
ML Math Derivations (2): Linear Algebra and Matrix Theory
The language of machine learning is linear algebra. This article derives vector spaces, eigendecomposition, SVD, and matrix calculus from first principles -- every tool you need for ML optimization.

Why this chapter, and what’s different#
If you have already worked through a standard linear-algebra course you have seen most of these objects. This chapter is not that course. It is the ML practitioner’s slice of linear algebra: the half-dozen ideas that actually appear when you implement gradient descent, run PCA, train a neural net, or read a paper.
Concretely the goals are:
- Build a geometric intuition for what matrices do (rotate, stretch, project, kill).
- Learn the four decompositions that show up everywhere — spectral, SVD, QR, Cholesky — and which one to reach for.
- Master enough matrix calculus to derive any neural-net gradient on the back of an envelope.
We skim the algebra of row reduction, determinants by cofactor, and abstract vector-space proofs. If you need those, the references at the bottom give the standard treatments. Here, every concept comes back to a picture or a line of NumPy.
Prerequisites: matrix multiplication, transpose, the idea of a determinant, partial derivatives.
Vector spaces, subspaces, rank — the geometric core#
Forget the eight axioms for a moment. The mental model that pays off in ML is:
A vector space is a flat, infinite “sheet” through the origin. A subspace is a smaller flat sheet through the origin sitting inside it.
A line through the origin is a 1D subspace of $\mathbb{R}^2$ . A plane through the origin is a 2D subspace of $\mathbb{R}^3$ . The crucial word is through the origin — shift the sheet and you lose closure under addition and scaling.
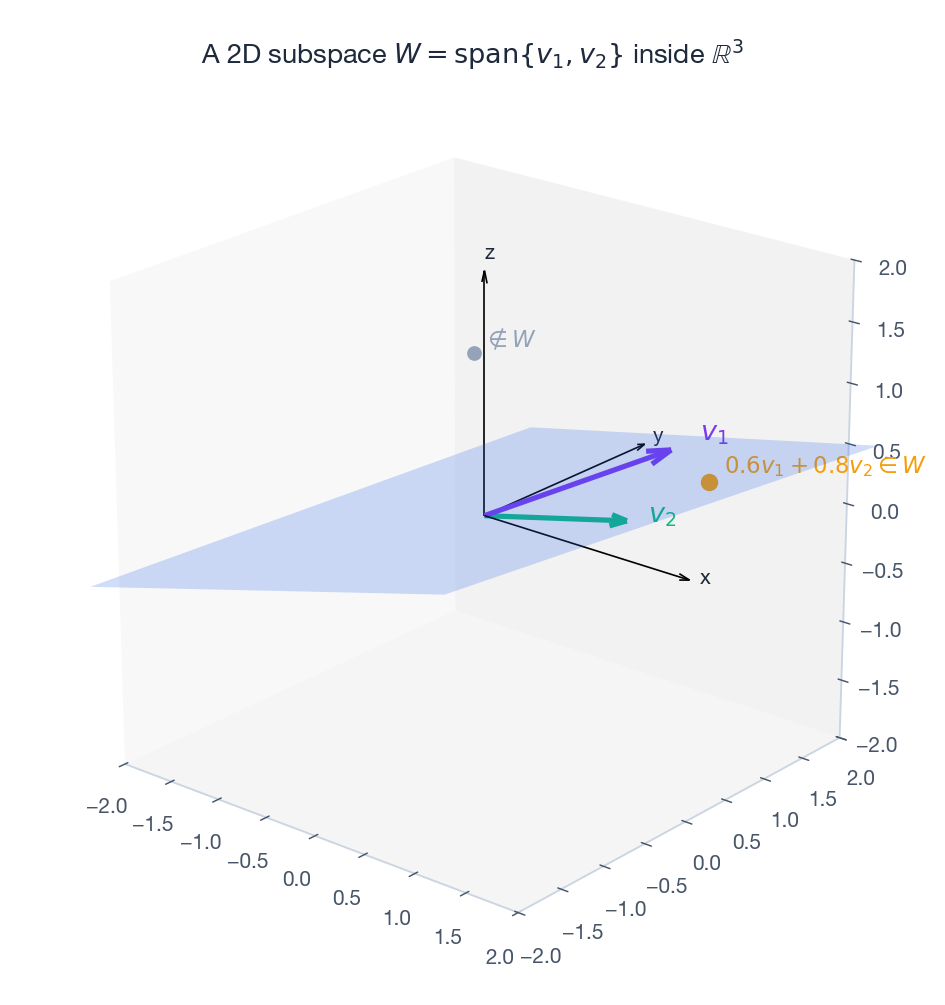
Span, independence, basis — in one breath#
Pick a few vectors $v_1, \ldots, v_k$ . The set of all linear combinations $\sum \alpha_i v_i$ is their span — it is always a subspace.
$$\alpha_1 v_1 + \cdots + \alpha_k v_k = 0 \;\implies\; \alpha_1 = \cdots = \alpha_k = 0. \tag{1}$$A basis is an independent spanning set. Its size is the dimension of the subspace.
