
ML Math Derivations (7): Decision Trees
From information entropy to the Gini index, from ID3 to CART — a complete derivation of decision-tree mathematics: split criteria, continuous and missing values, pruning, and feature importance, with sklearn-verified figures.
Hook. A decision tree mimics how humans actually decide things: ask a question, branch on the answer, ask the next question. The math under that intuition is surprisingly rich — entropy from information theory tells us which question to ask first, the Gini index gives a cheaper proxy that lands on essentially the same trees, and cost-complexity pruning gives a principled way to stop the tree from memorising noise. Almost every modern boosted ensemble (XGBoost, LightGBM, CatBoost) is just a clever sum of these objects, so getting the foundations right pays off many times over.

What You Will Learn#
- Why a decision tree is mathematically a piecewise-constant function on a recursive partition of the feature space.
- The information-theoretic motivation for entropy, conditional entropy, and information gain — and why gain ratio was invented.
- Why Gini and entropy almost always pick the same split (a one-line Taylor argument).
- How CART handles continuous features, missing values, and categorical features without breaking its greedy framework.
- Pre-pruning vs post-pruning, and the exact derivation of the cost-complexity threshold $\alpha$ .
- How feature importance is defined (not just printed by sklearn) and where it can be misleading.
Decision Tree Fundamentals#
Model Representation#
$$f(x) = \sum_{m=1}^{M} c_m \, \mathbb{1}[x \in R_m].$$ $$\text{node test:} \quad x_j \leq \tau \quad \text{vs.} \quad x_j > \tau.$$The tree itself is a hierarchical structure of:
- Root node: contains all training samples.
- Internal nodes: each tests one feature against a threshold (or one categorical value).
- Branches: outcomes of the test.
- Leaf nodes: terminal regions storing the prediction.
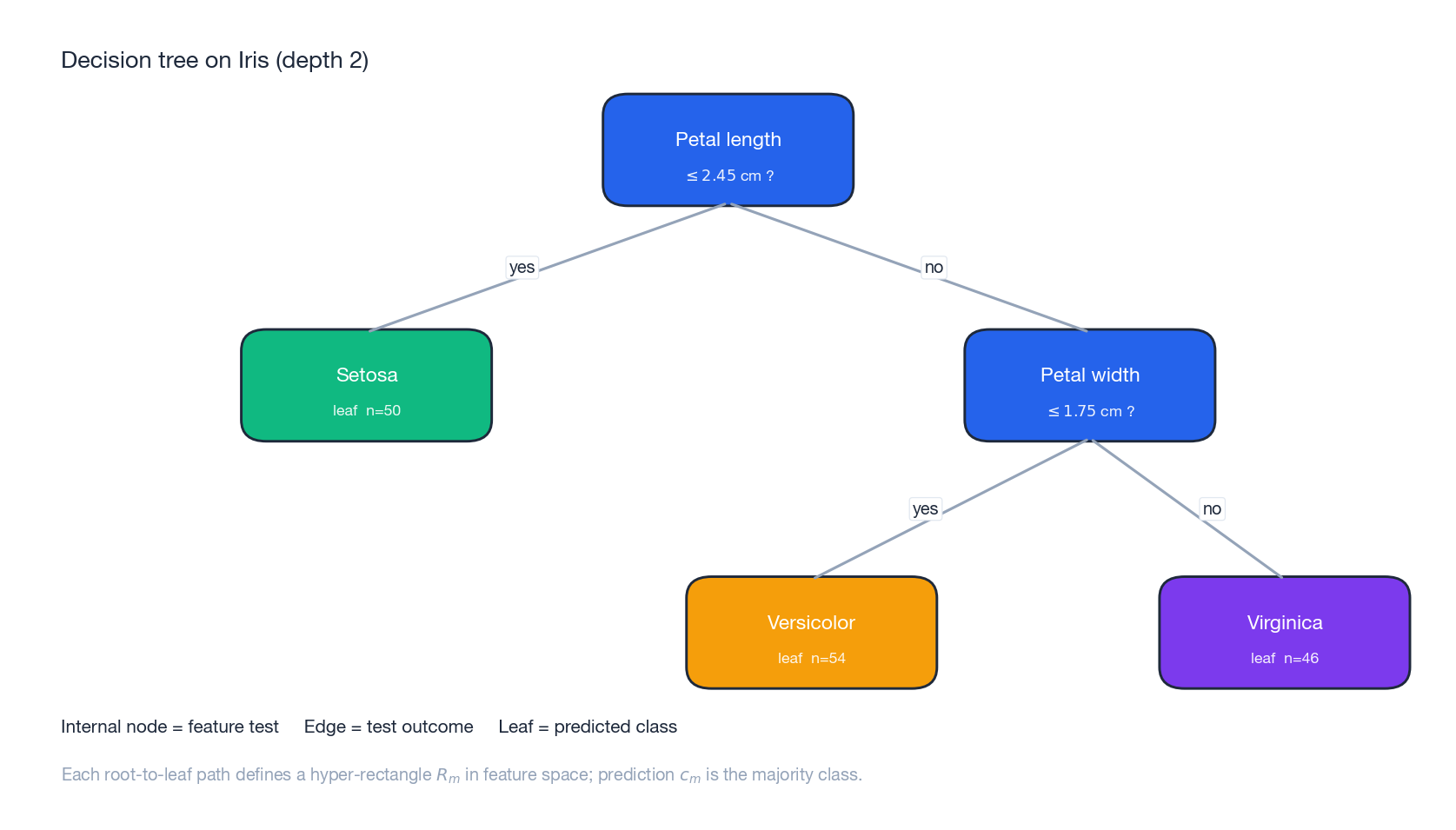
The figure above shows a depth-2 CART trained on Iris. Two questions about petal dimensions are enough to separate three species — every root-to-leaf path corresponds to one rectangle in feature space.
Why Trees Are Useful — and Where They Fail#
| Strengths | Weaknesses |
|---|---|
| Decision paths are human-readable | Greedy growth can miss the global optimum |
| Insensitive to feature scaling and monotone transforms | High variance — small data perturbations change the tree drastically |
| Handles numerical and categorical features uniformly | Linear relationships need many axis-aligned steps to approximate |
| Naturally captures feature interactions | Single trees usually overfit unless pruned |
The variance problem is exactly what bagging (random forests) and boosting (GBDT, XGBoost) were designed to fix.
