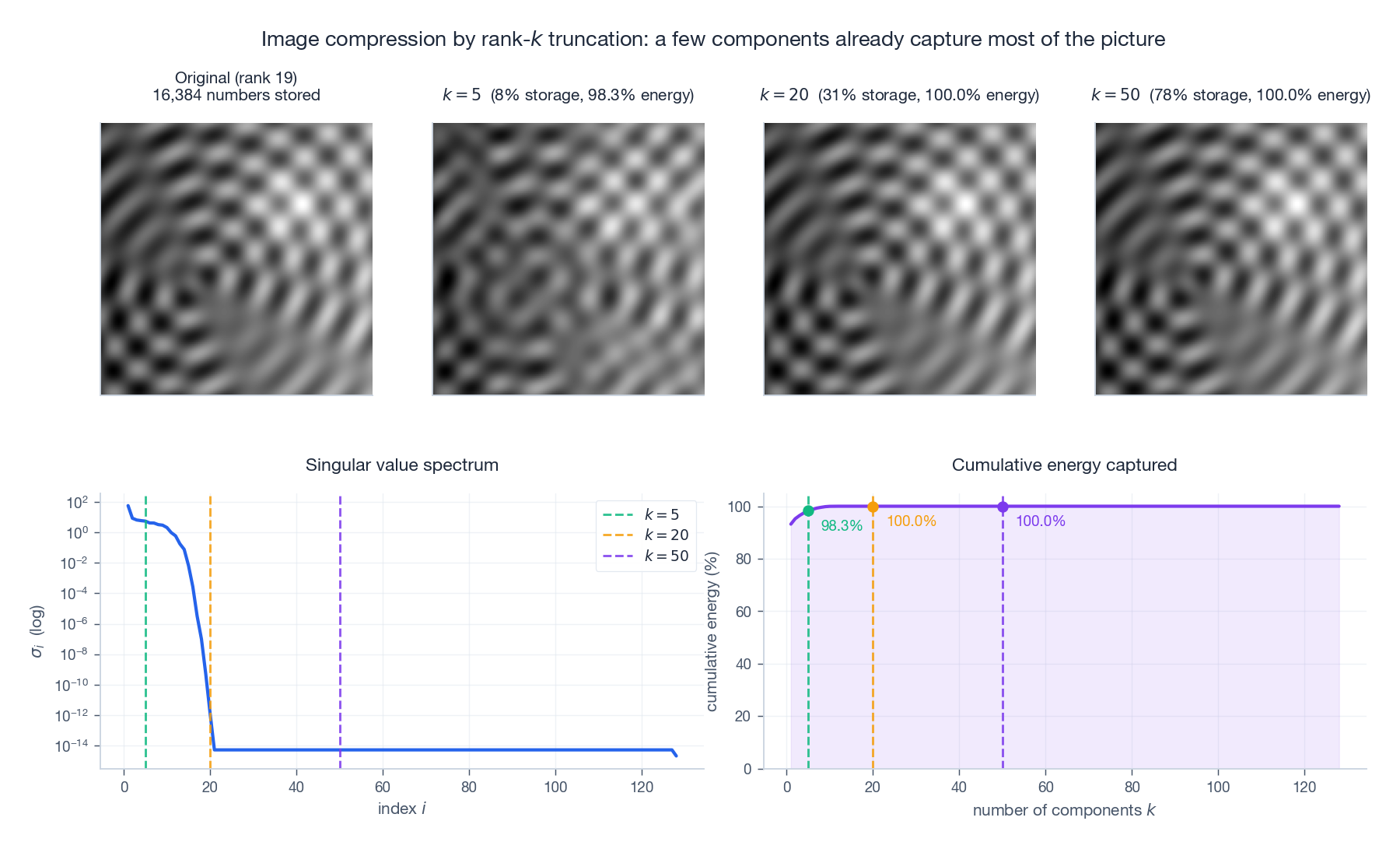
The bottom-left log-scale plot is the most important diagnostic in all of applied SVD: it tells you how aggressively you can truncate before quality collapses.
| |
The Pseudoinverse#
When the inverse does not exist#
For $A x = b$ , we usually want $x = A^{-1} b$ . But $A^{-1}$ exists only when $A$ is square and full-rank. The Moore–Penrose pseudoinverse $A^{+}$ supplies a universal “best alternative.”