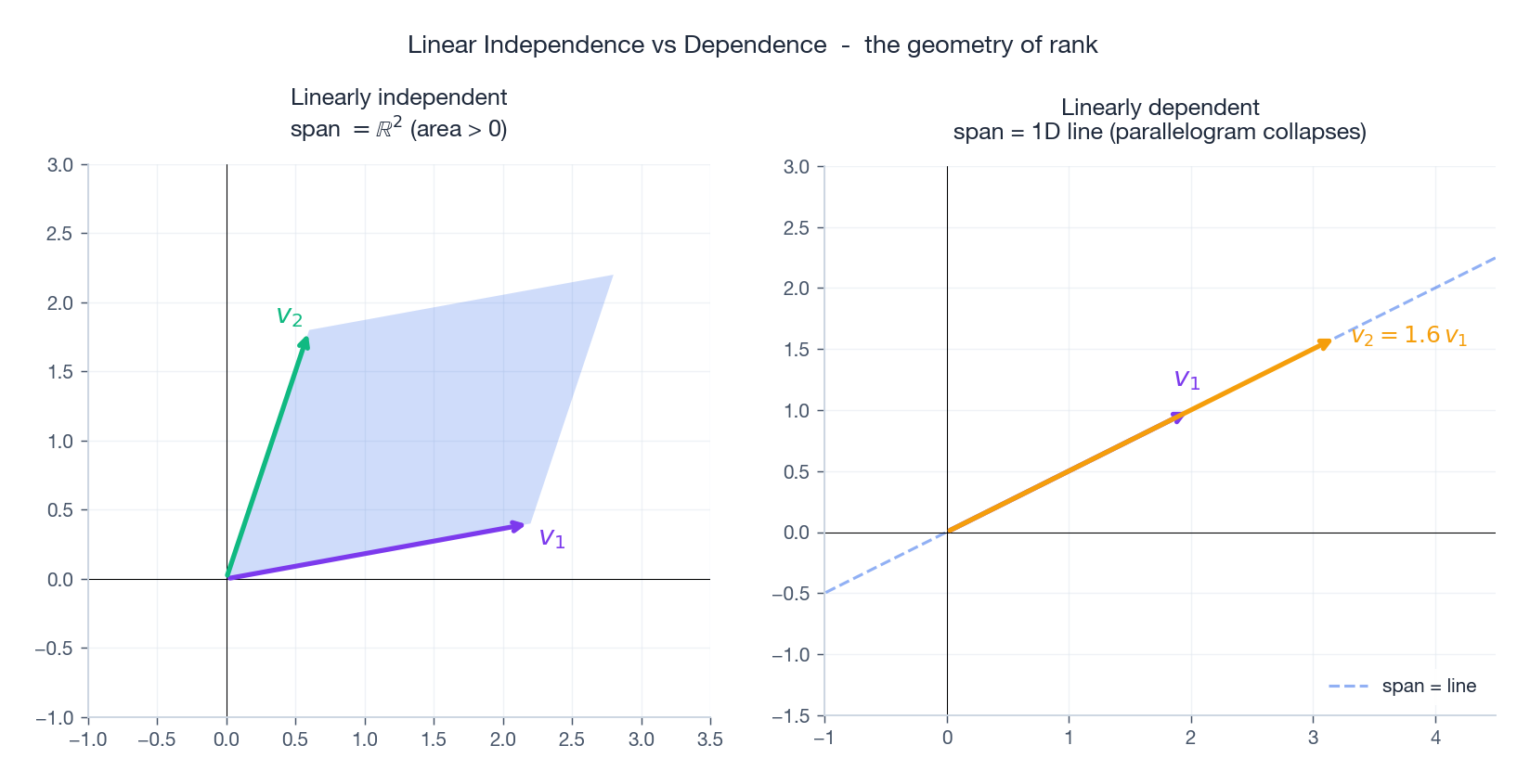
The picture is the whole story: independent vectors enclose a non-degenerate parallelogram (positive area, so they actually span a 2D piece). Dependent vectors lie on a line, and the parallelogram collapses.
Matrices ARE linear maps#
Every matrix $A \in \mathbb{R}^{m \times n}$ is the same object as a linear map $T : \mathbb{R}^n \to \mathbb{R}^m$ , and conversely. The dictionary is one rule:
The $j$ -th column of $A$ is $T(e_j)$ — where the $j$ -th standard basis vector lands.
So when you see a weight matrix $W \in \mathbb{R}^{h \times d}$ in a neural net layer, you are looking at a function “take a $d$ -dimensional input, output an $h$ -dimensional vector”; the columns of $W$ tell you what each input feature contributes.
Rank and the four fundamental subspaces#
For an $m \times n$ matrix $A$ of rank $r$ :
| Subspace | Definition | Lives in | Dimension | “Meaning” |
|---|---|---|---|---|
| Column space $\text{Col}(A)$ | $\{A x : x \in \mathbb{R}^n\}$ | $\mathbb{R}^m$ | $r$ | reachable outputs |
| Null space $\text{Null}(A)$ | $\{x : A x = 0\}$ | $\mathbb{R}^n$ | $n-r$ | inputs that get killed |
| Row space $\text{Row}(A)$ | $\text{Col}(A^\top)$ | $\mathbb{R}^n$ | $r$ | inputs $A$ “sees” |
| Left null space | $\{y : A^\top y = 0\}$ | $\mathbb{R}^m$ | $m-r$ | outputs unreachable |
Orthogonality (the fundamental theorem). $\text{Null}(A) \perp \text{Row}(A)$ .
Proof. If $Ax = 0$ and $w = A^\top z$ is in the row space, then $\langle x, w \rangle = z^\top (A x) = 0$ . $\square$
So $\mathbb{R}^n$ splits into two orthogonal pieces: the row space (where $A$ does interesting work) and the null space (where $A$ throws information away). Whenever a model has more parameters than equations — which is always in modern ML — the null space is non-trivial and many parameter vectors give the same predictions.
Norms and inner products — the geometry layer#
An inner product $\langle \cdot, \cdot \rangle$ gives you angles; a norm $\|\cdot\|$ gives you lengths. On $\mathbb{R}^n$ the standard one is $\langle x, y \rangle = x^\top y$ , with induced norm $\|x\|_2 = \sqrt{x^\top x}$ .