Information-Theoretic Foundations#
Entropy: Uncertainty as a Function of a Distribution#
$$H(Y) = -\sum_{k=1}^{K} p_k \log_2 p_k, \qquad 0 \log_2 0 \triangleq 0.$$In bits, $H$ measures the average number of yes/no questions needed to identify $Y$ .
Three properties we will use repeatedly.
- Non-negativity. $H(Y) \geq 0$ with equality iff some $p_k = 1$ (no uncertainty).
- Maximum at the uniform distribution. Maximising $-\sum_k p_k \log p_k$ subject to $\sum_k p_k = 1$ via Lagrange multipliers yields $p_k = 1/K$ and $H_{\max} = \log_2 K$ .
- Concavity. $H$ is strictly concave on the probability simplex, so mixing distributions never decreases entropy.
Conditional Entropy and Information Gain#
$$H(Y \mid X) = \sum_{v=1}^{V} p(X = v) \, H(Y \mid X = v)$$ $$IG(Y, X) \;=\; H(Y) - H(Y \mid X) \;\geq\; 0.$$Non-negativity follows from concavity of $H$ (Jensen’s inequality), and $IG = 0$ iff $X$ is independent of $Y$ .
ID3 chooses the feature with the largest $IG$ at every node.
Gain Ratio: Penalising Many-Valued Features#
Information gain has a well-known pathology: a feature with one unique value per sample (e.g. an ID column) sends every sample into its own bucket, makes every child entropy zero, and so wins every split — yet it has zero generalisation value.
$$IV(X) = -\sum_{v=1}^{V} \frac{|S_v|}{|S|} \log_2 \frac{|S_v|}{|S|},$$ $$GR(Y, X) = \frac{IG(Y, X)}{IV(X)}.$$Many-valued features have large $IV$ and so are penalised. C4.5’s actual heuristic is to first restrict to features whose $IG$ is above the average, then pick the highest gain ratio inside that pool — this avoids favouring under-informative features whose tiny $IV$ artificially inflates $GR$ .
Splitting Criteria#

Gini Index#
$$G(S) = 1 - \sum_{k=1}^{K} p_k^2 = \sum_{k=1}^{K} p_k (1 - p_k),$$where $p_k$ is the proportion of class $k$ in node $S$ . It is the probability that two samples drawn at random from $S$ belong to different classes. Like entropy, it is zero for a pure node and maximal at the uniform distribution.
$$\Delta G = G(S) - \sum_{v=1}^{V} \frac{|S_v|}{|S|} G(S_v).$$CART picks the split that maximises $\Delta G$ .
Why Gini and Entropy Behave Almost Identically#
$$G(p) = 2p(1-p), \qquad H(p) = -p \log_2 p - (1-p) \log_2 (1-p).$$ $$H_\mathrm{nat}(p) \;\approx\; \ln 2 \;-\; 2\,(p - \tfrac{1}{2})^2,$$ $$G(p) = \tfrac{1}{2} - 2\,(p - \tfrac{1}{2})^2.$$After the constant scaling $H_\mathrm{nat}/(2 \ln 2)$ , the two functions have the same leading-order shape near $p = \tfrac{1}{2}$ . In practice CART trained with Gini and with entropy almost always splits the same way.
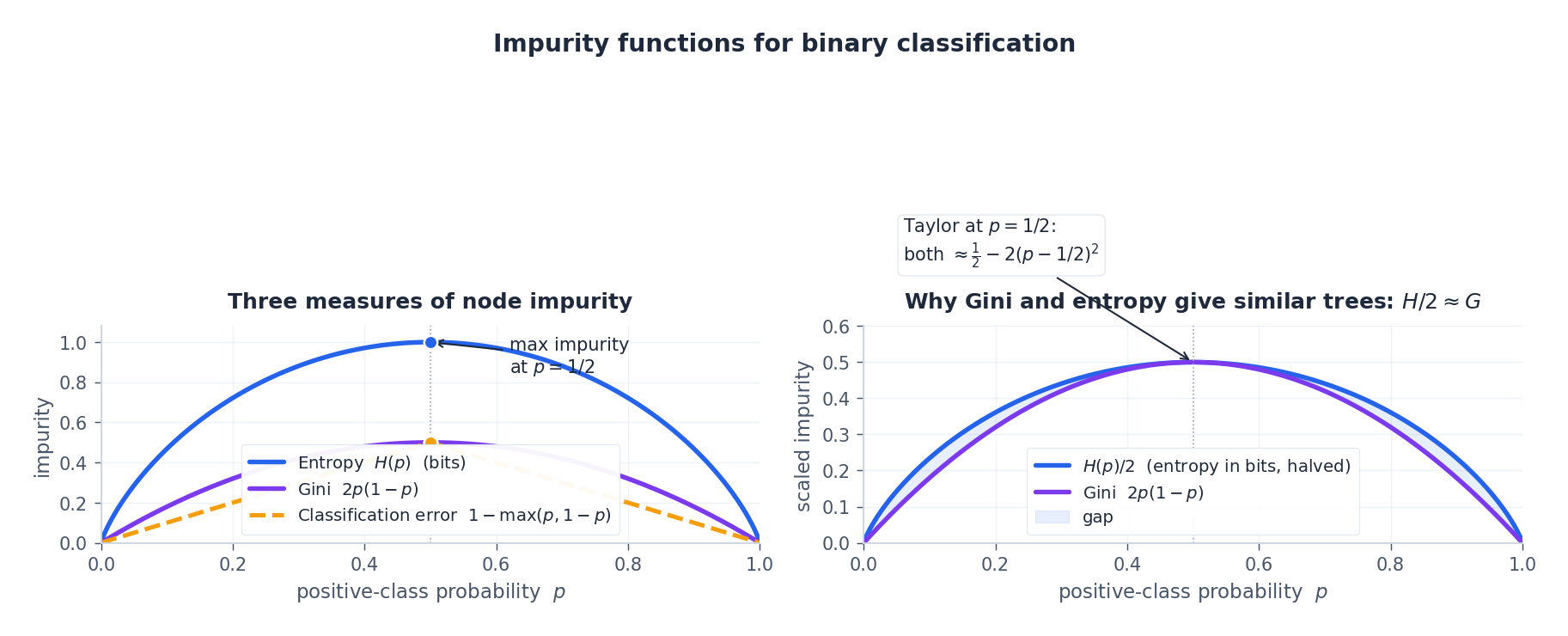
The left panel compares all three: Gini, entropy, and the classification error rate $1 - \max(p, 1-p)$ . The right panel overlays $H/2$ on $G$ — they agree to second order around $p = 1/2$ and differ only near the corners. Classification error, by contrast, is piecewise linear and not strictly concave, which is exactly why it makes a bad splitting criterion: a split that improves both children’s purity may leave the weighted error unchanged, so the search gets stuck.
