ML Math Derivations (10): Semi-Naive Bayes and Bayesian Networks
From SPODE, TAN and AODE to full Bayesian networks: how relaxing the conditional-independence assumption -- through one-dependence trees, ensembles of super-parents and graphical structure learning -- closes the gap between Naive Bayes and the full joint distribution.
Hook. Naive Bayes assumes every feature is conditionally independent given the class. It is a convenient lie — one that lets us train in a single pass over the data, but one that classifiers based on tree structures and small graphs can systematically beat by a few accuracy points on virtually every UCI benchmark. This part walks the spectrum from “no dependencies” (Naive Bayes) to “all dependencies” (full joint), showing the three sweet spots that practitioners actually use: SPODE, TAN and AODE. The same factorisation idea, taken to its general form, is the Bayesian network.
What You Will Learn#
- Why the full conditional-independence assumption fails, and how the parameter count grows when we relax it.
- SPODE: every feature gets a single shared “super-parent”.
- TAN: each feature gets its own parent, chosen by a maximum-spanning-tree on conditional mutual information (Chow-Liu).
- AODE: averaging over all eligible super-parents to remove model-selection variance.
- Bayesian networks: DAG factorisation, d-separation, the Markov blanket, variable elimination and the junction tree.
- A practical decision rule: NB vs TAN vs AODE vs full BN.
Prerequisites#
- Joint distributions, conditional independence, mutual information.
- Spanning trees, dynamic programming on trees.
- Familiarity with Part 9: Naive Bayes .
Why relax the independence assumption?#
The convenient lie#
$$P(\mathbf{x}\mid c_k) \;=\; \prod_{j=1}^{d} P(x^{(j)}\mid c_k).$$The lie is that, conditioned on the class, features are unrelated. Three places it almost always breaks:
- Text. The bigram “machine learning” is far more frequent than $P(\text{machine})\,P(\text{learning})$ would suggest, even within the “ML article” class.
- Medicine. Within “flu”, fever and cough remain strongly correlated — they share a downstream physiological pathway.
- Vision. Adjacent pixels are nearly identical regardless of the class label.
When the conditional-independence assumption is severely violated, the posterior $P(c\mid\mathbf{x})$ is miscalibrated: the “winning” class still wins, but with wildly inflated confidence, and the ranking among close runners-up degrades.
The spectrum#
| Model | Allowed dependencies | # parameters | Accuracy |
|---|---|---|---|
| Naive Bayes | none | $O(K d)$ | baseline |
| 1-dependence (SPODE / TAN / AODE) | each feature has 1 parent | $O(K d S)$ | better |
| $k$ -dependence | each feature has $k$ parents | $O(K d S^{k})$ | even better |
| Full joint | everything | $O(K S^{d})$ | best, but intractable |
Here $S$ is the average number of values per feature and $K$ the number of classes. The empirical sweet spot is $k=1$ : the cost is one extra factor of $S$ , and the accuracy gain over Naive Bayes is consistently the largest single jump.
One-dependence estimators (ODE)#
SPODE: a single super-parent#
$$P(\mathbf{x}\mid c_k) \;=\; P(x^{(p)}\mid c_k)\prod_{j\neq p} P(x^{(j)}\mid c_k,\,x^{(p)}).$$Reading the model. Instead of “all features are independent given the class”, SPODE says “all features share one hidden moderator beyond the class”. For spam detection that moderator might be the token free: knowing whether free appears changes the probabilities of click, offer, winner.
$$ \hat{P}(x^{(j)}{=}v\mid c_k,\,x^{(p)}{=}u) \;=\; \frac{\#\{x^{(j)}{=}v,\,c_k,\,x^{(p)}{=}u\}+\alpha} {\#\{c_k,\,x^{(p)}{=}u\}+\alpha\,S_j}. $$ $$ p^{*} \;=\; \arg\max_{p}\, I(X^{(p)};Y), \qquad p^{*} \;=\; \arg\max_{p}\,\sum_{j\neq p} I(X^{(j)};Y\mid X^{(p)}). $$The first picks the feature most informative about $Y$ ; the second picks the feature whose conditioning maximally explains the rest. Both are heuristics — and the next two models exist precisely because this single choice is fragile.
