
ML Math Derivations (9): Naive Bayes
Rigorous derivation of Naive Bayes from Bayes theorem through conditional independence, parameter estimation, Laplace smoothing, three model variants, and why it works despite violated assumptions.
Hook: A spam filter that trains in milliseconds, scales to a million features, has no hyperparameters worth tuning, and still beats much fancier models on short-text problems. Naive Bayes pulls this off by making one outrageous assumption — every feature is independent given the class — and refusing to apologise for it. The assumption is wrong on essentially every real dataset, yet the classifier works. Understanding why is a tour through generative modelling, MAP estimation, Dirichlet priors, and the bias–variance tradeoff. This article walks the entire path.

What You Will Learn#
- Bayes’ theorem as the foundation of probabilistic classification, and why the Bayes-optimal rule is the target every classifier secretly imitates.
- The conditional-independence assumption — what it costs us geometrically, and what it buys us statistically.
- Maximum-likelihood estimates for priors and likelihoods, and the MAP / Dirichlet view of Laplace smoothing.
- The three workhorse variants — Multinomial, Bernoulli, Gaussian — and a decision rule for picking between them.
- Why Naive Bayes survives violated assumptions: the Domingos–Pazzani result, error cancellation, and the bias–variance tradeoff.
- A clean, NumPy-only implementation of all three variants, validated against scikit-learn.
Prerequisites#
- Probability: Bayes’ theorem, conditional probability, joint vs. marginal distributions, Gaussian density.
- Calculus: logarithms, partial derivatives.
- Familiarity with Part 8: Support Vector Machines , especially the discriminative perspective on classification.
Bayesian Decision Theory#
Before we add the “naive” piece, we need to be honest about what an optimal probabilistic classifier looks like. Naive Bayes is one specific way of approximating it.
Bayes’ Theorem, Reread as a Belief Update#
$$P(c_k \mid \mathbf{x}) \;=\; \frac{P(\mathbf{x} \mid c_k)\, P(c_k)}{P(\mathbf{x})}.$$Each term has a job:
| Term | Name | What it answers |
|---|---|---|
| $P(c_k \mid \mathbf{x})$ | Posterior | After seeing the data, how likely is class $c_k$ ? |
| $P(\mathbf{x} \mid c_k)$ | Likelihood | If the class were $c_k$ , how plausible is this $\mathbf{x}$ ? |
| $P(c_k)$ | Prior | Before any features, how common is $c_k$ ? |
| $P(\mathbf{x})$ | Evidence | How likely is $\mathbf{x}$ overall? Pure normaliser. |
Why the prior matters. Suppose a test for a rare disease is 99% accurate. With prevalence $0.1\%$ , a positive test moves the posterior from $0.001$ to roughly $0.09$ — still ten-to-one against the disease. Bayes’ theorem makes the prior do its arithmetic out loud; ignoring priors is the canonical statistics mistake.
The Bayes-Optimal Classifier#
$$ \hat{y}(\mathbf{x}) \;=\; \arg\max_{c_k} P(c_k \mid \mathbf{x}) \;=\; \arg\max_{c_k} P(\mathbf{x} \mid c_k)\, P(c_k), $$ $$R^\star(\mathbf{x}) \;=\; 1 - \max_k P(c_k \mid \mathbf{x}),$$is the Bayes risk, and the data-set-level $\mathbb{E}_{\mathbf{x}}[R^\star(\mathbf{x})]$ is the Bayes error rate — the irreducible noise floor of the problem itself.
Every probabilistic classifier we will study is, in one way or another, an approximation of this rule under finite data.
Generative vs. Discriminative#
There are two ways to approach $P(c_k \mid \mathbf{x})$ :
- Discriminative models (logistic regression, SVM, neural nets) learn $P(y \mid \mathbf{x})$ directly. They put all their statistical power into the decision surface and ignore how features are distributed.
- Generative models (Naive Bayes, LDA, HMMs) model $P(\mathbf{x}, y) = P(\mathbf{x} \mid y)P(y)$ and derive the posterior via Bayes’ theorem.
| Discriminative | Generative | |
|---|---|---|
| Learns | $P(y \mid \mathbf{x})$ | $P(\mathbf{x}, y)$ |
| Asymptotic accuracy | Often higher | Often lower |
| Sample efficiency | Worse | Better when assumptions hold |
| Missing features | Hard | Natural — marginalise them out |
| Generates new $\mathbf{x}$ | No | Yes |
| Uses unlabeled data | No | Easy |
A classic result of Ng & Jordan (2001) shows the trade-off cleanly: with infinite data the discriminative model wins, but the generative model converges faster. Naive Bayes is the generative side at its most extreme.
The Naive Bayes Classifier#
Why We Need a Drastic Simplification#
Estimating $P(\mathbf{x} \mid c_k)$ for $\mathbf{x} \in \{0,1\}^d$ requires $2^d - 1$ free parameters per class. For a vocabulary of $d = 10{,}000$ words this is utterly hopeless. The joint distribution is the bottleneck.
The Conditional-Independence Assumption#
$$P(\mathbf{x} \mid c_k) \;=\; \prod_{j=1}^{d} P\!\left(x^{(j)} \mid c_k\right).$$Two readings of this:
- Causal. Treat the class as a hidden cause that produces each feature independently. Once you know the patient has the flu, fever and cough are no longer informative about each other.
- Statistical. Each feature contributes its own one-dimensional conditional density. The joint is the product. Parameter count drops from exponential in $d$ to linear in $d$ .
The geometric cost of this assumption is illustrated below: the true joint density (left) has correlated, tilted contours; the Naive Bayes approximation (right) has the same marginals but axis-aligned, factorised contours.