Mean Squared Error for Regression Trees#
$$\hat{c}_m = \frac{1}{|R_m|} \sum_{i \in R_m} y_i,$$ $$\min_{j, \tau} \;\Big[\sum_{i: x_{ij} \leq \tau} (y_i - \hat{c}_L)^2 \;+\; \sum_{i: x_{ij} > \tau} (y_i - \hat{c}_R)^2\Big].$$ $$\Delta = \mathrm{Var}(y_S) - \frac{|S_L|}{|S|}\mathrm{Var}(y_{S_L}) - \frac{|S_R|}{|S|}\mathrm{Var}(y_{S_R}).$$Decision Tree Algorithms#
ID3, C4.5, CART at a Glance#
| ID3 (1986) | C4.5 (1993) | CART (1984) | |
|---|---|---|---|
| Tree shape | Multi-way | Multi-way | Binary |
| Criterion (cls) | Information gain | Gain ratio | Gini |
| Criterion (reg) | — | — | MSE |
| Continuous features | Discretise upfront | Threshold search | Threshold search |
| Missing values | — | Surrogate weighting | Surrogate splits |
| Pruning | None | Pessimistic post-pruning | Cost-complexity |
The descriptions below follow CART because that is what scikit-learn implements.
Greedy Recursive Construction#
| |
stop typically combines: a class is pure, no informative split exists, depth limit reached, or fewer than min_samples_split samples remain. The greedy choice is locally optimal — finding the globally optimal tree is NP-hard, even for binary features.
Continuous Features and Missing Values#
Continuous Features by Threshold Search#
Sort the values of feature $j$ as $x_{(1)} < \dots < x_{(n)}$ . Candidate thresholds sit at the midpoints $\tau_i = (x_{(i)} + x_{(i+1)})/2$ . For each $\tau_i$ we evaluate the split criterion. Naively this is $O(n^2)$ per feature; pre-sorting once and updating impurity counts incrementally as samples cross the threshold brings it down to $O(n \log n)$ .
A well-known property: the optimal threshold always lies between two adjacent samples whose labels differ. This shrinks the candidate set substantially for skewed labels.
Missing Values#
C4.5 and CART handle missing values without imputation.
$$IG_\text{adj}(Y, X_j) = \frac{|S_\text{obs}|}{|S|} \cdot IG(Y_\text{obs}, X_j),$$so features with many missing values are automatically discounted.
$$w_L = \frac{|S_L^\text{obs}|}{|S^\text{obs}|}, \qquad w_R = \frac{|S_R^\text{obs}|}{|S^\text{obs}|}.$$CART additionally stores surrogate splits at each node — backup features that approximate the primary split — and uses the best surrogate when the primary feature is missing at prediction time.
Categorical Features#
For an unordered feature with $K$ categories, an exhaustive binary split requires testing $2^{K-1} - 1$ subsets. For binary classification (or regression with squared loss) there is a remarkable shortcut: sort categories by their mean response (Fisher 1958, Breiman 1984), and the optimal binary split is along that ordering. This brings the search from exponential to linear in $K$ .
Splits as Information#
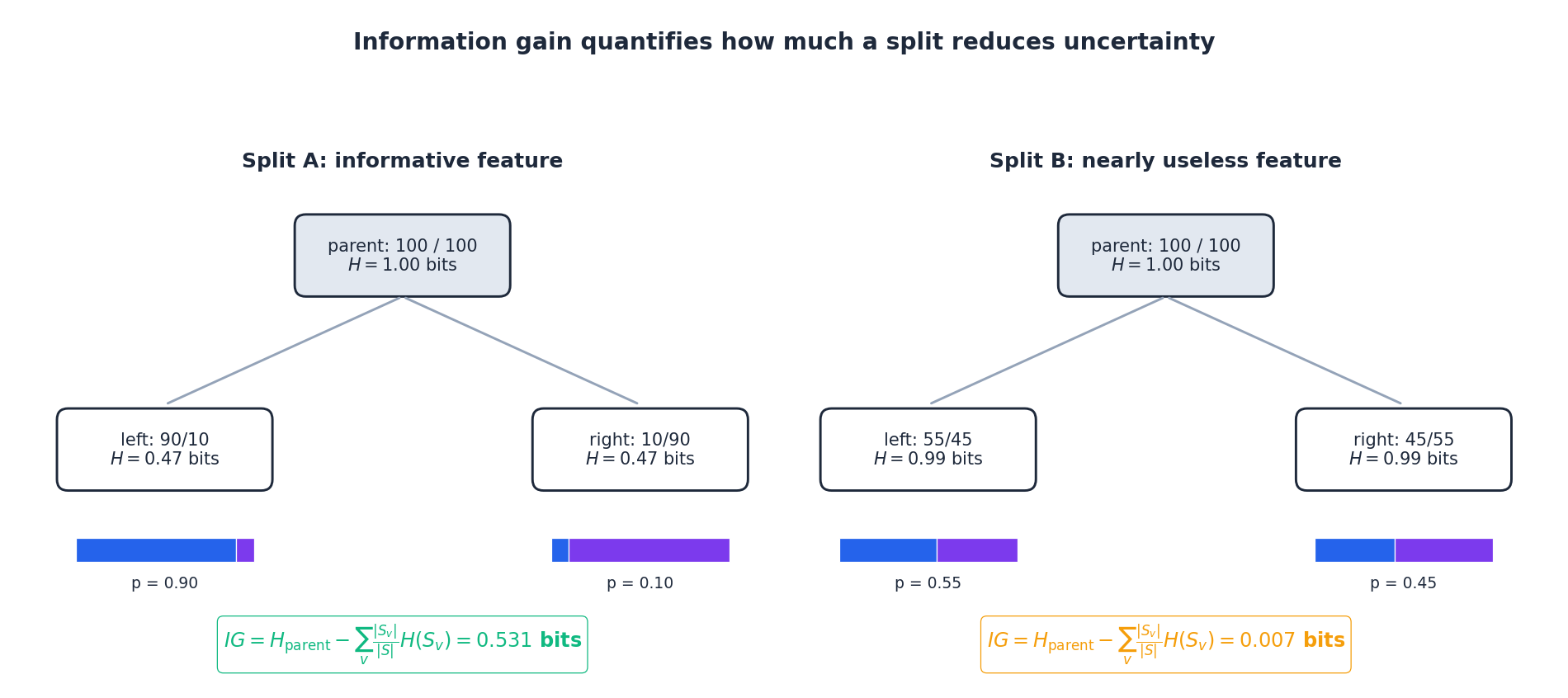
The figure illustrates the gain calculation on two synthetic splits of the same parent ($H = 1$ bit). The informative feature on the left drives both children near purity, recovering most of the parent’s entropy. The uninformative feature on the right barely moves the needle — its weighted child entropy is almost as high as the parent’s, so $IG$ is near zero. CART picks the feature/threshold pair with the largest such bar.