The $\ell_p$ family is the workhorse for regularisation:
| Norm | Formula | Unit ball | ML use |
|---|---|---|---|
| $\ell_1$ | $\sum_i \lvert x_i\rvert$ | diamond | Lasso, sparsity |
| $\ell_2$ | $\sqrt{\sum_i x_i^2}$ | circle / sphere | Ridge, weight decay |
| $\ell_\infty$ | $\max_i \lvert x_i\rvert$ | square / cube | adversarial bounds |
Cauchy-Schwarz and the cosine#
Theorem (Cauchy-Schwarz). $|\langle u, v \rangle| \le \|u\| \cdot \|v\|$ , with equality iff $u, v$ are parallel.
Proof. The quadratic $\|u + tv\|^2 = \|u\|^2 + 2t\langle u, v\rangle + t^2 \|v\|^2$ is non-negative for every $t$ , so its discriminant must be $\le 0$ . $\square$
Divide both sides by $\|u\|\|v\|$ and you have the cosine of the angle $\cos\theta = \langle u, v\rangle / (\|u\|\|v\|)$ — and the proof that it lives in $[-1, 1]$ . This is the formula behind cosine similarity in retrieval, attention, contrastive learning — everywhere.
Matrix norms you actually use#
| Norm | Formula | What it measures |
|---|---|---|
| Frobenius $\mid A\mid_F$ | $\sqrt{\sum_{ij} a_{ij}^2}$ | the “Euclidean length” of $A$ as a vector |
| Spectral $\mid A\mid_2$ | $\sigma_1(A)$ | maximum stretch over all unit inputs |
| Nuclear $\mid A\mid_*$ | $\sum_i \sigma_i(A)$ | convex surrogate for $\text{rank}(A)$ |
The spectral norm is the right thing for Lipschitz analysis (e.g. spectral normalisation in GANs). The nuclear norm is what you minimise for low-rank matrix completion (Netflix-style problems).
Eigendecomposition — “the directions that survive”#
Definition and intuition#
$$A v = \lambda v. \tag{2}$$Geometrically: $A$ usually rotates inputs, but along an eigenvector all $A$ does is rescale. The eigenvector’s direction survives the transformation.
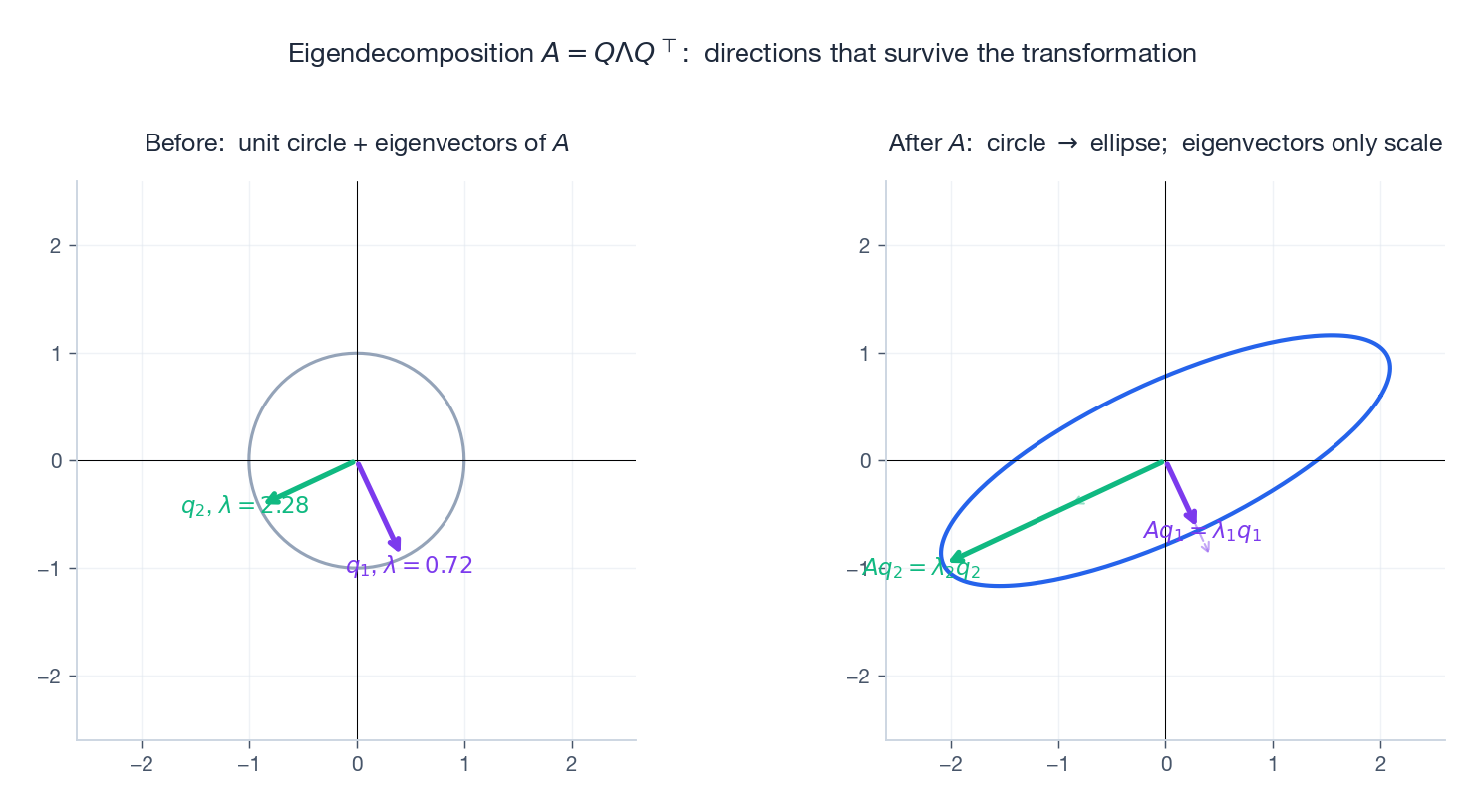
Eigenvalues solve the characteristic polynomial $\det(A - \lambda I) = 0$ . Two free identities:
- $\sum_i \lambda_i = \operatorname{tr}(A)$
- $\prod_i \lambda_i = \det(A)$
The spectral theorem (the only one you’ll use daily)#
Theorem. If $A$ is real and symmetric ($A = A^\top$ ), then:
- all eigenvalues are real,
- eigenvectors for distinct eigenvalues are orthogonal,
- $A$ is orthogonally diagonalisable: $A = Q \Lambda Q^\top$ with $Q^\top Q = I$ and $\Lambda$ diagonal.