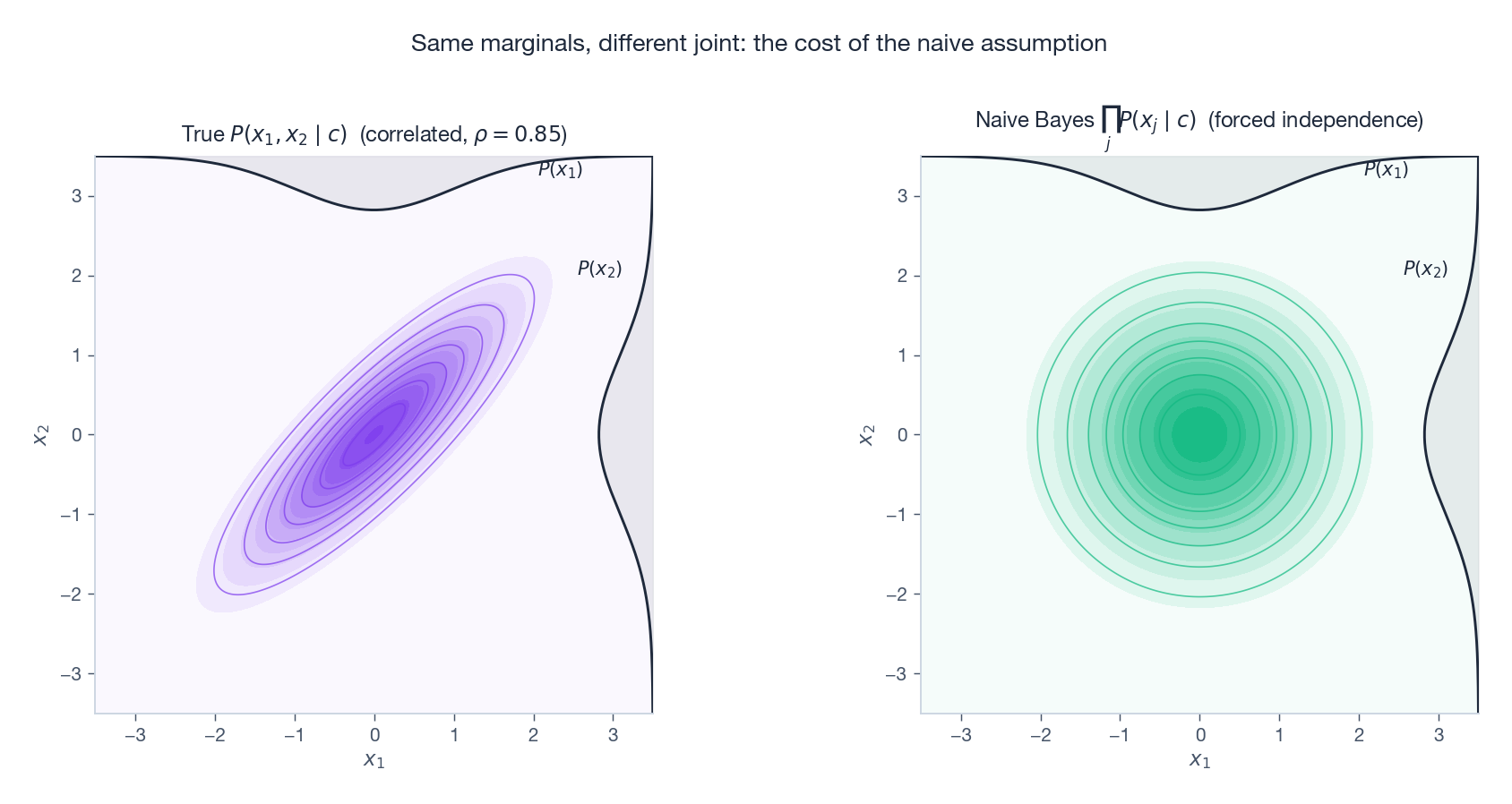
The two densities can disagree dramatically, yet — as we will see in §5 — the classification often does not.
The Classification Rule#
$$\hat{y}(\mathbf{x}) \;=\; \arg\max_{c_k}\; P(c_k) \prod_{j=1}^{d} P\!\left(x^{(j)} \mid c_k\right).$$ $$\boxed{\;\hat{y}(\mathbf{x}) \;=\; \arg\max_{c_k} \left[\, \ln P(c_k) \;+\; \sum_{j=1}^{d} \ln P\!\left(x^{(j)} \mid c_k\right) \right].\;}$$This is the entire algorithm. Everything else — three “variants”, smoothing, calibration — is just a discussion of how to estimate the per-feature conditionals $P(x^{(j)} \mid c_k)$ .
For a 2D toy problem with two Gaussian classes, the picture is concrete:
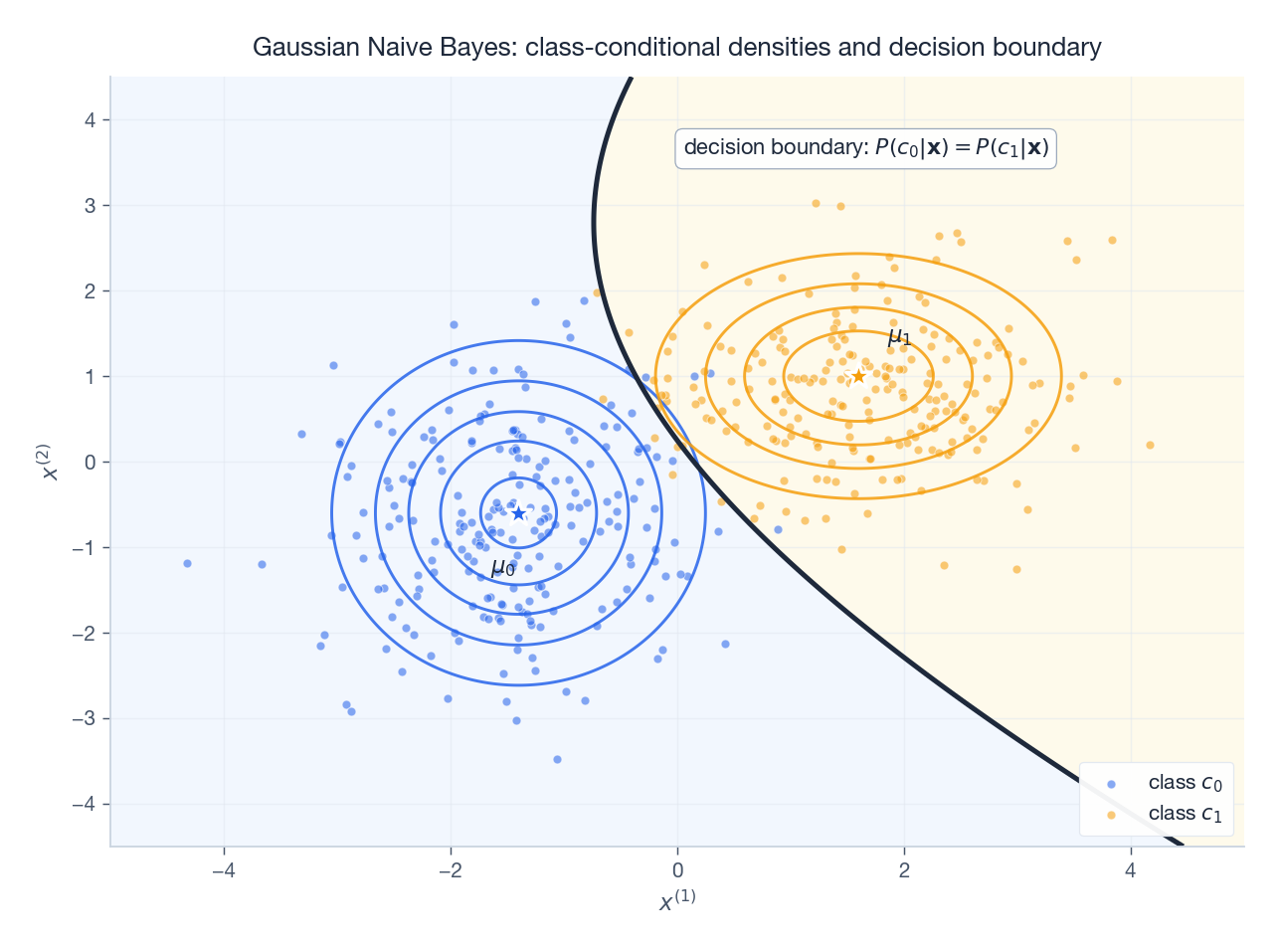
Each class has a diagonal-covariance Gaussian (axis-aligned ellipses, the Naive Bayes hypothesis class for continuous features). The decision boundary is the locus of points where the two posteriors agree — quadratic in general, linear when the classes share a covariance.
The same picture viewed probabilistically (rather than geometrically) is the posterior surface:
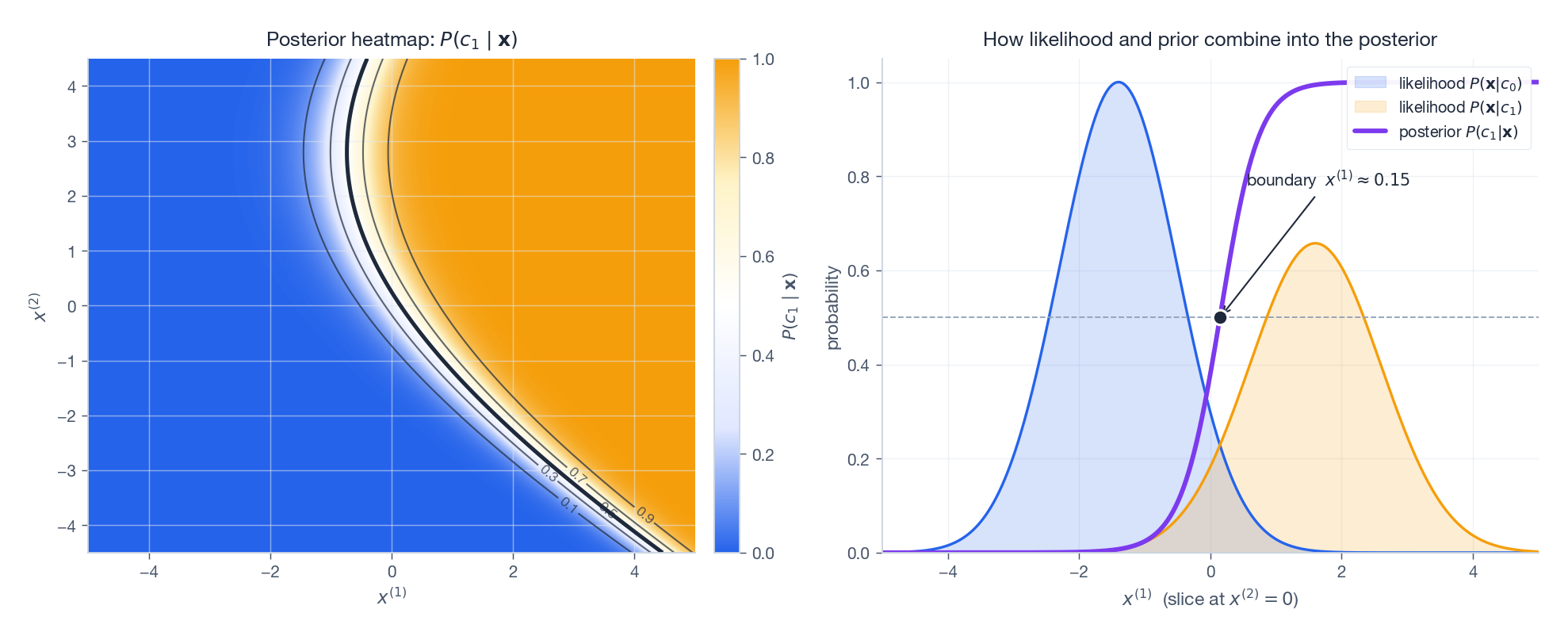
The right panel makes the soft transition explicit: along a slice through the data, the posterior $P(c_1 \mid \mathbf{x})$ rises smoothly from $0$ to $1$ , crossing $0.5$ exactly where the two scaled likelihoods cross. That crossover is the decision boundary.