Definition via SVD#
$$ A^{+} = V \Sigma^{+} U^{\!\top}, \qquad \Sigma^{+}_{ii} = \begin{cases} 1/\sigma_i & \sigma_i > 0,\\ 0 & \sigma_i = 0,\end{cases} $$and $\Sigma^{+}$ is transposed to have shape $n \times m$ . When $A$ is invertible, $A^{+} = A^{-1}$ .
What $A^{+}$ does#
For any $b$ , the vector $\hat x = A^{+} b$ is
- a least-squares solution: it minimises $\|A x - b\|_2$ ;
- among all least-squares solutions, the one with minimum norm $\|x\|_2$ .
The two cases:
- Overdetermined ($m > n$ ): typically no exact solution. $A^{+} b$ delivers the least-squares fit.
- Underdetermined ($m < n$ ): infinitely many solutions. $A^{+} b$ picks the shortest.
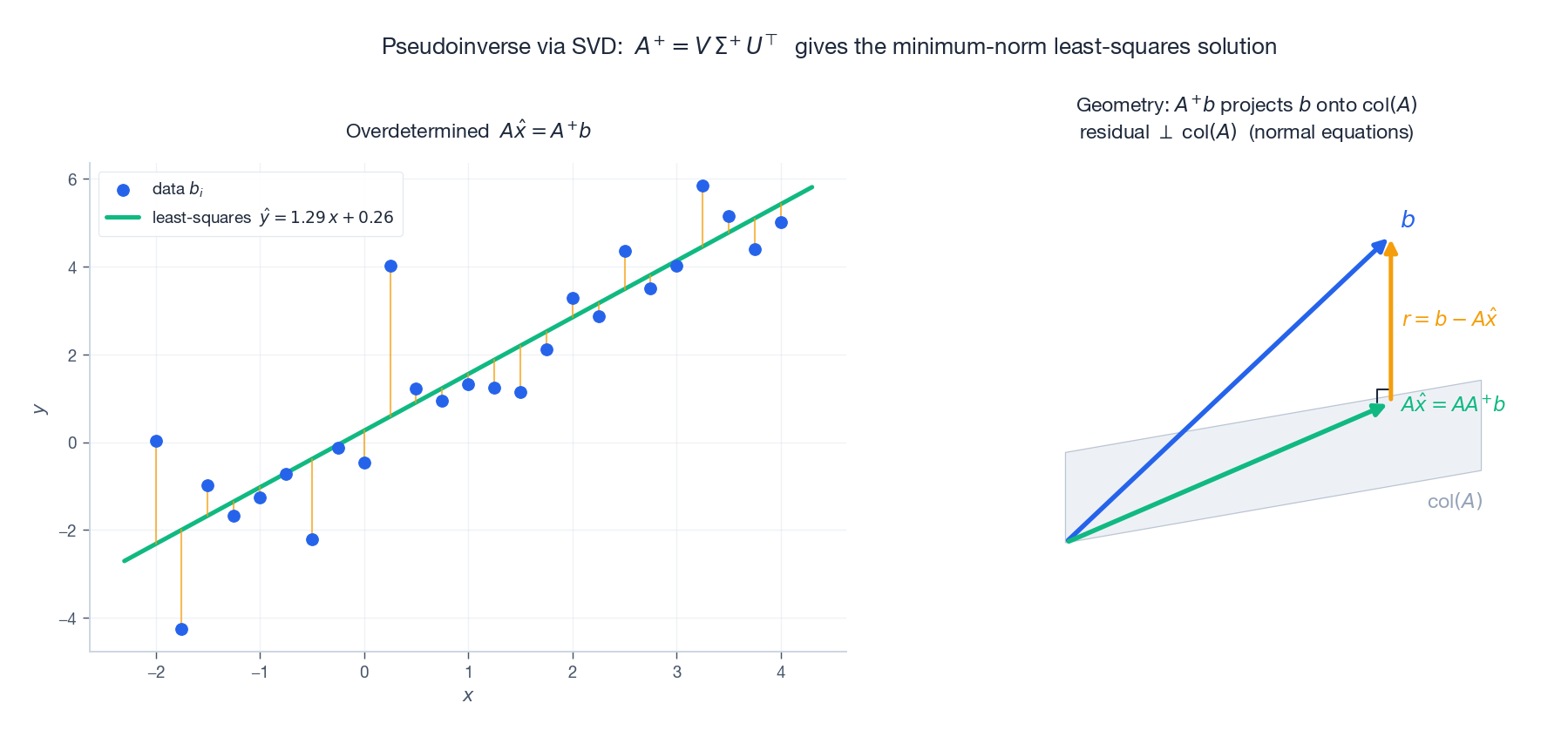
The left panel shows the canonical least-squares line fit; the right panel shows the geometry that makes it least-squares: the projection of $b$ onto the column space of $A$ , with the residual orthogonal to it. That orthogonality is the normal equation, and SVD gives it for free.