Sketch (eigenvalues real). Take $A v = \lambda v$ with $v \neq 0$ . Compute $\bar v^\top A v$ two ways and use $A = A^\top$ : you get $\lambda = \bar\lambda$ . $\square$
$$A = \sum_{i=1}^n \lambda_i\, q_i q_i^\top. \tag{3}$$Read this carefully — $A$ is literally a weighted sum of projections onto orthogonal directions. This is the ML lens on every symmetric matrix: covariance, Gram, Hessian, graph Laplacian.
Positive (semi-)definite matrices#
A symmetric $A$ is positive definite ($A \succ 0$ ) when $x^\top A x > 0$ for all $x \neq 0$ . Equivalently, all eigenvalues are positive. Positive semi-definite ($A \succeq 0$ ) replaces $> 0$ with $\ge 0$ .
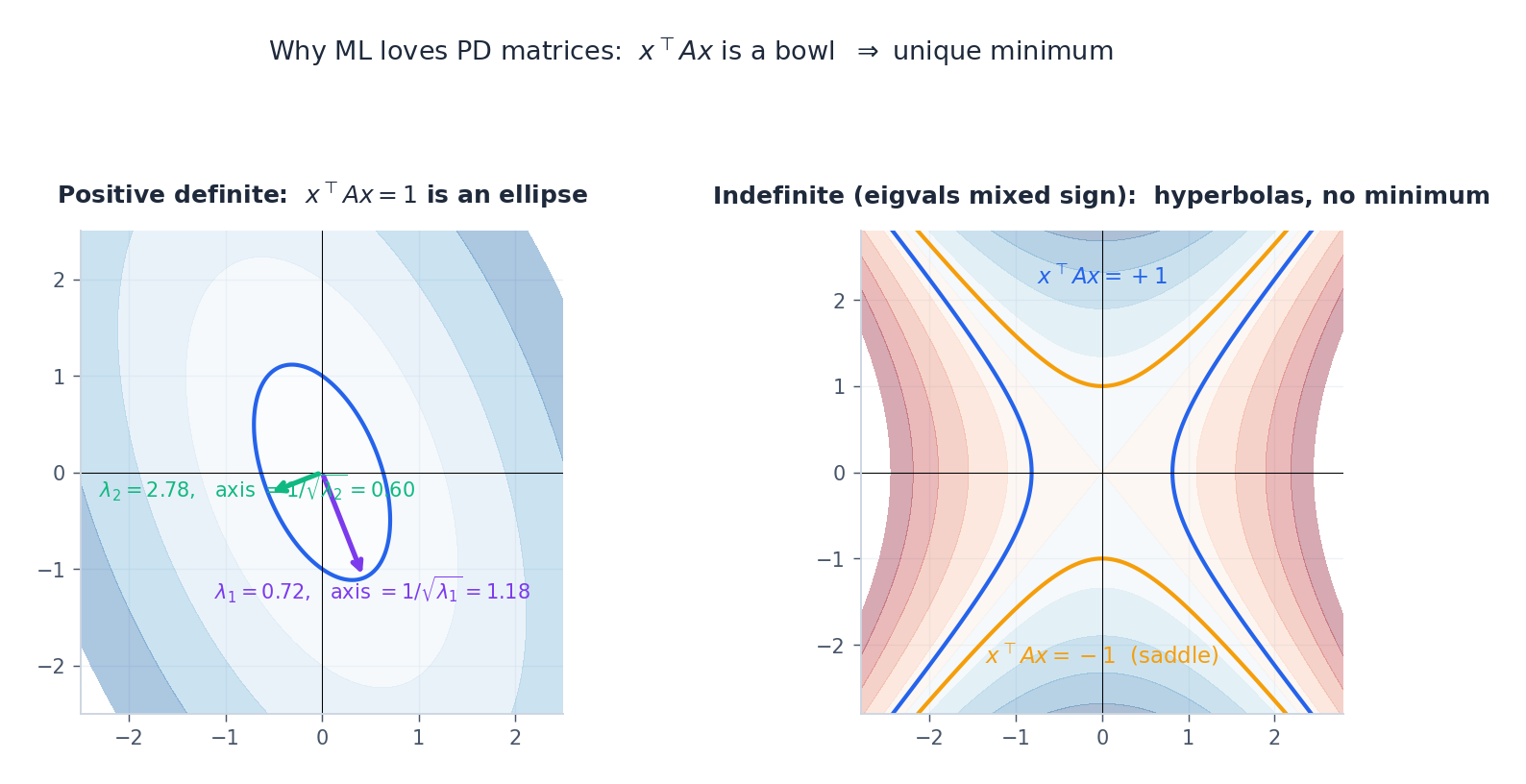
The geometry: $\{x : x^\top A x = 1\}$ is an ellipsoid, with principal axes along the eigenvectors and semi-axis lengths $1/\sqrt{\lambda_i}$ . When some eigenvalue is negative, you get a hyperboloid (saddle) and there is no minimum. This is why convex ML insists on PD Hessians.
Where PD matrices show up:
- Covariance $\Sigma = \mathbb{E}[(x - \mu)(x - \mu)^\top]$ is always PSD.
- Gram matrices $X^\top X$ are PSD; PD iff columns of $X$ are independent.
- Kernels must be PSD (Mercer’s theorem) or you can’t reproduce them in any feature space.