Parameter Estimation by Maximum Likelihood#
Given training data $\mathcal{D} = \{(\mathbf{x}_i, y_i)\}_{i=1}^{N}$ with $N_k$ samples in class $c_k$ :
$$\hat{P}(c_k) \;=\; \frac{N_k}{N}.$$ $$ \hat{P}\!\left(x^{(j)} = v \mid c_k\right) \;=\; \frac{\#\{i : x_i^{(j)} = v \text{ and } y_i = c_k\}}{N_k}. $$ $$P\!\left(x^{(j)} \mid c_k\right) \;=\; \frac{1}{\sqrt{2\pi}\,\sigma_{jk}} \exp\!\left(-\frac{(x^{(j)} - \mu_{jk})^2}{2\sigma_{jk}^2}\right),$$ $$ \hat{\mu}_{jk} \;=\; \frac{1}{N_k}\!\sum_{i:\, y_i=c_k}\! x_i^{(j)}, \qquad \hat{\sigma}_{jk}^2 \;=\; \frac{1}{N_k}\!\sum_{i:\, y_i=c_k}\! \bigl(x_i^{(j)} - \hat{\mu}_{jk}\bigr)^2. $$That is all the training. There is no iterative optimiser, no learning rate, no convergence check.
Laplace Smoothing — and Its Bayesian Soul#
The catastrophe of zero counts. Suppose the word excellent never appears in your spam training set. Then $\hat{P}(\text{excellent} \mid \text{spam}) = 0$ , and a single occurrence of excellent in a future email forces $P(\text{spam} \mid \mathbf{x}) = 0$ regardless of how spammy the other 199 words are. One missing word kills everything.
$$ \hat{P}\!\left(x^{(j)} = v \mid c_k\right) \;=\; \frac{\#\{i : x_i^{(j)} = v,\, y_i=c_k\} + \alpha}{N_k + \alpha\, S_j}, \qquad \hat{P}(c_k) \;=\; \frac{N_k + \alpha}{N + \alpha K}. $$With $\alpha = 1$ this is Laplace (add-one) smoothing; smaller $\alpha$ is sometimes called Lidstone smoothing.
Why it works. Place a symmetric Dirichlet prior $\mathrm{Dir}(\alpha, \dots, \alpha)$ on the multinomial $P(\cdot \mid c_k)$ . The Dirichlet is conjugate to the multinomial, so the posterior is also Dirichlet, with parameters $\alpha + \text{counts}$ . The MAP estimate of that posterior is exactly the smoothed formula above. Laplace smoothing is MAP estimation under a uniform Dirichlet prior — every category has been “seen” $\alpha$ times before any data arrives.
The visual story:
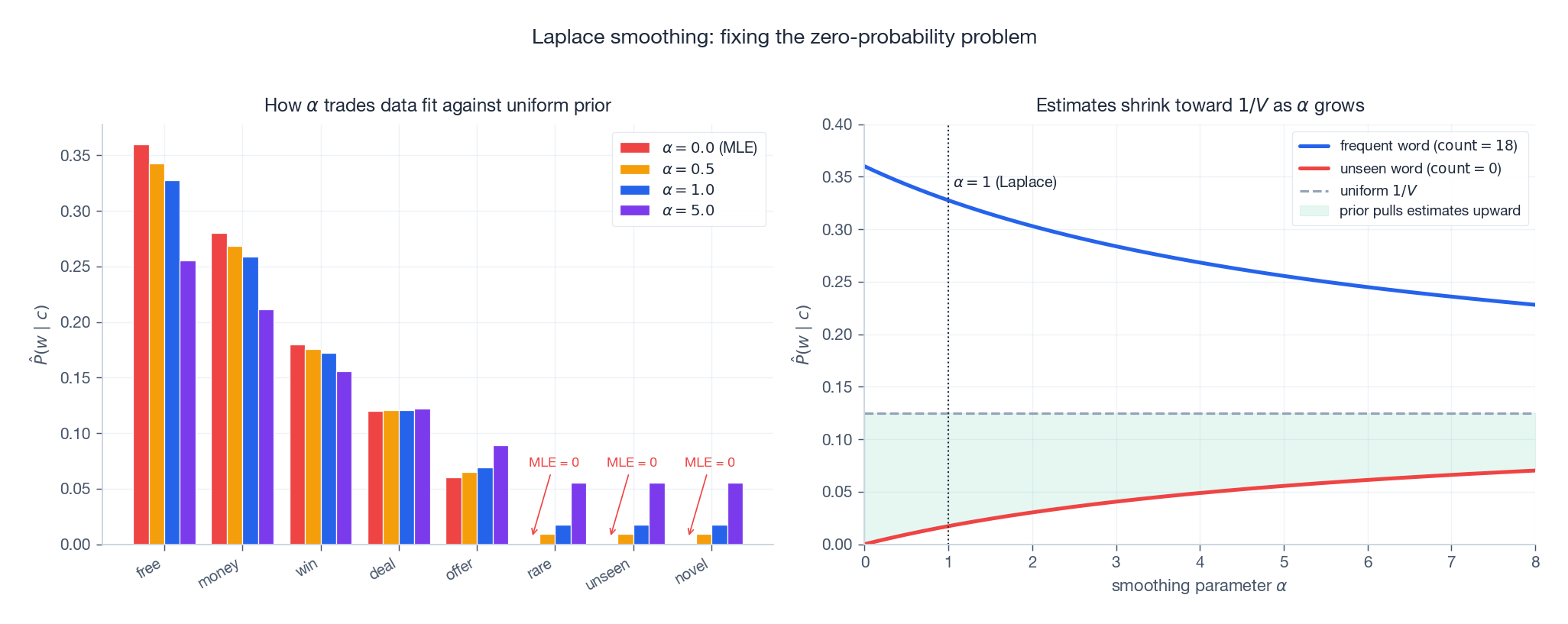
- Left: as $\alpha$ grows, common words give up probability mass to rare and unseen words.
- Right: the estimate of every word is pulled toward the uniform $1/V$ . Frequent words shrink down, unseen words rise from $0$ . The pull strength is proportional to $\alpha / N_k$ .
Choosing $\alpha$ is a bias–variance dial: small $\alpha$ trusts the data (low bias, high variance); large $\alpha$ trusts the uniform prior (high bias, low variance). Cross-validate.