AODE: average all eligible super-parents#
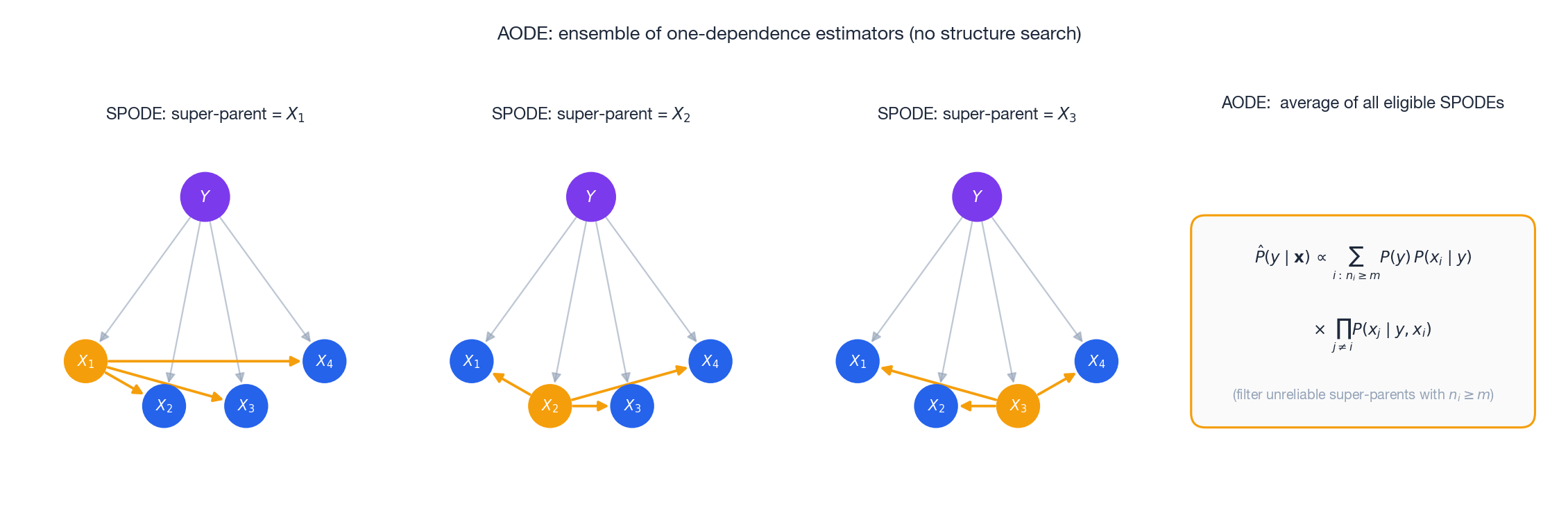
Here $n_i$ is the number of training samples with attribute value $x^{(i)}$ and $m$ (typically 30) is the minimum support for a super-parent to be trusted.
Why it works.
- No structure search. The model is fixed; only counts are estimated.
- Variance reduction. Averaging $d$ correlated SPODE estimates is a variance-shrinker, just like bagging.
- Trivial to ship. Same counters as SPODE, summed over all $i$ .