- Hessians of convex losses are PSD; strict convexity = PD.
Singular Value Decomposition — the universal factorisation#

The theorem#
$$A = U \Sigma V^\top, \tag{4}$$with $U \in \mathbb{R}^{m \times m}$ , $V \in \mathbb{R}^{n \times n}$ orthogonal, and $\Sigma$ diagonal with singular values $\sigma_1 \ge \cdots \ge \sigma_r > 0$ (and zeros after).
Why SVD always exists. $A^\top A$ is symmetric PSD, so the spectral theorem gives an orthonormal eigenbasis $V$ with eigenvalues $\sigma_i^2$ . Define $u_i = A v_i / \sigma_i$ for $\sigma_i > 0$ and complete to an orthonormal basis $U$ . Then $A v_i = \sigma_i u_i$ , i.e. $A V = U \Sigma$ , i.e. $A = U \Sigma V^\top$ . $\square$
Geometric picture: rotate, scale, rotate#
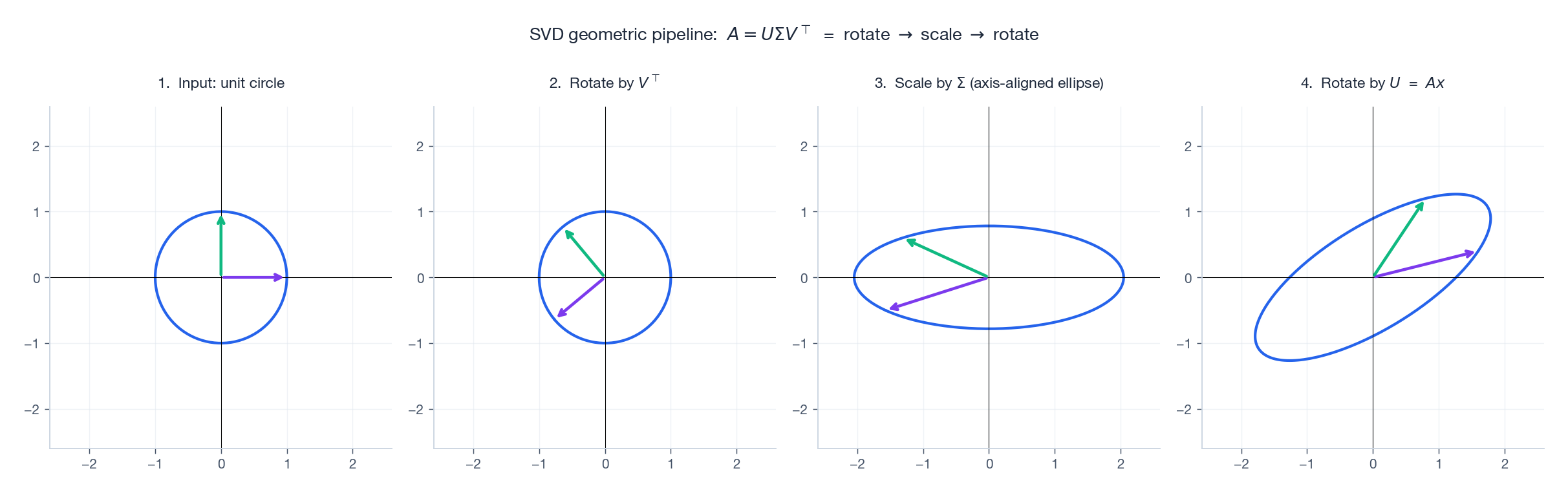
Every linear map decomposes into three motions:
- $V^\top$ : rotate input so its principal axes align with the coordinate axes,
- $\Sigma$ : stretch each axis by $\sigma_i$ (this is the only “non-rigid” step),
- $U$ : rotate the result into the output space.
This is the cleanest mental model in linear algebra: any matrix is a rotation, then an axis-aligned stretch, then another rotation.
What goes wrong without rank: visualised#
When some $\sigma_i = 0$ , that direction in the input is annihilated. The output ellipse collapses to a lower-dimensional shape:
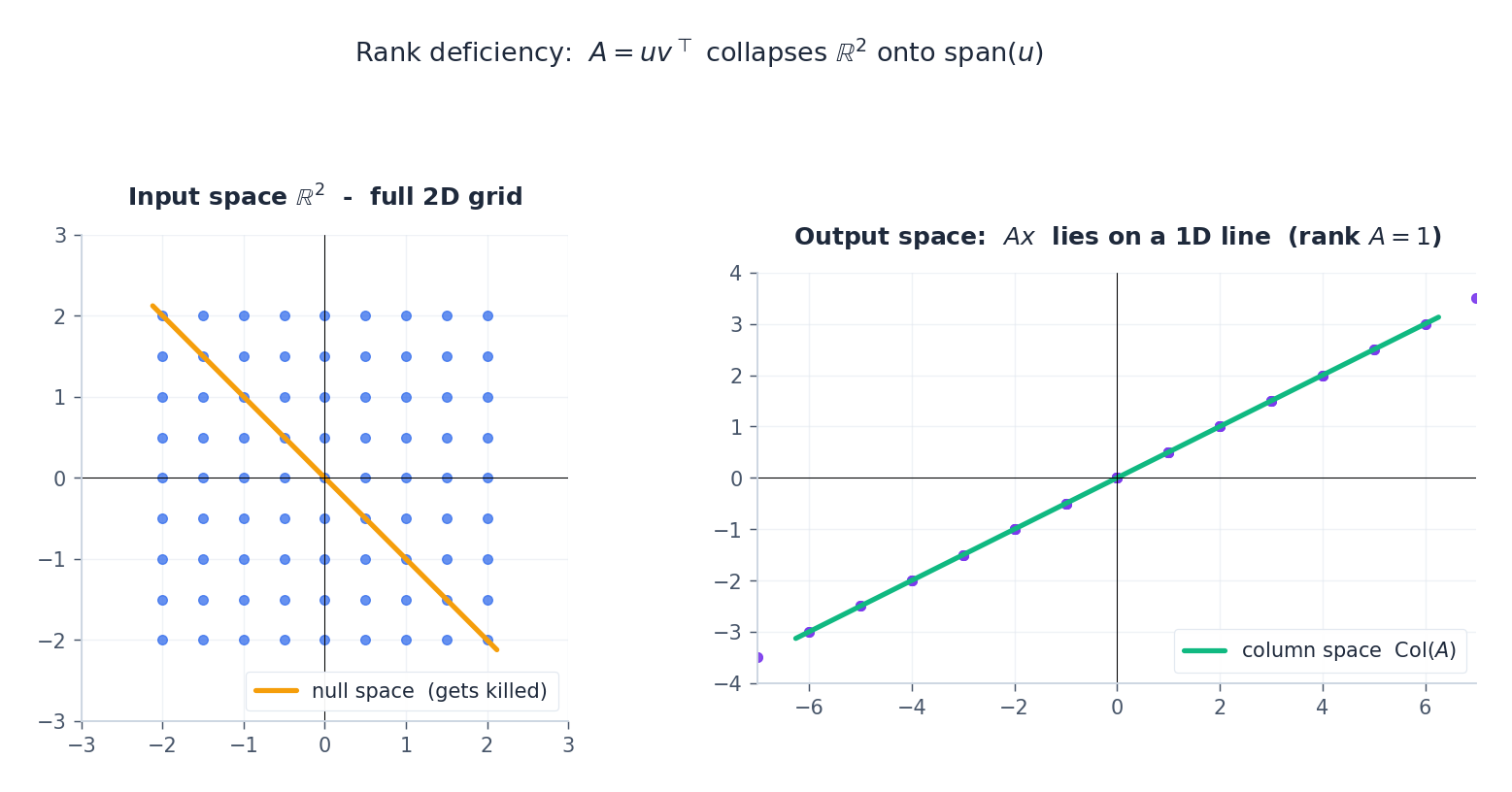
A rank-1 matrix $A = u v^\top$ sends all of $\mathbb{R}^n$ onto the single line $\text{span}(u)$ . Everything orthogonal to $v$ is in the null space and gets sent to $0$ . This is exactly what makes low-rank approximations compress data.
Eckart-Young: best low-rank approximation#
$$A_k = \sum_{i=1}^k \sigma_i u_i v_i^\top, \tag{5}$$with errors $\|A - A_k\|_F = \sqrt{\sum_{i>k} \sigma_i^2}$ and $\|A - A_k\|_2 = \sigma_{k+1}$ .