Three Variants — Same Bayes Rule, Different Likelihood#

The classification rule never changes. Only the model for $P(x^{(j)} \mid c_k)$ changes.
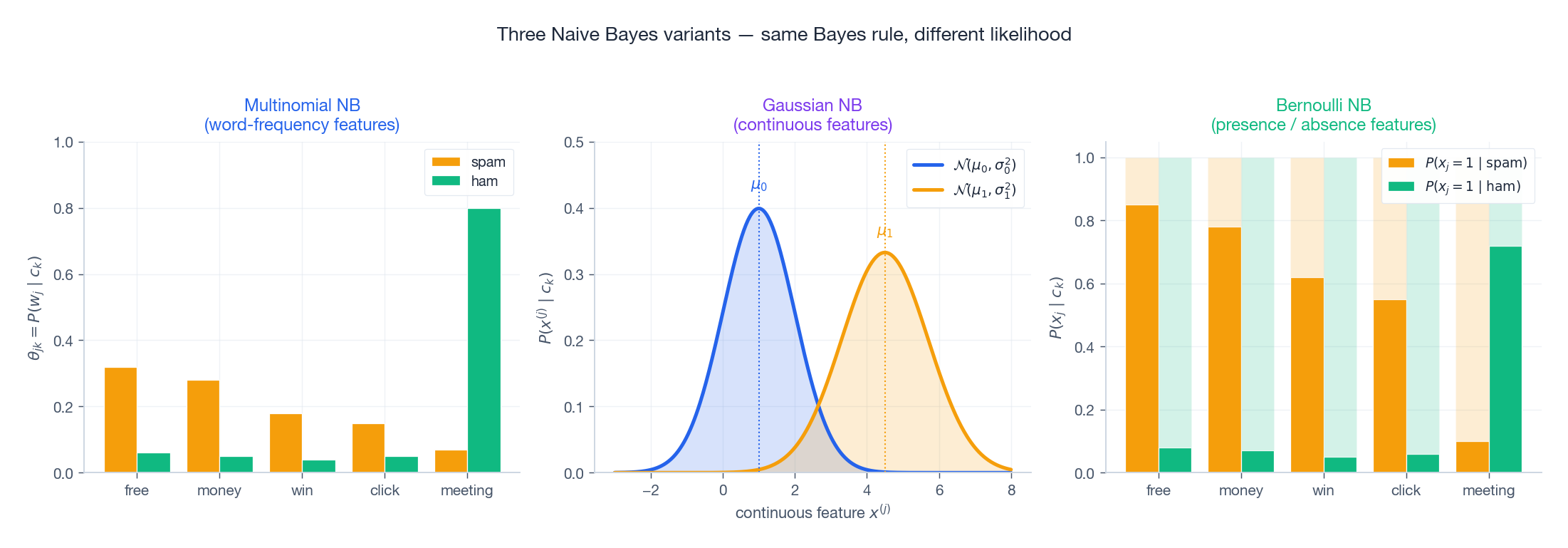
Multinomial Naive Bayes — for word counts#
Use when features are non-negative integer counts: term frequencies, $n$ -grams, hashed buckets.
$$P(\mathbf{x} \mid c_k) \;\propto\; \prod_{j=1}^{d} \theta_{jk}^{x^{(j)}}.$$ $$\hat{\theta}_{jk} \;=\; \frac{\sum_{i:\, y_i=c_k} x_i^{(j)} + \alpha}{\sum_{j'=1}^{d}\sum_{i:\, y_i=c_k} x_i^{(j')} + \alpha d}.$$In words: the fraction of all word tokens in class-$c_k$ documents that happen to be word $j$ .
Bernoulli Naive Bayes — for word presence#
Use when features are binary: word present (1) or absent (0). Especially good for short texts.
$$P(\mathbf{x} \mid c_k) \;=\; \prod_{j=1}^{d} p_{jk}^{x^{(j)}} (1 - p_{jk})^{1 - x^{(j)}}.$$The crucial difference from Multinomial. Bernoulli explicitly multiplies in $(1 - p_{jk})$ for every absent word. Absence is evidence. If “free” is missing from an email and “free” is normally present in 85% of spam, that absence is strong evidence against spam. Multinomial NB simply ignores absent words.
Rule of thumb. Short, sparse documents (tweets, log lines, short queries): Bernoulli usually wins. Long documents (news, reviews): Multinomial is typically better because frequency carries real information.
Gaussian Naive Bayes — for continuous features#
Use when features are real-valued and roughly unimodal per class: physical measurements, embeddings, sensor readings.
$$P(x^{(j)} \mid c_k) \;=\; \mathcal{N}\!\left(x^{(j)};\, \mu_{jk},\, \sigma_{jk}^2\right).$$The total per-class density is the product of $d$ univariate Gaussians — i.e. a multivariate Gaussian with diagonal covariance. This is exactly the picture in Figure 1: axis-aligned ellipses.
A surprising fact: when the two classes have the same per-feature variances ($\sigma_{jk}^2 = \sigma_j^2$ for all $k$ ), the log-posterior ratio becomes linear in $\mathbf{x}$ , and Gaussian NB reproduces the same boundary as logistic regression — but estimated generatively rather than discriminatively.