| |
TAN: tree-augmented Naive Bayes#

Each feature picks its own parent#
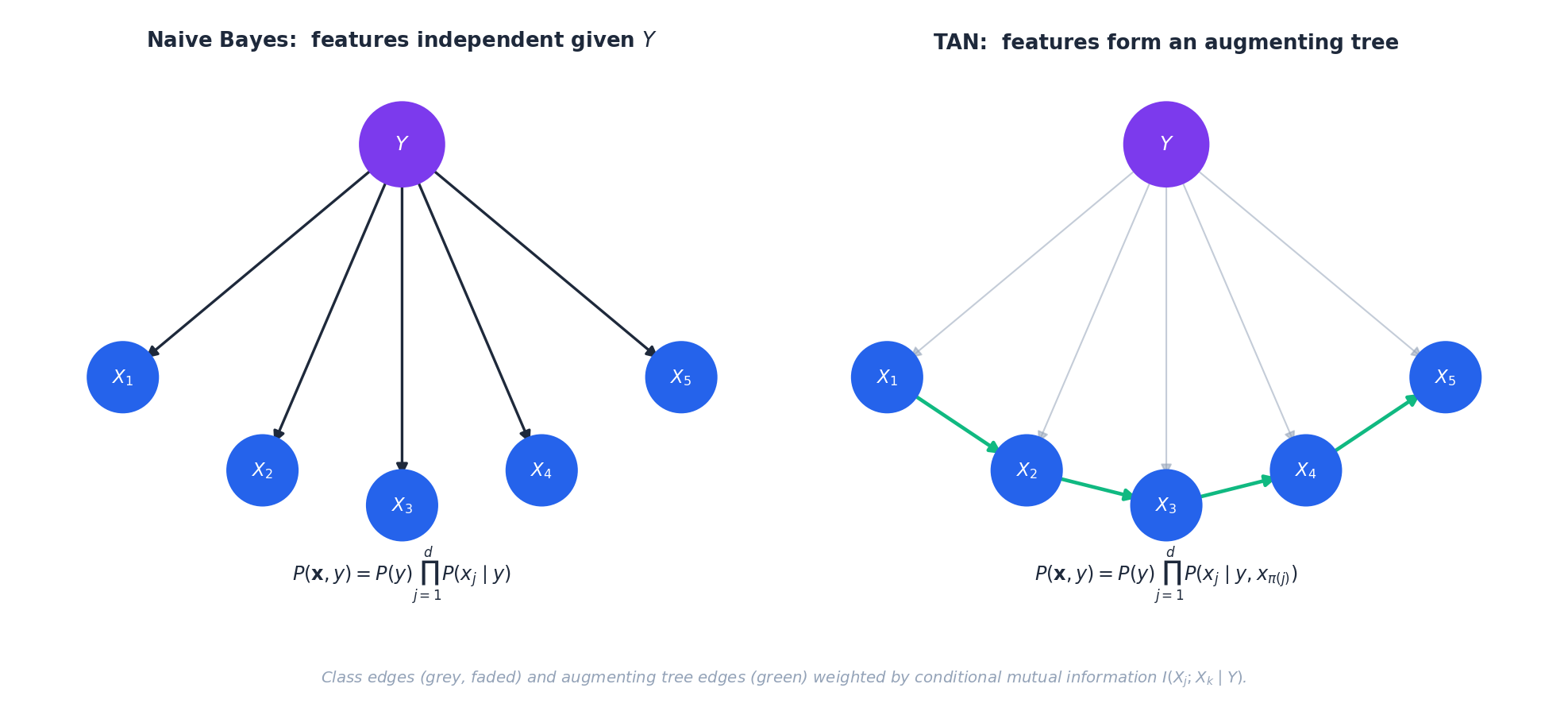
where $\pi_j$ is the unique parent of feature $j$ in the augmenting tree (the root has no parent, reducing to $P(x^{(j)}\mid c_k)$ ).
Learning the tree — Chow-Liu#
$$ w_{jk} \;=\; I(X^{(j)};X^{(k)}\mid Y) \;=\; \sum_{y,x_j,x_k} P(x_j,x_k,y)\, \log\frac{P(x_j,x_k\mid y)}{P(x_j\mid y)\,P(x_k\mid y)}. $$Reading the weight. $I(X^{(j)};X^{(k)}\mid Y)$ measures the residual correlation between $X^{(j)}$ and $X^{(k)}$ that the class has not already explained. Edges with high weight are exactly the dependencies most worth modelling.
Algorithm.
- Build a complete graph over the $d$ features.
- Set edge weight $(j,k)$ to $I(X^{(j)};X^{(k)}\mid Y)$ .
- Run Kruskal or Prim to obtain the maximum spanning tree — $O(d^{2}\log d)$ .
- Pick a root (commonly $\arg\max_j I(X^{(j)};Y)$ ).
- Orient edges away from the root.
| |
The CMI for the dependent pair $(x_0,x_1)$ in the synthetic example above will be visibly larger than for the independent pair $(x_0,x_2)$ — that ordering is exactly what Kruskal will exploit.
Bayesian networks#
Both NB and TAN are special cases of a general object: a Bayesian network. We now zoom out to the full graphical-models picture.