This is the mathematical engine of PCA. Center your data matrix $X$ , take its SVD; the top-$k$ right singular vectors are the principal components, and the singular values squared (over $n$ ) are the explained variances. It is also the engine of every low-rank technique you have heard of: image compression, latent semantic analysis, recommender systems, LoRA.
Pseudo-inverse and condition number#
The Moore-Penrose pseudo-inverse is computed via SVD: $A^+ = V \Sigma^+ U^\top$
where $\Sigma^+$
inverts the non-zero singular values. Then $x^\star = A^+ b$
is the minimum-norm least-squares solution to $Ax = b$
— exactly what numpy.linalg.lstsq returns.
The condition number $\kappa(A) = \sigma_1 / \sigma_r$ measures how much $A$ amplifies relative perturbations. Roughly, if $\kappa(A) \approx 10^k$ you lose $k$ digits of precision when solving $Ax = b$ . Always log it before trusting a least-squares fit.
Eigendecomposition vs SVD#
| Eigendecomposition | SVD | |
|---|---|---|
| Works on | square matrices | any $m \times n$ |
| Form | $A = P \Lambda P^{-1}$ | $A = U \Sigma V^\top$ |
| Always exists? | only if diagonalisable | always |
| “Values” | may be complex | non-negative real |
| Right factor = inverse of left? | no (general $P$ ) | yes ($U, V$ orthogonal) |
For a symmetric PSD matrix the two coincide ($U = V = Q$ , $\Sigma = \Lambda$ ). Otherwise prefer SVD — it’s stable, total, and applies to rectangular matrices.
Matrix calculus — the engine of optimisation#
This is where most ML readers want a clean reference. We use numerator layout: the derivative has the shape of the output (so a gradient of a scalar w.r.t. a vector is a column vector, matching the input).