Worked Example: Spam Classification#
Bayes’ theorem, conditional independence, and Laplace smoothing all collide in the canonical Naive Bayes use case: spam filtering. The pipeline is:
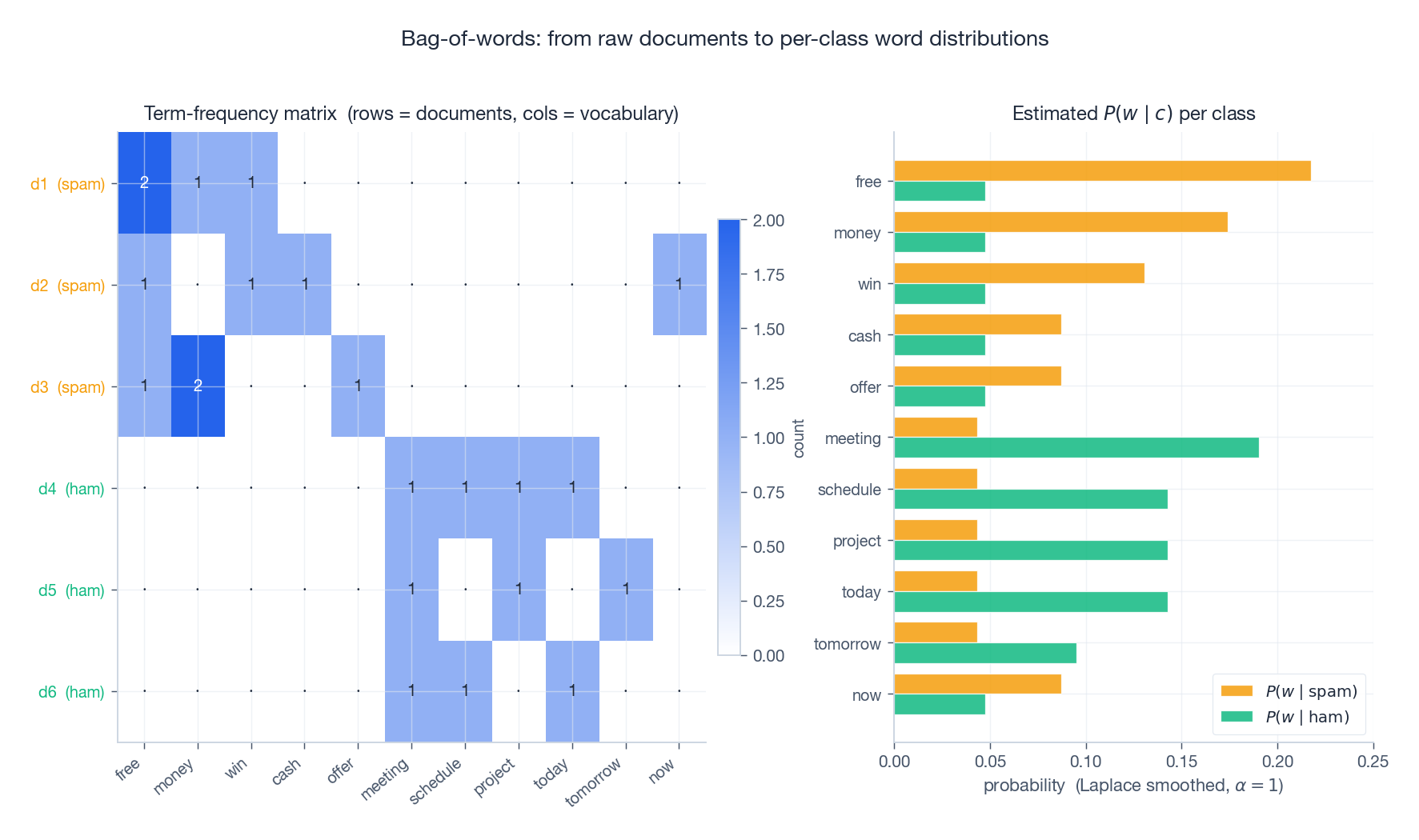
- Tokenise documents into words.
- Build a term-frequency matrix of shape $(N \times d)$ — rows are documents, columns are vocabulary entries.
- Estimate $\hat{P}(c_k)$ from class frequencies and $\hat{\theta}_{jk}$ (Multinomial) or $\hat{p}_{jk}$ (Bernoulli) with Laplace smoothing.
- Predict by computing the per-class log score and taking the argmax.
The right panel shows the estimated per-class word distributions: spam concentrates probability on “free”, “money”, “win”; ham on “meeting”, “schedule”, “project”. A test email’s classification is just an additive accumulation of per-feature evidence:
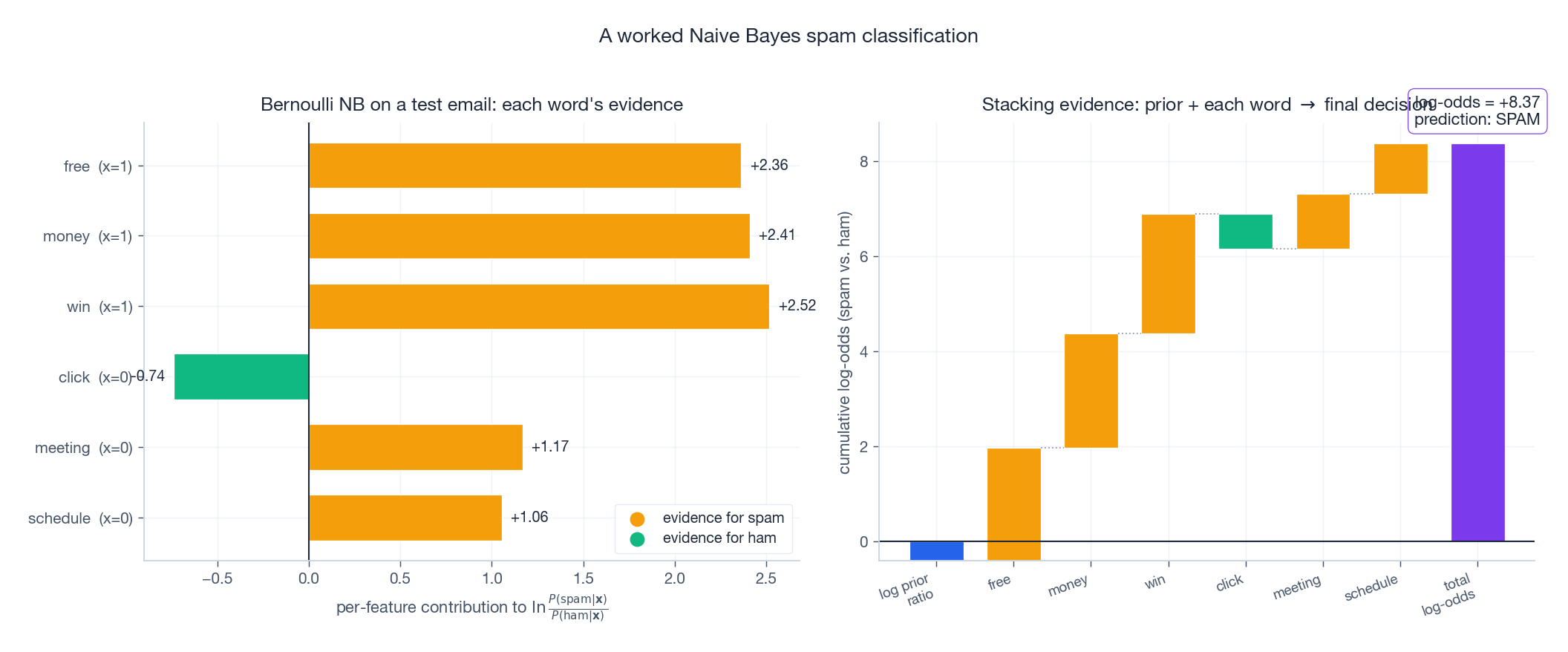
Read the waterfall left to right: start from the log prior ratio $\ln \frac{P(\mathrm{spam})}{P(\mathrm{ham})}$ , then add the contribution of each word (positive bars push toward spam, negative toward ham). The final height is the total log-odds. If it is positive, predict spam.