| |
PCA via SVD#
The connection#
Principal Component Analysis is SVD wearing a statistics hat.
$$ X_c = U \Sigma V^{\!\top}. $$Then:
- Principal directions = columns of $V$ (right singular vectors). These are the orthogonal axes of maximum variance.
- Principal scores = $X_c V = U \Sigma$ . These are the data re-expressed in the new basis.
- Variance along PC$_i$ = $\sigma_i^2 / (n - 1)$ .
- Dimensionality reduction to $k$ components: $X_k = U_k \Sigma_k$ .
Why PCA works#
The first principal direction maximises $\operatorname{Var}(X_c w)$ over unit vectors $w$ . A short calculation shows this is $w^{\!\top}\!(X_c^{\!\top} X_c)\,w / (n-1)$ , which is maximised at the top eigenvector of $X_c^{\!\top} X_c$ — exactly the top right singular vector $v_1$ .
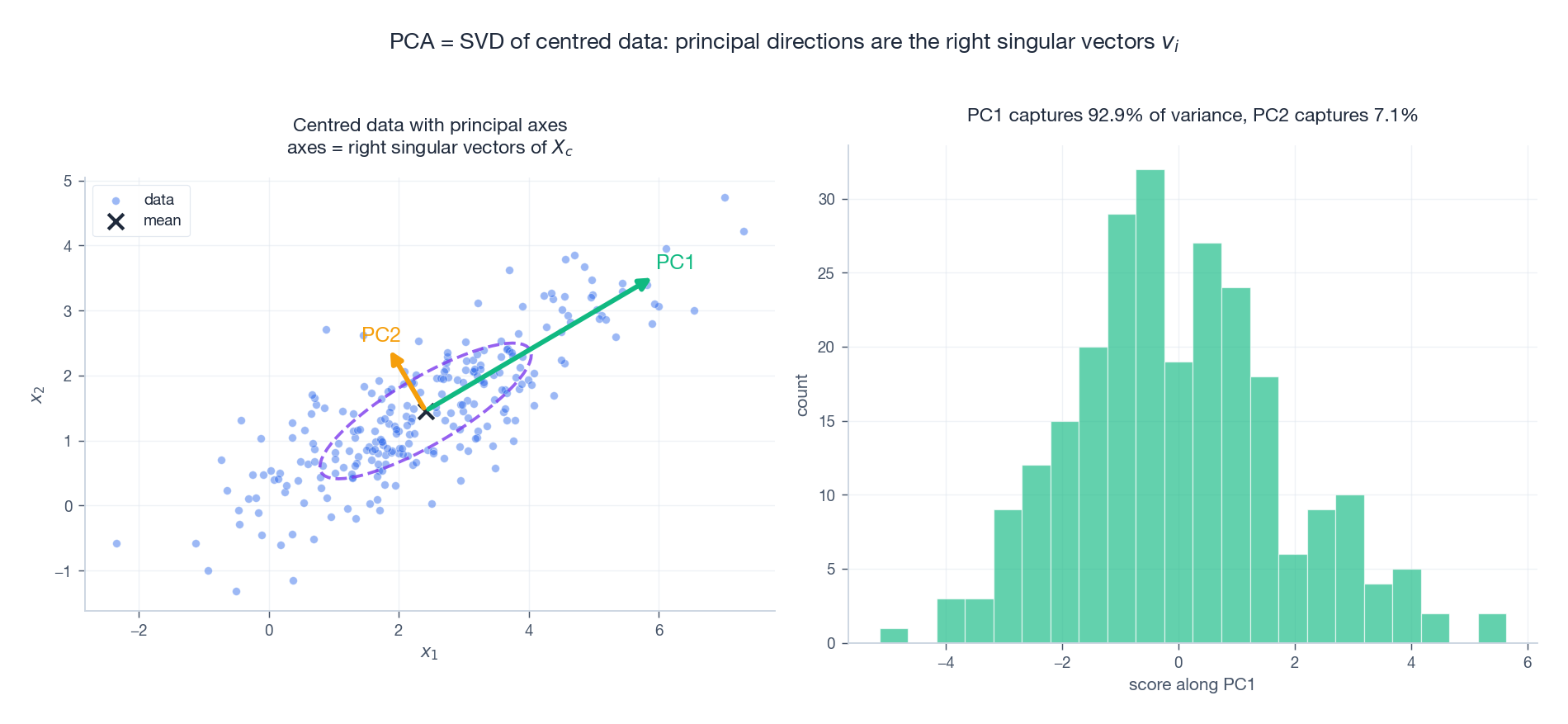
The dashed ellipse is the 1-$\sigma$ Gaussian fit. The two arrows are the principal axes drawn proportional to the standard deviations $\sigma_i / \sqrt{n-1}$ . Projecting onto PC1 collapses 2D to 1D while preserving the lion’s share of variance.
| |
Recommender Systems#
The setup#
Netflix, Amazon, and Spotify all face the same question: how do we predict ratings for items a user has not yet seen? The user–item rating matrix $R \in \mathbb{R}^{m\times n}$ is enormous and mostly missing.
Matrix factorisation#
$$ R \approx U_k \Sigma_k V_k^{\!\top}. $$- A row of $U_k \Sigma_k$ is one user’s taste vector.
- A row of $V_k$ is one item’s characteristic vector.
- The predicted rating is just their dot product.
This idea — learn $U_k$ and $V_k$ to fit the observed entries, then read off predictions for the rest — powered the winning entries of the Netflix Prize (2006–2009).
SVD vs. Eigendecomposition#
| Property | Eigendecomposition | SVD |
|---|---|---|
| Applicable to | square (often symmetric) | any matrix |
| Form | $A = P\Lambda P^{-1}$ | $A = U\Sigma V^{\!\top}$ |
| Values | eigenvalues; can be negative or complex | singular values; non-negative real |