Decision Boundaries in 2D#
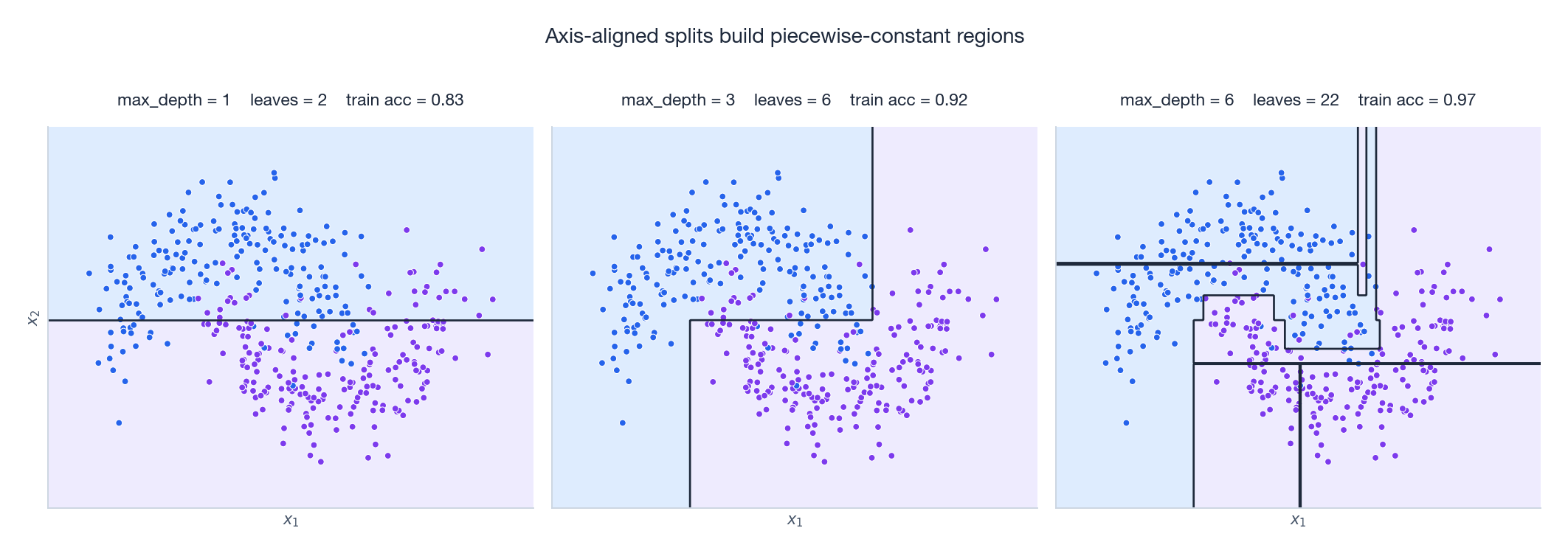
These boundaries make the piecewise-constant nature of trees visceral. At depth 1 we have a single vertical or horizontal cut — the tree behaves like a one-dimensional threshold. By depth 3 the tree carves the moons out roughly. At depth 6 the boundary becomes ragged: each tiny rectangle is chasing one or two training points, the classic signature of overfitting on a noisy dataset.
Pruning: Controlling Variance#
Why Unpruned Trees Overfit#
A fully grown tree has zero training error: every leaf contains samples of one class (or one sample). The model has effectively memorised the training set, including its noise. Two structural causes:
- Depth. Deep trees express ever finer feature interactions, eventually fitting noise.
- Tiny leaves. Class proportions in a 3-sample leaf are statistically meaningless.