This is, in a strong sense, the whole algorithm: every Naive Bayes prediction is a sum of per-feature log-likelihood ratios.
Why Does It Work When the Assumption Is Wrong?#
The independence assumption is essentially never true. Words co-occur, symptoms cluster, sensor readings correlate. So why does Naive Bayes routinely win on text classification benchmarks against models that know about correlations?
Four reinforcing reasons:
- Classification needs ranking, not calibration. The argmax depends only on which posterior is largest, not on its absolute value. A model can be wildly miscalibrated yet rank classes correctly.
- Error cancellation in ratios. When we compute the log-posterior ratio, repeated correlated features inflate both class scores by similar amounts. The bias largely cancels.
- Bias–variance tradeoff. The independence assumption injects bias but slashes variance: instead of $\mathcal{O}(K \cdot d^2)$ correlation parameters, we estimate $\mathcal{O}(K \cdot d)$ marginals. With limited data, lower variance often wins outright.
- The Domingos–Pazzani theorem (1997). They showed Naive Bayes is 0-1-loss optimal whenever the class with the highest true posterior also has the highest estimated posterior — which is a much weaker condition than the independence assumption itself. Naive Bayes can be globally wrong about probabilities and still locally right about every prediction.
This is not a license to skip diagnostics. Naive Bayes is not well calibrated and its probability outputs should not be trusted at face value (see Q&A). But for ranking and argmax decisions, it punches well above its weight.
Reference Implementation#
A clean NumPy implementation of all three variants. The Gaussian version is validated against scikit-learn at the end.
| |
Exercises#
Exercise 1 — Conditional independence, by definition#
Problem. Given $P(c_1) = 0.6$ , $P(x_1=1 \mid c_1) = 0.8$ , $P(x_2=1 \mid c_1) = 0.5$ , compute $P(x_1=1, x_2=1 \mid c_1)$ under the Naive Bayes assumption.
$$P(x_1=1, x_2=1 \mid c_1) = P(x_1=1 \mid c_1)\, P(x_2=1 \mid c_1) = 0.8 \times 0.5 = 0.4.$$The prior $P(c_1)$ is irrelevant inside the conditional — it would only enter if we also wanted $P(x_1, x_2)$ unconditionally.
Exercise 2 — Laplace smoothing in numbers#
Problem. Vocabulary size $V = 1000$ . In the positive class, the word excellent appears 5 times in a corpus of 200 word tokens. Compute (a) the MLE estimate, (b) the Laplace-smoothed estimate ($\alpha = 1$ ), (c) the smoothed estimate for an unseen word.
Solution.
(a) MLE: $\hat{P}(\text{excellent} \mid +) = 5 / 200 = 0.025$ .
(b) Smoothed: $\hat{P}(\text{excellent} \mid +) = (5+1) / (200 + 1000) = 6 / 1200 = 0.005$ .
(c) Unseen word: $\hat{P}(w \mid +) = (0+1) / (200 + 1000) = 1/1200 \approx 8.3 \times 10^{-4}$ .
Notice how aggressively smoothing flattens the distribution when $\alpha V \gg N_k$ (here $1000 \gg 200$ ). This is the dial: with little class-conditional data, the prior dominates.
Exercise 3 — Multinomial vs. Bernoulli on the same email#
Problem. A 100-word email contains win 5 times and free 3 times. Under spam: $P(\text{win} \mid \text{spam}) = 0.05$ , $P(\text{free} \mid \text{spam}) = 0.03$ (Multinomial); $P(\text{win present} \mid \text{spam}) = 0.7$ , $P(\text{free present} \mid \text{spam}) = 0.6$ (Bernoulli). Compare log-likelihood contributions.
Solution.
$$\ln L_{\mathrm{mult}} = 5\ln(0.05) + 3\ln(0.03) = 5(-2.996) + 3(-3.507) = -25.5.$$ $$\ln L_{\mathrm{bern}} = \ln(0.7) + \ln(0.6) = -0.357 + (-0.511) = -0.87.$$These numbers are not directly comparable (different probabilistic objects), but the shape tells the story: Multinomial amplifies repeated occurrences linearly with count, while Bernoulli flattens 5 occurrences and 1 occurrence into the same evidence. For long documents that dynamic range matters; for tweets it usually does not.
Exercise 4 — Gaussian NB parameters by hand#
Problem. Class $+1$ contains three 2D points: $(1, 2), (2, 3), (3, 1)$ . Compute the Gaussian NB parameters.
$$ \hat\mu_1 = \tfrac{1+2+3}{3} = 2, \quad \hat\sigma_1^2 = \tfrac{(1{-}2)^2 + (2{-}2)^2 + (3{-}2)^2}{3} = \tfrac{2}{3}, $$ $$ \hat\mu_2 = \tfrac{2+3+1}{3} = 2, \quad \hat\sigma_2^2 = \tfrac{(2{-}2)^2 + (3{-}2)^2 + (1{-}2)^2}{3} = \tfrac{2}{3}. $$So $\hat\sigma_1 = \hat\sigma_2 = \sqrt{2/3} \approx 0.816$ . Gaussian NB has no off-diagonal covariance term; the (1,3) sample being on the “wrong” diagonal does not enter the model.