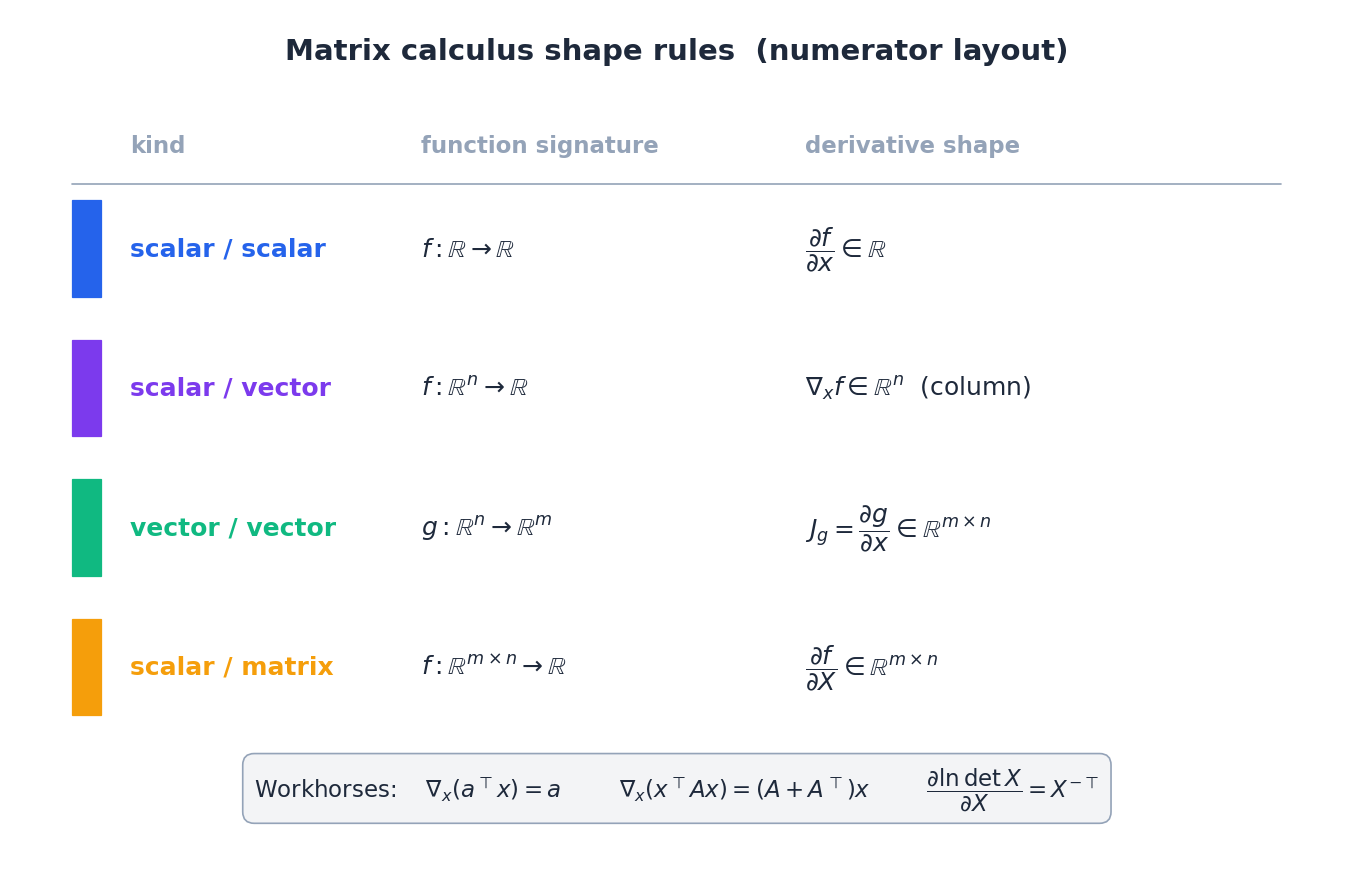

The four formulas you reach for daily#
| # | $f$ | $\nabla f$ | Comment |
|---|---|---|---|
| 1 | $a^\top x$ | $a$ | linear |
| 2 | $x^\top A x$ | $(A + A^\top) x$ | $= 2 A x$ if $A$ symmetric |
| 3 | $\mid A x - b\mid_2^2$ | $2 A^\top (A x - b)$ | least squares |
| 4 | $\ln \det X$ | $X^{-\top}$ | log-det — shows up in Gaussian MLE |
Proof of (2). Write $f = \sum_{i,j} A_{ij} x_i x_j$ . Then $\partial f / \partial x_k = \sum_j A_{kj} x_j + \sum_i A_{ik} x_i = [(A + A^\top) x]_k$ . $\square$
Proof of (3). Apply the chain rule: $\nabla_x \tfrac12 \|Ax - b\|^2 = A^\top (A x - b)$ . Multiplying by 2 absorbs the $\tfrac12$ that ML papers conventionally drop. The optimum solves the normal equations $A^\top A x = A^\top b$ .
Chain rule and backprop#
$$\nabla_x L = J_g^\top \cdot \nabla_g f, \tag{6}$$where $J_g$ is the $m \times n$ Jacobian of $g$ .
$$\delta := \frac{\partial L}{\partial a} \odot \sigma'(z), \quad \frac{\partial L}{\partial W} = \delta\, x^\top, \quad \frac{\partial L}{\partial b} = \delta. \tag{7}$$That’s the entire algorithm — everything PyTorch’s autograd engine does is run (6) on a computation graph.
Sanity check by finite differences#
$$\hat g_i = \frac{f(x + \varepsilon e_i) - f(x - \varepsilon e_i)}{2 \varepsilon}.$$Match against the analytic $\nabla f$ to relative error $\approx 10^{-6}$ at $\varepsilon = 10^{-5}$ . The code in section 7 does exactly this.
Numerical decompositions — which one and when#
QR for least squares#
Any full-column-rank $A$ factors as $A = QR$ with $Q$ orthonormal columns and $R$ upper triangular. To solve $\min_x \|Ax - b\|$ , compute $Q^\top b$ and back-substitute on $R x = Q^\top b$ .
Why prefer QR over normal equations. Normal equations form $A^\top A$ , whose condition number is $\kappa(A)^2$ . If $\kappa(A) = 10^4$ that’s $10^8$ — you’ve squared your error budget. QR works directly on $A$ , so it’s $\kappa(A)$ .
Cholesky for PD systems#
For PD $A$ , the Cholesky factorisation $A = L L^\top$ exists with $L$ lower triangular and positive diagonal. Cost $O(n^3 / 3)$ , half of LU. Solving $Ax = b$ is then two triangular solves. This is the workhorse for Gaussian-process inference, kernel methods, and any quadratic program with PD Hessian.
Decision table#
| Situation | Use |
|---|---|
| Symmetric matrix (covariance, Hessian) | Eigendecomposition / spectral |
| Any matrix; want “honest” rank, PCA, pseudo-inverse | SVD |
| Tall least squares $\min \mid Ax - b\mid$ | QR |
| PD linear system $Ax = b$ | Cholesky |
| Detect rank deficiency / monitor numerical health | $\kappa(A)$ via SVD |
Code: verify everything#
| |
Three things to notice in the output: the Eckart-Young error matches theory to machine precision; QR is one to two orders of magnitude more accurate than normal equations even on benign matrices; the analytic gradient agrees with finite differences to ~$10^{-9}$ .