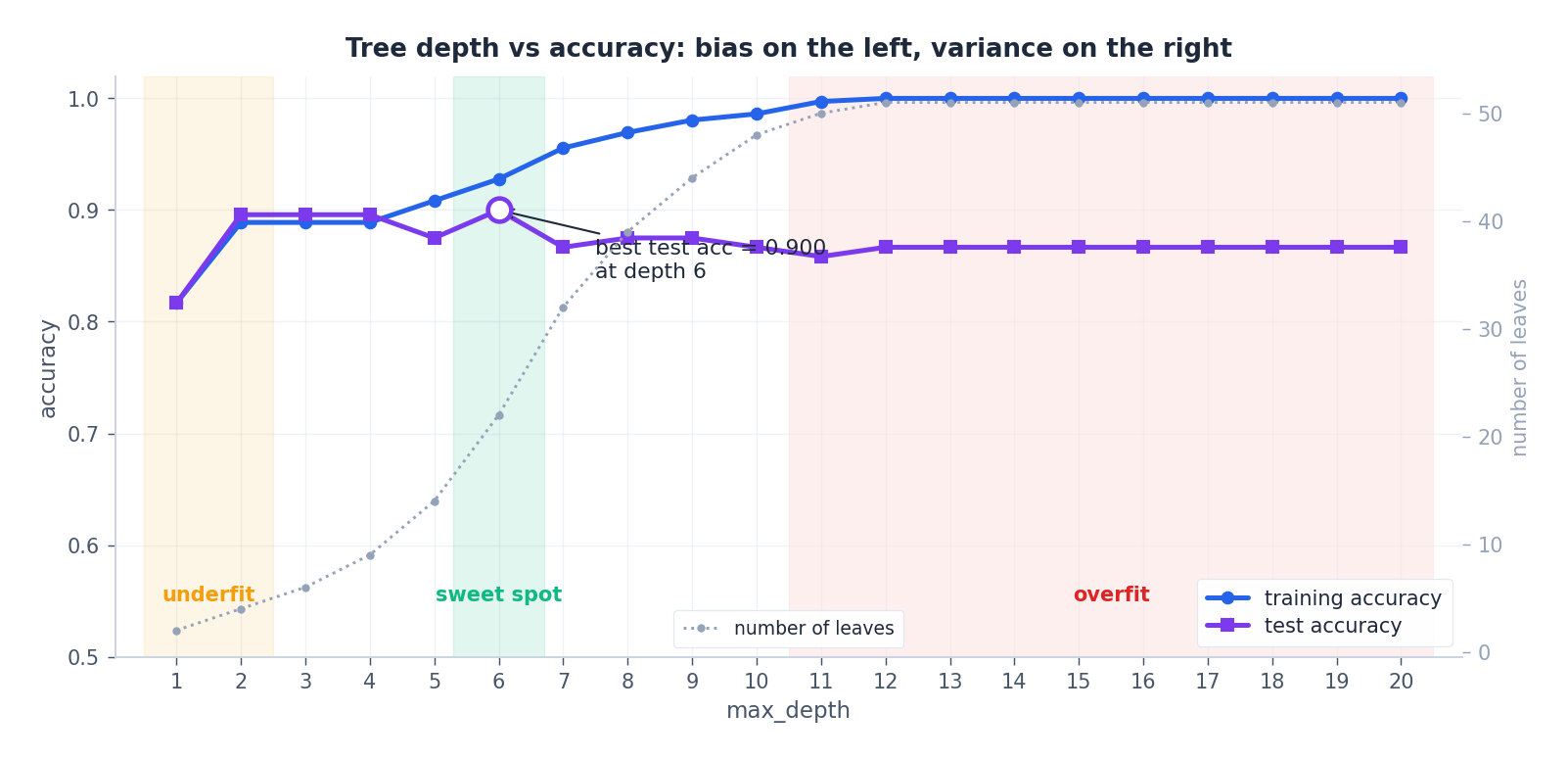
This is the canonical decision-tree overfitting curve, computed on a noisy moons dataset with depth ranging from 1 to 20. Training accuracy rises monotonically toward 1.0, while test accuracy peaks at a moderate depth and then drifts downward. The grey dotted curve on the secondary axis tracks the number of leaves — its near-exponential growth past the sweet spot is the variance the test curve is paying for.
Pre-Pruning#
Stop growing early. Standard knobs:
max_depthmin_samples_split(don’t split nodes with fewer samples)min_samples_leaf(every leaf must have at least this many samples)min_impurity_decrease(require a minimum gain)max_leaf_nodes(cap leaves and split greedily by best gain)
Pre-pruning is fast — it avoids ever building the full tree — but myopic. A split that looks weak now might enable two great splits below; pre-pruning forecloses on it.
Post-Pruning: Cost-Complexity#
$$R_\alpha(T) = R(T) + \alpha \, |T|,$$where $R(T)$ is the training risk (sum of leaf impurities weighted by leaf size) and $|T|$ is the number of leaves. Larger $\alpha$ favours simpler trees.
$$\boxed{\;\alpha_\text{eff}(t) \;=\; \frac{R(t) - R(T_t)}{|T_t| - 1}\;}$$For $\alpha$
above this threshold, collapsing the subtree strictly improves $R_\alpha$
. Sweeping $\alpha$
from $0$
upward and repeatedly collapsing the weakest link (smallest $\alpha_\text{eff}$
) produces a finite, nested sequence of trees $T_0 \supset T_1 \supset \dots \supset T_\text{root}$
. The corresponding $\alpha$
values are exactly what scikit-learn returns from cost_complexity_pruning_path.
The final $\alpha$ is selected by cross-validation on the training set.
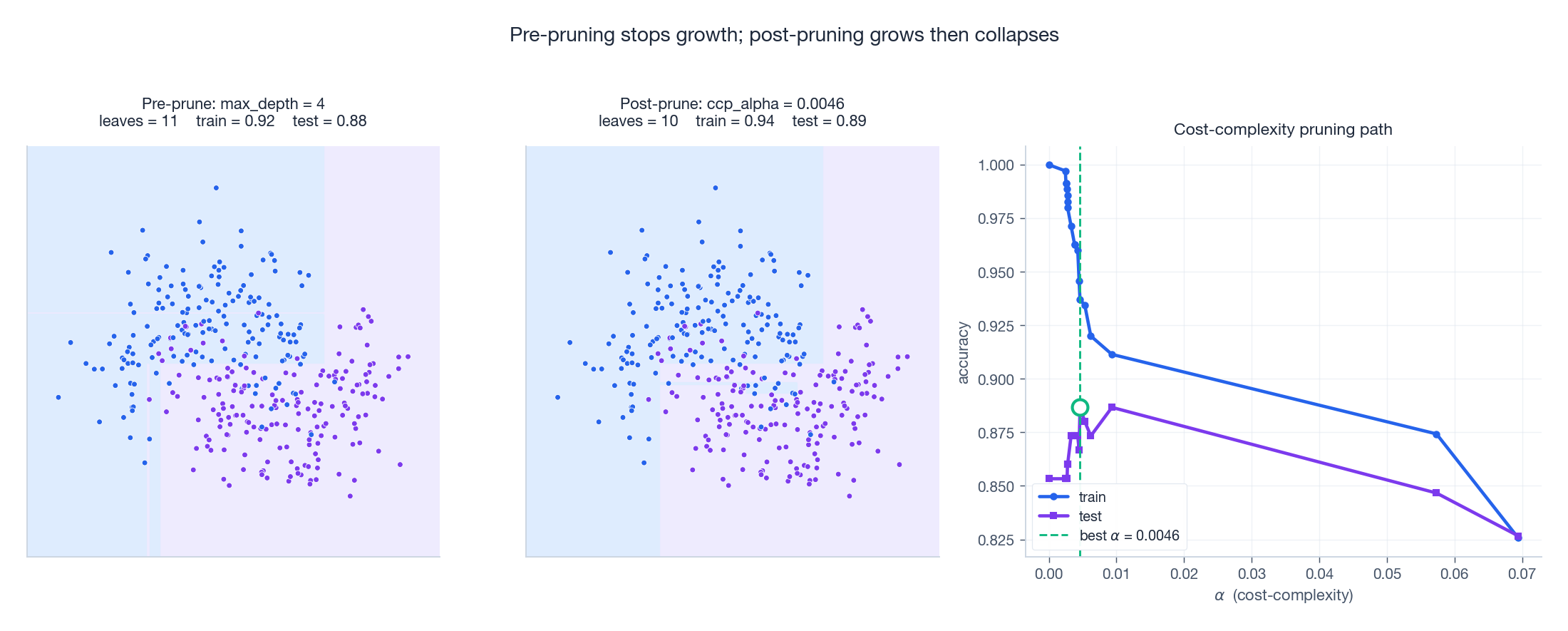
The left and middle panels show the resulting decision regions — pre-pruning at max_depth=4 produces neat large blocks; post-pruning grows fully and then collapses the weakest links, ending with a tree that often has fewer leaves than the depth-bounded one yet generalises better on this dataset. The right panel is the pruning path: as $\alpha$
grows, training accuracy decays smoothly while test accuracy rises, peaks, and then collapses when the tree shrinks too far.