| Vectors | eigenvectors; not always orthogonal | singular vectors; always orthogonal |
| Geometric story | invariant directions + scaling | rotate + stretch + rotate |
| Always exists? | no | yes |
Why SVD earns “crown jewel” status#
- Universal — works on any matrix, square or not, full rank or not.
- Stable — numerically robust; the gold standard for rank, conditioning, and least squares.
- Optimal — gives provably best low-rank approximations (Eckart–Young).
- Insightful — exposes rank, the four subspaces, and the operator norm in one shot.
- Practical — image compression, NLP, recommender systems, denoising, control, statistics.
Other Applications#
Latent Semantic Analysis (LSA)#
Build a document–term matrix (rows = documents, columns = vocabulary, entries = TF-IDF scores). Apply SVD and keep the top $k$ components. The right singular vectors play the role of “latent topics,” and document similarity becomes a cosine similarity in this much smaller space.
Signal denoising#
Low-rank signal + full-rank noise: SVD the observation, keep the large singular values, discard the small ones, reconstruct. This is the workhorse trick behind everything from astronomical image cleaning to seismic-data processing.
Eigenfaces#
Run PCA on a database of aligned face images. The principal components (“eigenfaces”) form a basis of typical facial appearance. Any new face is a linear combination of eigenfaces, and recognition reduces to comparing coefficient vectors.
Python Implementation#
Manual SVD via eigendecomposition#
| |
Image compression demo#
| |
Exercises#
Basics#
- Explain why singular values are always non-negative, while eigenvalues can be negative or complex.
- Compute the SVD of $A = \begin{pmatrix} 1 & 0 \\ 0 & 0 \end{pmatrix}$ by hand.
- If $A$ is $3 \times 5$ , what are the shapes of $U$ , $\Sigma$ , and $V^{\!\top}$ in the full SVD? In the economy SVD?