Q&A — the questions that come up#
Q1. When should I trust normal equations? When $\kappa(A) < 10^3$ or so — e.g. small problems with well-scaled features. Anything ill-conditioned: use QR or SVD. Squaring the condition number really does eat your precision.
Q2. Why does $\ell_1$ produce sparse solutions? The $\ell_1$ unit ball has corners on the coordinate axes. Generic optimisation contours hit these corners first, and at a corner some coordinates are exactly zero. The smooth $\ell_2$ ball has no corners and so generically the optimum lies off the axes.
Q3. SVD vs PCA — the same thing? Yes, modulo bookkeeping. With centred data $X \in \mathbb{R}^{n \times d}$ :
- PCA via covariance: $\Sigma = \tfrac{1}{n} X^\top X$ , top eigenvectors = principal directions.
- PCA via SVD: $X = U \Sigma V^\top$ , columns of $V$ = principal directions; $\sigma_i^2 / n$ = explained variances. For high-dimensional data ($d \gg n$ ) the SVD path is dramatically cheaper because you never materialise the $d \times d$ covariance.
Q4. Why are eigenvalues of symmetric matrices special? They are real (so we can order them), and the eigenvectors form an orthonormal basis. That orthonormality is what lets us write any vector as $\sum \langle v, q_i\rangle q_i$ — a clean coordinate system for the action of $A$ . The Hessians, covariances, Grams, and Laplacians of ML are all symmetric, which is why this matters every day.
Q5. What does “rank” really mean operationally? The rank is how much information $A$ preserves about its input. A rank-$r$ matrix in $\mathbb{R}^{m \times n}$ throws away an $(n - r)$ -dimensional subspace (its null space) and outputs into an $r$ -dimensional subspace (its column space). In ML this is exactly the bottleneck dimension of an autoencoder, the rank in matrix completion, or the latent dimension of an embedding.
Summary#
| Tool | Key formula | When to reach for it |
|---|---|---|
| Spectral theorem | $A = Q \Lambda Q^\top$ | symmetric matrices: covariance, Hessian |
| SVD | $A = U \Sigma V^\top$ | any matrix: PCA, pseudo-inverse, low-rank |
| QR | $A = QR$ | least squares with bad conditioning |
| Cholesky | $A = LL^\top$ | PD linear systems (Gaussian MLE, GP) |
| Quadratic gradient | $\nabla (x^\top A x) = 2 A x$ | every quadratic objective |
| Chain rule | $\nabla_x L = J_g^\top \nabla_g f$ | backpropagation in one line |
What’s next#
Linear algebra makes “parameter” and “gradient” concrete as vectors and matrices, but it is purely deterministic. Real data is noisy, models make mistakes, and estimates need confidence intervals — all the language of probability. The next chapter swaps vocabulary: random variable, expectation, variance, likelihood, posterior.
I deliberately walk the main line — Bayes’ rule, law of large numbers, central limit theorem, MLE/MAP — end to end, because every later chapter assumes you can switch fluently between these ideas. Linear regression will use MLE to explain least squares; logistic regression will use MLE to explain cross-entropy; EM will use MLE to explain latent variables; regularization will use MAP to explain penalties. So this chapter is “statistical inference through a probabilistic lens” — not a probability course in disguise, but a way to reduce every ML algorithm to “estimating parameters of some likelihood”.
References#
- Strang, G. (2023). Introduction to Linear Algebra (6th ed.). Wellesley-Cambridge Press.
- Trefethen, L. N. & Bau III, D. (1997). Numerical Linear Algebra. SIAM. — the cleanest treatment of stability.
- Golub, G. H. & Van Loan, C. F. (2013). Matrix Computations (4th ed.). Johns Hopkins University Press. — the algorithmic bible.
- Petersen, K. B. & Pedersen, M. S. (2012). The Matrix Cookbook. Technical University of Denmark. — the gradient lookup table everyone uses.
- Eckart, C. & Young, G. (1936). The approximation of one matrix by another of lower rank. Psychometrika, 1(3), 211-218.
- Boyd, S. & Vandenberghe, L. (2018). Introduction to Applied Linear Algebra. Cambridge University Press. — great companion focused on data.
ML Math Derivations 20 parts
- 01 ML Math Derivations (1): Introduction and Mathematical Foundations
- 02 ML Math Derivations (2): Linear Algebra and Matrix Theory you are here
- 03 ML Math Derivations (3): Probability Theory and Statistical Inference
- 04 ML Math Derivations (4): Convex Optimization Theory
- 05 ML Math Derivations (5): Linear Regression
- 06 ML Math Derivations (6): Logistic Regression and Classification
- 07 ML Math Derivations (7): Decision Trees
- 08 ML Math Derivations (8): Support Vector Machines
- 09 ML Math Derivations (9): Naive Bayes
- 10 ML Math Derivations (10): Semi-Naive Bayes and Bayesian Networks
- 11 ML Math Derivations (11): Ensemble Learning
- 12 ML Math Derivations (12): XGBoost and LightGBM
- 13 ML Math Derivations (13): EM Algorithm and GMM
- 14 ML Math Derivations (14): Variational Inference and Variational EM
- 15 ML Math Derivations (15): Hidden Markov Models
- 16 ML Math Derivations (16): Conditional Random Fields
- 17 ML Math Derivations (17): Dimensionality Reduction and PCA
- 18 ML Math Derivations (18): Clustering Algorithms
- 19 ML Math Derivations (19): Neural Networks and Backpropagation
- 20 ML Math Derivations (20): Regularization and Model Selection