Decision-Tree Theory#
VC Dimension#
$$\text{VCdim} = O(L \log L).$$ $$R(T) \leq \hat{R}(T) + \mathcal{O}\!\left(\sqrt{\frac{L \log L \cdot \log n + \log(1/\delta)}{n}}\right).$$The leaf count $L$ is the natural complexity term — exactly what cost-complexity pruning is regularising.
Why Trees Are Unstable#
If two features have nearly equal information gain at the root, an arbitrarily small perturbation can flip the choice and cascade into a completely different tree below. Formally, let the gains be $g_1, g_2$ with $g_1 - g_2 = \epsilon$ ; any data perturbation that changes a single child impurity by more than $\epsilon$ may invert the order. This high variance is exactly why bagging multiple bootstrap-sampled trees — i.e. random forests — works so well: averaging predictions from many trees with decorrelated errors cuts the variance roughly in proportion to the number of trees.
Feature Interactions and Tree Depth#
$$[x_{j_1} \leq \tau_1] \wedge [x_{j_2} \leq \tau_2] \wedge \dots \wedge [x_{j_d} \leq \tau_d].$$So a depth-$d$ tree expresses interactions of order at most $d$ . This is why even shallow trees beat linear models on data where two features only become predictive together.
Implementation: A Minimal CART#
| |
Feature Importance#
$$\Delta I(t) \;=\; \frac{N_t}{N} \left( I(t) - \frac{N_{t,L}}{N_t} I(t_L) - \frac{N_{t,R}}{N_t} I(t_R) \right).$$The importance of feature $j$ is the sum of $\Delta I(t)$ over all nodes that split on $j$ , normalised so the importances sum to one.
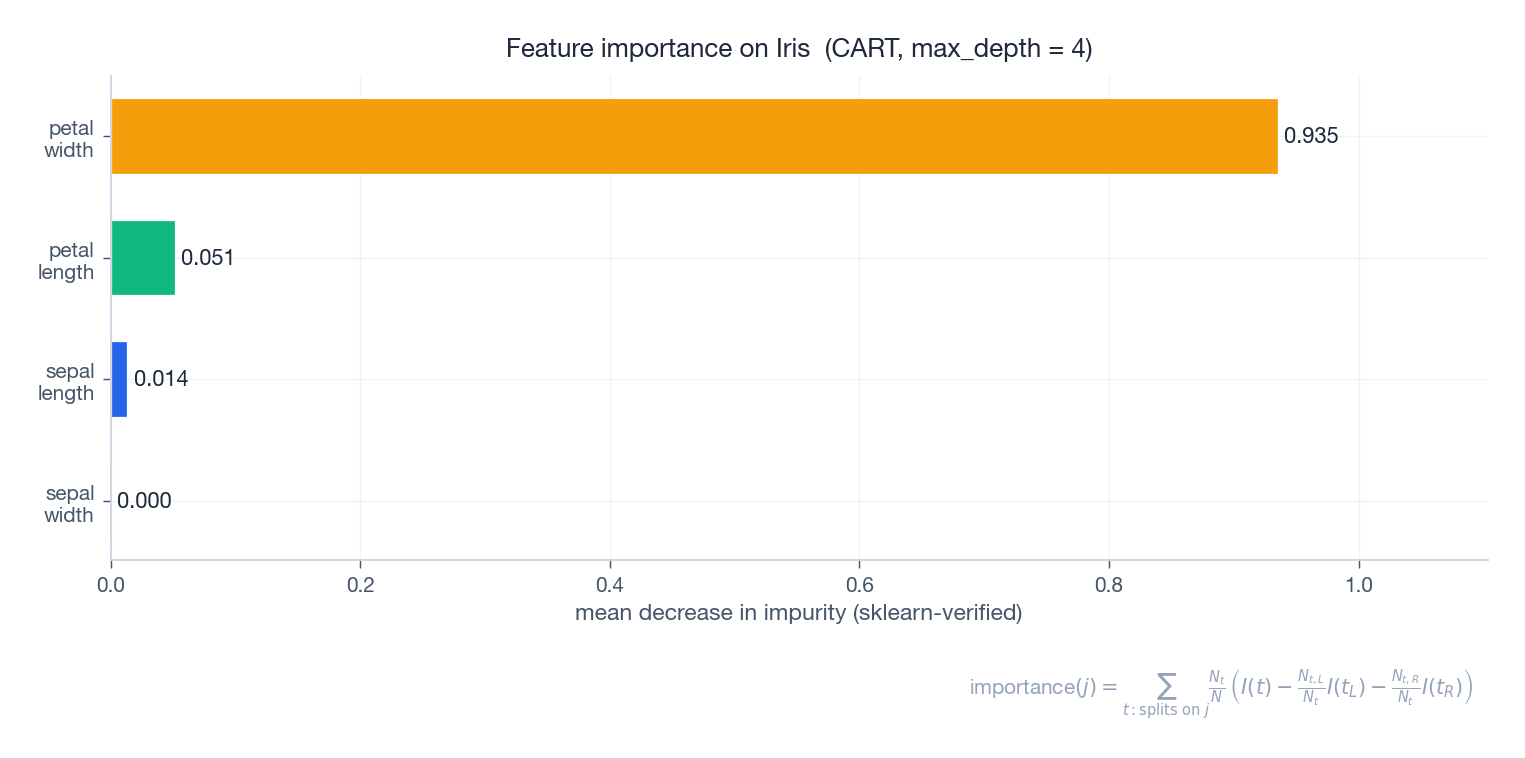
On Iris, petal-length and petal-width dominate, in agreement with the depth-2 tree we drew earlier — those two features alone separate the three species. The figure was produced by sklearn’s feature_importances_ and the formula in the caption matches one-to-one.