Advanced#
- Prove that $\operatorname{rank}(A)$ equals the number of nonzero singular values.
- Prove $\|A\|_F^2 = \sigma_1^2 + \cdots + \sigma_r^2$ .
- If $Q$ is orthogonal, what are its singular values? Why?
- Show that $U_r U_r^{\!\top}$ is the projection matrix onto the column space, and $V_r V_r^{\!\top}$ is the projection onto the row space.
- Prove the operator-norm version of Eckart–Young: $\|A - A_k\|_2 = \sigma_{k+1}$ .
Programming#
- Compression curve. Load a grayscale image, compute its SVD, and plot rank-$k$ reconstructions for $k \in \{5, 20, 50, 100\}$ together with the singular-value decay on a log scale.
- PCA on Iris. Apply PCA to the Iris dataset. Plot the first two components, colour-coded by species, and report the explained-variance ratio.
- Toy recommender. Build a $5 \times 10$ rating matrix with some entries missing, fill missing values with the row mean, run a rank-3 SVD, and inspect the predicted ratings.
Summary#
| Concept | Key formula | Intuition |
|---|---|---|
| SVD | $A = U\Sigma V^{\!\top}$ | rotate + stretch + rotate |
| Singular values | $\sigma_i \ge 0$ | stretching factors of the ellipse |
| Outer-product form | $A = \sum_i \sigma_i u_i v_i^{\!\top}$ | weighted sum of rank-1 layers |
| Low-rank approximation | $A_k = U_k \Sigma_k V_k^{\!\top}$ | optimal rank-$k$ matrix (Eckart–Young) |
| Pseudoinverse | $A^{+} = V \Sigma^{+} U^{\!\top}$ | minimum-norm least-squares solution |
| PCA | SVD of centred $X$ | maximum-variance directions = right singular vectors |
References#
- Strang, G. (2019). Introduction to Linear Algebra, Chapter 7 .
- Trefethen, L. N. & Bau, D. (1997). Numerical Linear Algebra. SIAM.
- Golub, G. H. & Van Loan, C. F. (2013). Matrix Computations, 4th ed. Johns Hopkins.
- Eckart, C. & Young, G. (1936). “The approximation of one matrix by another of lower rank.” Psychometrika, 1(3).
- Hastie, T., Tibshirani, R. & Friedman, J. (2009). The Elements of Statistical Learning. Springer.
- Koren, Y., Bell, R. & Volinsky, C. (2009). “Matrix Factorization Techniques for Recommender Systems.” Computer, 42(8).
- 3Blue1Brown. Essence of Linear Algebra series.
Linear Algebra 18 parts
- 01 Essence of Linear Algebra (1): The Essence of Vectors — More Than Just Arrows
- 02 Essence of Linear Algebra (2): Linear Combinations and Vector Spaces
- 03 Essence of Linear Algebra (3): Matrices as Linear Transformations
- 04 Essence of Linear Algebra (4): The Secrets of Determinants
- 05 Essence of Linear Algebra (5): Linear Systems and Column Space
- 06 Essence of Linear Algebra (6): Eigenvalues and Eigenvectors
- 07 Essence of Linear Algebra (7): Orthogonality and Projections — When Vectors Mind Their Own Business
- 08 Essence of Linear Algebra (8): Symmetric Matrices and Quadratic Forms — The Best Matrices in Town
- 09 Essence of Linear Algebra (9): Singular Value Decomposition — The Crown Jewel of Linear Algebra you are here
- 10 Essence of Linear Algebra (10): Matrix Norms and Condition Numbers — Is Your Linear System Healthy?
- 11 Essence of Linear Algebra (11): Matrix Calculus and Optimization — The Engine Behind Machine Learning
- 12 Essence of Linear Algebra (12): Sparse Matrices and Compressed Sensing — Less Is More
- 13 Essence of Linear Algebra (13): Tensors and Multilinear Algebra
- 14 Essence of Linear Algebra (14): Random Matrix Theory
- 15 Essence of Linear Algebra (15): Linear Algebra in Machine Learning
- 16 Essence of Linear Algebra (16): Linear Algebra in Deep Learning
- 17 Essence of Linear Algebra (17): Linear Algebra in Computer Vision
- 18 Essence of Linear Algebra (18): Frontiers and Summary