Exercise 5 — Why does Naive Bayes work?#
Problem. Give four independent reasons Naive Bayes performs well despite violated independence.
Solution. The reasons developed in §5:
- Argmax-only decisions. Classification needs the right ranking, not the right probability magnitudes.
- Bias cancellation in log-ratios. Correlated features inflate the log-likelihoods of both classes by similar amounts, and these cancel in the posterior ratio.
- Bias–variance favourability. Trading a small amount of bias for a large reduction in variance is often a net win, especially when $N$ is small relative to $d$ .
- Domingos–Pazzani (1997). A formal result: Naive Bayes is 0-1-loss optimal whenever its argmax matches the true posterior’s argmax — a much weaker condition than the joint distribution actually factorising.
FAQ#
Why “naive”?#
The conditional-independence assumption is rarely literally true — words co-occur, symptoms cluster — yet the model is built as if they did. The label refers to that strong, almost ostentatiously simple, assumption.
How is Naive Bayes related to logistic regression?#
Under Gaussian features with shared per-feature variance, Gaussian NB produces a linear decision boundary $\mathbf{w}^\top\mathbf{x} + b = 0$ — the same form as logistic regression. The coefficients differ because NB estimates $\mathbf{w}$ by generative MLE on $P(\mathbf{x}, y)$ , while LR estimates it discriminatively on $P(y \mid \mathbf{x})$ . Asymptotically LR is at least as good; NB converges with fewer samples.
Are Naive Bayes probability outputs trustworthy?#
No. Because correlated features get their log-likelihoods double-counted, the posteriors are typically too extreme (close to 0 or 1). For ranking tasks (argmax, top-$K$ , ROC) this is fine. For decisions that depend on the magnitude (cost-sensitive thresholds, risk computations) calibrate with Platt scaling or isotonic regression on a held-out set.
What is the time complexity?#
Training: $\mathcal{O}(Nd)$ — one pass through the training data. Prediction: $\mathcal{O}(Kd)$ per sample. There are no iterations and no hyperparameters whose tuning costs more than a CV sweep over $\alpha$ .
How does Naive Bayes handle missing features?#
Naturally: when feature $j$ is missing for sample $i$ , omit the term $\ln P(x_i^{(j)} \mid c_k)$ from the sum. This is mathematically equivalent to marginalising over the missing feature and is a real advantage of generative modelling.
When should I not use Naive Bayes?#
When (i) features are heavily correlated and the correlation differs across classes (so error cancellation fails), (ii) the application needs calibrated probabilities, or (iii) you have plenty of data and a discriminative model is feasible — logistic regression, gradient boosting, or transformers will usually pull ahead.
What’s next#
Naive Bayes’ “given the class, features are independent” assumption is almost cheating, yet often survives in practice. The trouble shows up when features really are correlated — symptoms in medical diagnosis, item co-occurrence in recommenders — at which point the assumption visibly cracks. The next chapter is the natural extension: relax the independence, but not all at once.
Semi-naive Bayes (TAN, AODE) lets each feature depend on at most one other feature — expressivity goes up while estimation stays tractable. One step further is the full Bayesian network: a directed acyclic graph that explicitly encodes the conditional-independence structure, factoring the joint into local conditionals. I climb the ladder NB → TAN → AODE → general Bayesian network and ask the same question at every rung: how much extra expressivity did I buy, and how much inference / learning cost did I pay?
References#
- Domingos, P., & Pazzani, M. (1997). On the optimality of the simple Bayesian classifier under zero-one loss. Machine Learning, 29(2-3), 103-130.
- Ng, A. Y., & Jordan, M. I. (2001). On discriminative vs. generative classifiers: A comparison of logistic regression and naive Bayes. NeurIPS.
- McCallum, A., & Nigam, K. (1998). A comparison of event models for naive Bayes text classification. AAAI Workshop on Learning for Text Categorization.
- Manning, C. D., Raghavan, P., & Schutze, H. (2008). Introduction to Information Retrieval. Cambridge University Press. Chapter 13 .
- Bishop, C. M. (2006). Pattern Recognition and Machine Learning. Springer. Section 8 .2.
- Murphy, K. P. (2012). Machine Learning: A Probabilistic Perspective. MIT Press. Chapter 3 .
ML Mathematical Derivations Series
< Part 8: Support Vector Machines | Part 9: Naive Bayes | Part 10: Semi-Naive Bayes >
ML Math Derivations 20 parts
- 01 ML Math Derivations (1): Introduction and Mathematical Foundations
- 02 ML Math Derivations (2): Linear Algebra and Matrix Theory
- 03 ML Math Derivations (3): Probability Theory and Statistical Inference
- 04 ML Math Derivations (4): Convex Optimization Theory
- 05 ML Math Derivations (5): Linear Regression
- 06 ML Math Derivations (6): Logistic Regression and Classification
- 07 ML Math Derivations (7): Decision Trees
- 08 ML Math Derivations (8): Support Vector Machines
- 09 ML Math Derivations (9): Naive Bayes you are here
- 10 ML Math Derivations (10): Semi-Naive Bayes and Bayesian Networks
- 11 ML Math Derivations (11): Ensemble Learning
- 12 ML Math Derivations (12): XGBoost and LightGBM
- 13 ML Math Derivations (13): EM Algorithm and GMM
- 14 ML Math Derivations (14): Variational Inference and Variational EM
- 15 ML Math Derivations (15): Hidden Markov Models
- 16 ML Math Derivations (16): Conditional Random Fields
- 17 ML Math Derivations (17): Dimensionality Reduction and PCA
- 18 ML Math Derivations (18): Clustering Algorithms
- 19 ML Math Derivations (19): Neural Networks and Backpropagation
- 20 ML Math Derivations (20): Regularization and Model Selection