Caveats.
- MDI is biased toward continuous and high-cardinality features — they get more chances to be selected as split candidates.
- Importances on a single tree are unstable. Aggregating across a random forest (or using permutation importance) gives more trustworthy rankings.
- Importance is not causality. A feature can be highly important because it correlates with the true cause.
Multivariate Decision Trees#
$$\text{node test:} \quad w^\top x \leq \tau,$$a hyperplane in the feature space. They can represent diagonal boundaries with one node instead of dozens of axis-aligned steps. The trade-off:
- Finding the optimal $w$ at each node is NP-hard in general (heuristics: linear discriminant, perceptron, soft optimisation).
- Interpretability drops: each node is now a small linear model rather than a single threshold.
Q&A Highlights#
Why can decision trees model non-linear relationships?#
Each individual split is linear, but the composition of axis-aligned splits is a piecewise-constant function on a recursive partition. Any continuous function can be approximated arbitrarily well in $L^1$ by such piecewise constants — so trees are universal approximators for bounded, integrable functions.
Why does information gain favour many-valued features?#
Because finer partitions are mechanically more pure: in the limit of one sample per child, every child entropy is zero and the gain equals the parent entropy. This says nothing about generalisation. C4.5’s gain ratio divides by $IV(X)$ — itself an entropy term — to normalise away this artefact.
Gini or entropy — does it matter?#
Almost never. Their second-order Taylor expansions around $p = 1/2$ agree, and empirically the resulting trees differ in only a small minority of splits. Gini is slightly faster (no log) and is the default in scikit-learn’s CART.
How are categorical features handled?#
Three options: (i) one-hot encode and let the tree split each indicator; (ii) multi-way split with one branch per category (ID3/C4.5); (iii) optimal binary split — for binary classification this reduces to sorting categories by their mean response and choosing the best ordered cut, an $O(K)$ rather than $O(2^K)$ search.
Multi-output trees?#
Yes. For multi-output regression, leaves store mean vectors and impurity is the trace of the within-leaf covariance. For multi-label classification, leaves store class-probability vectors and impurity is the average per-label Gini. sklearn supports both natively.
Pre-pruning or post-pruning?#
Pre-pruning is fast and good for prototyping. Post-pruning (cost-complexity) is more accurate because it sees the fully grown tree before deciding what to remove, but it is more expensive. In practice many people just tune max_depth and min_samples_leaf because trees are usually inside an ensemble that absorbs the rest of the variance.
Why are trees scale-invariant?#
Splits compare $x_j \leq \tau$ — only the ordering of values matters. Any monotone transform of a feature leaves the tree unchanged. This is also why one-hot encoding categorical variables works without further preprocessing.
How do I visualise a tree?#
| |
Can a tree do feature selection?#
Yes — sort by feature_importances_ and threshold. But the rankings are unstable on a single tree. Use a random forest or permutation importance for production feature selection.
Can tree training be parallelised?#
A single tree’s growth is sequential by node. Within a node, candidate splits across features are embarrassingly parallel; across trees in a forest, the trees themselves are independent. XGBoost and LightGBM additionally pipeline split-finding by histograms over feature buckets.
What is a sensible search range for max_depth?#
A practical default is $3 \leq d \leq 12$ for tabular data with $n$ in the thousands. Cross-validate. For ensembles, use shallower trees — depth 4–8 is typical for boosting.
Do trees scale to high-dimensional data?#
Naively, yes — but split-finding cost is $O(d \cdot n \log n)$
per node, and with sparse signals most features add noise. In high $d$
, prefer regularised linear models, or random forests with feature subsampling at each split (controlled by max_features).
Variants and Extensions#
- Cost-sensitive trees. Replace classification error by an asymmetric loss $L(y, \hat{y})$ — useful when false negatives cost more than false positives (medical diagnosis, fraud detection).
- Fuzzy / soft trees. Each split sends a fraction of a sample left and right, allowing differentiable training with gradient descent.
- Incremental trees. Hoeffding trees (Domingos & Hulten, 2000) use the Hoeffding bound to decide when enough samples have arrived to split with high confidence, enabling streaming training.
- Oblique trees. Linear-combination splits as discussed above (CART-LC, OC1).
Exercises#
Exercise 1. Information gain by hand. A 14-sample dataset has 9 positives and 5 negatives. Feature $X$ has two values: $X = a$ holds for 8 samples (6+, 2−), $X = b$ holds for 6 samples (3+, 3−). Compute $H(Y)$ , $H(Y \mid X)$ , $IG$ , $IV(X)$ , and $GR$ .
Solution. $H(Y) = -\tfrac{9}{14}\log_2 \tfrac{9}{14} - \tfrac{5}{14}\log_2\tfrac{5}{14} \approx 0.940$ . Conditional entropies: $H(Y \mid X=a) = -\tfrac{6}{8}\log_2\tfrac{6}{8} - \tfrac{2}{8}\log_2\tfrac{2}{8} \approx 0.811$ , $H(Y \mid X=b) = 1.000$ . Then $H(Y \mid X) = \tfrac{8}{14}(0.811) + \tfrac{6}{14}(1.000) \approx 0.892$ . So $IG \approx 0.048$ . $IV(X) = -\tfrac{8}{14}\log_2\tfrac{8}{14} - \tfrac{6}{14}\log_2\tfrac{6}{14} \approx 0.985$ , hence $GR \approx 0.049$ .