DAG factorisation#
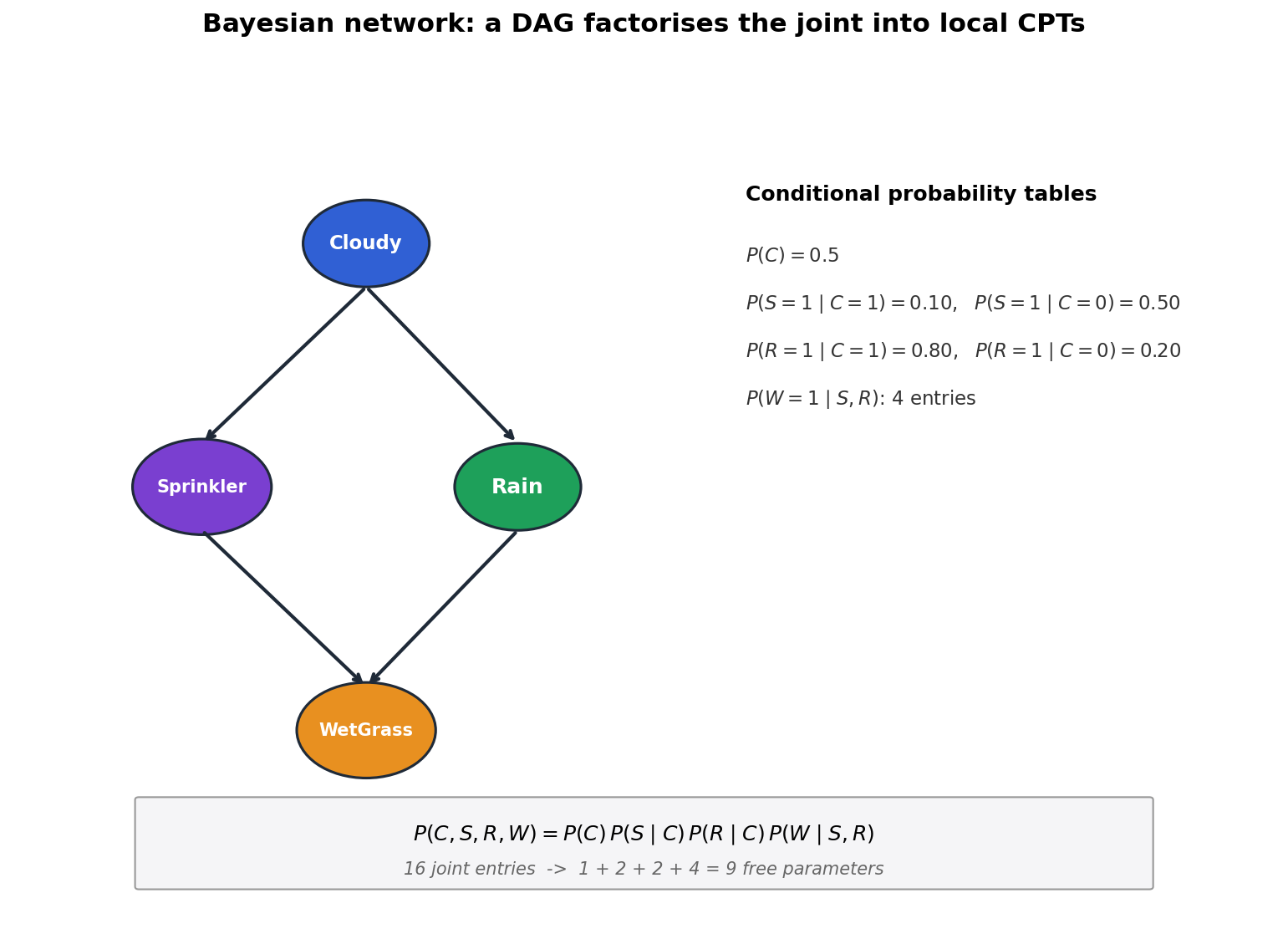
Why factorisation matters. In the four-variable network above, the joint over four binary variables would have $2^4 - 1 = 15$ free parameters. The factored form needs $1+2+2+4 = 9$ . With $n=20$ binary variables and average parent count 3, the joint has $\approx 10^{6}$ entries while the factored form has $\sim 20\cdot 2^{3}=160$ .