Exercise 2. Gini ≈ Entropy / 2. Show that for binary classification with natural log, $H_\mathrm{nat}(p) \approx 2\,G(p) \cdot \ln 2$ to second order around $p = \tfrac{1}{2}$ .
Solution. Expand $H_\mathrm{nat}(p) = -p \ln p - (1-p)\ln(1-p)$ around $p_0 = \tfrac{1}{2}$ . The first derivative vanishes at $p_0$ (maximum), and the second derivative is $-1/p_0 - 1/(1-p_0) = -4$ . Hence $H_\mathrm{nat}(p) \approx \ln 2 - 2(p - \tfrac{1}{2})^2$ . Similarly $G(p) = 2p(1-p) = \tfrac{1}{2} - 2(p - \tfrac{1}{2})^2$ . Both share the same quadratic shape; the constants differ by exactly the factor $\ln 2$ that converts nats to bits.
Exercise 3. The critical alpha. A subtree $T_t$ has 4 leaves and training error $0.10$ ; collapsing it to a leaf gives error $0.25$ . Compute $\alpha_\text{eff}(t)$ and decide whether to prune at $\alpha = 0.06$ .
$$\alpha_\text{eff} = \frac{0.25 - 0.10}{4 - 1} = 0.05.$$At $\alpha = 0.06 > 0.05$ , the cost of keeping the subtree exceeds the benefit, so we prune: $R_\alpha(\text{leaf}) = 0.25 + 0.06 = 0.31$ vs $R_\alpha(T_t) = 0.10 + 4(0.06) = 0.34$ .
Exercise 4. Missing-value penalty. A dataset has 100 samples; feature $X$ is missing in 20 of them. On the 80 observed samples $H(Y) = 0.94$ and $H(Y \mid X) = 0.60$ . Compute the adjusted information gain.
Solution. The observed-fraction weight is $80/100 = 0.8$ , so $IG_\text{adj} = 0.8 \cdot (0.94 - 0.60) = 0.272$ bits. A feature with twice the missingness would receive half the credit at the same observed gain.
Exercise 5. Steps to approximate a diagonal. On $[0, 1]^2$ , how many axis-aligned splits does a binary tree need to approximate the line $y = x$ to within $\epsilon$ in $L^\infty$ ?
Solution. A staircase of $n$ rectangles each of width $1/n$ has step height $1/n$ , giving error $\Theta(1/n)$ . Setting $1/n \leq \epsilon$ yields $n = \lceil 1/\epsilon \rceil$ rectangles, so $\Theta(1/\epsilon)$ leaves. An oblique tree with the single split $y - x \leq 0$ achieves zero error with one node — illustrating the cost of axis alignment for diagonal structure.
What’s next#
Decision trees take “axis-aligned splits + information gain” to its limit on tabular data, but they have two hard weaknesses: a single tree overfits (high variance), and the boundary is forever staircase-shaped — no depth lets it draw a diagonal. Two ways forward: switch to a different inductive bias, or ensemble the tree itself. The next chapter takes the first road — support vector machines.
SVM’s core idea is almost the opposite of a tree: it does not slice the space, it finds a max-margin hyperplane; it is not greedy, it solves a quadratic program; it does not use depth for expressivity, it uses the kernel trick to lift data into a higher-dimensional space where the originally non-separable problem becomes linearly separable. I run hard margin → soft margin → dual → kernels → SMO end to end — it is the first time after chapter four that KKT conditions actually do load-bearing work. By the end you should have a concrete sense of when SVM beats trees, and when it doesn’t.
References#
- Quinlan, J. R. (1986). Induction of decision trees. Machine Learning, 1(1), 81–106.
- Quinlan, J. R. (1993). C4.5: Programs for Machine Learning. Morgan Kaufmann.
- Breiman, L., Friedman, J., Stone, C. J., & Olshen, R. A. (1984). Classification and Regression Trees. Wadsworth.
- Hastie, T., Tibshirani, R., & Friedman, J. (2009). The Elements of Statistical Learning (2nd ed.). Springer. Chapter 9.
- Mitchell, T. M. (1997). Machine Learning. McGraw-Hill. Chapter 3.
- Breiman, L. (2001). Random forests. Machine Learning, 45(1), 5–32.
- Loh, W. Y. (2011). Classification and regression trees. WIREs Data Mining and Knowledge Discovery, 1(1), 14–23.
- Domingos, P., & Hulten, G. (2000). Mining high-speed data streams. KDD.
Decision trees are the building blocks of the most successful tabular learners we have. Understanding entropy, the Gini index, and cost-complexity pruning is a prerequisite for understanding random forests, GBDT, XGBoost, and LightGBM — the next chapters build directly on the machinery developed here.
ML Math Derivations 20 parts
- 01 ML Math Derivations (1): Introduction and Mathematical Foundations
- 02 ML Math Derivations (2): Linear Algebra and Matrix Theory
- 03 ML Math Derivations (3): Probability Theory and Statistical Inference
- 04 ML Math Derivations (4): Convex Optimization Theory
- 05 ML Math Derivations (5): Linear Regression
- 06 ML Math Derivations (6): Logistic Regression and Classification
- 07 ML Math Derivations (7): Decision Trees you are here
- 08 ML Math Derivations (8): Support Vector Machines
- 09 ML Math Derivations (9): Naive Bayes
- 10 ML Math Derivations (10): Semi-Naive Bayes and Bayesian Networks
- 11 ML Math Derivations (11): Ensemble Learning
- 12 ML Math Derivations (12): XGBoost and LightGBM
- 13 ML Math Derivations (13): EM Algorithm and GMM
- 14 ML Math Derivations (14): Variational Inference and Variational EM
- 15 ML Math Derivations (15): Hidden Markov Models
- 16 ML Math Derivations (16): Conditional Random Fields
- 17 ML Math Derivations (17): Dimensionality Reduction and PCA
- 18 ML Math Derivations (18): Clustering Algorithms
- 19 ML Math Derivations (19): Neural Networks and Backpropagation
- 20 ML Math Derivations (20): Regularization and Model Selection