Special cases we have already met:
- Naive Bayes is the star graph: $Y$ is the parent of every $X_j$ , no edges among the $X_j$ .
- TAN is a star plus a tree on the $X_j$ .
d-separation#
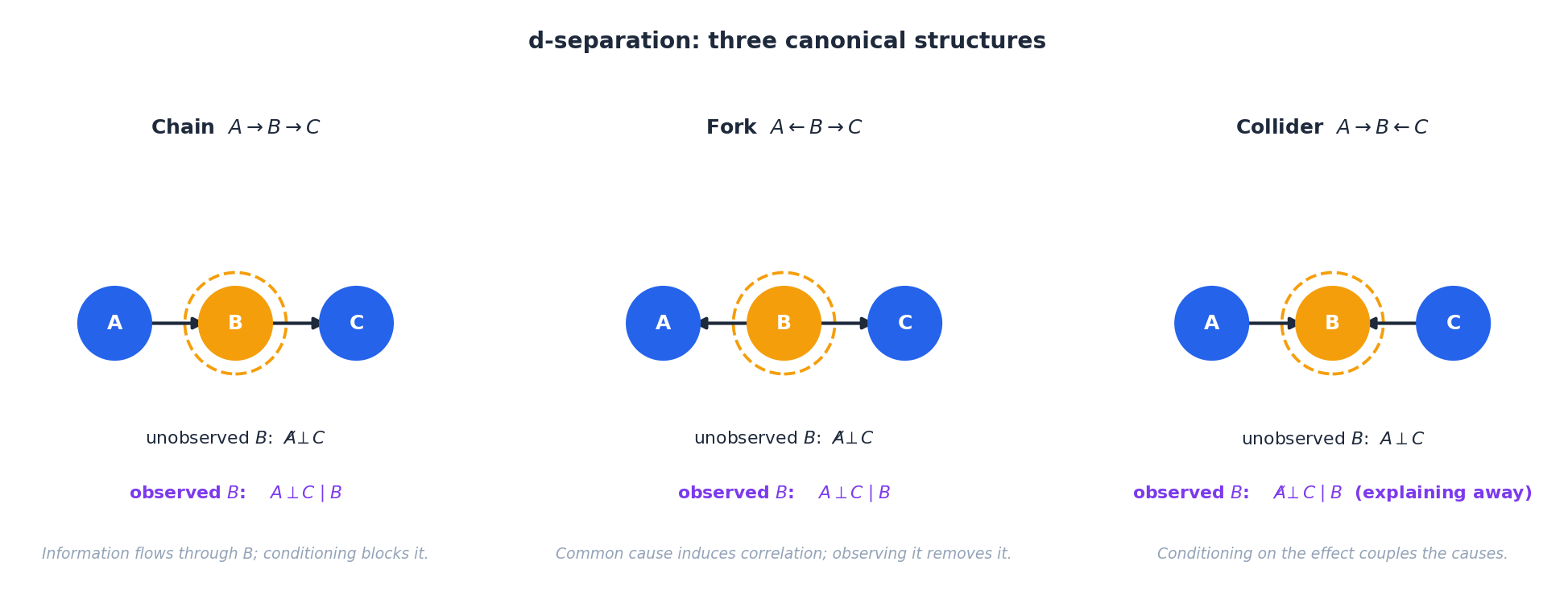
Conditional independence in a DAG is decided by the graphical criterion d-separation. Three structures generate every case:
- Chain $A\to B\to C$ . With $B$ unobserved, $A\not\!\perp C$ . Conditioning on $B$ blocks the path: $A\perp C\mid B$ .
- Fork $A\leftarrow B\to C$ . Same behaviour as the chain: a common cause makes $A,C$ correlated, observing it removes the correlation.
- Collider $A\to B\leftarrow C$ . Behaves oppositely. Without $B$ , $A\perp C$ . Observing $B$ activates the path, coupling its two causes — the famous explaining-away effect.
A path is blocked if any chain or fork node on it is observed, or any collider on it is not observed (and none of its descendants are). Two sets are d-separated by $Z$ iff every path between them is blocked by $Z$ . This is the engine that drives every conditional-independence claim in graphical models.
Markov blanket#
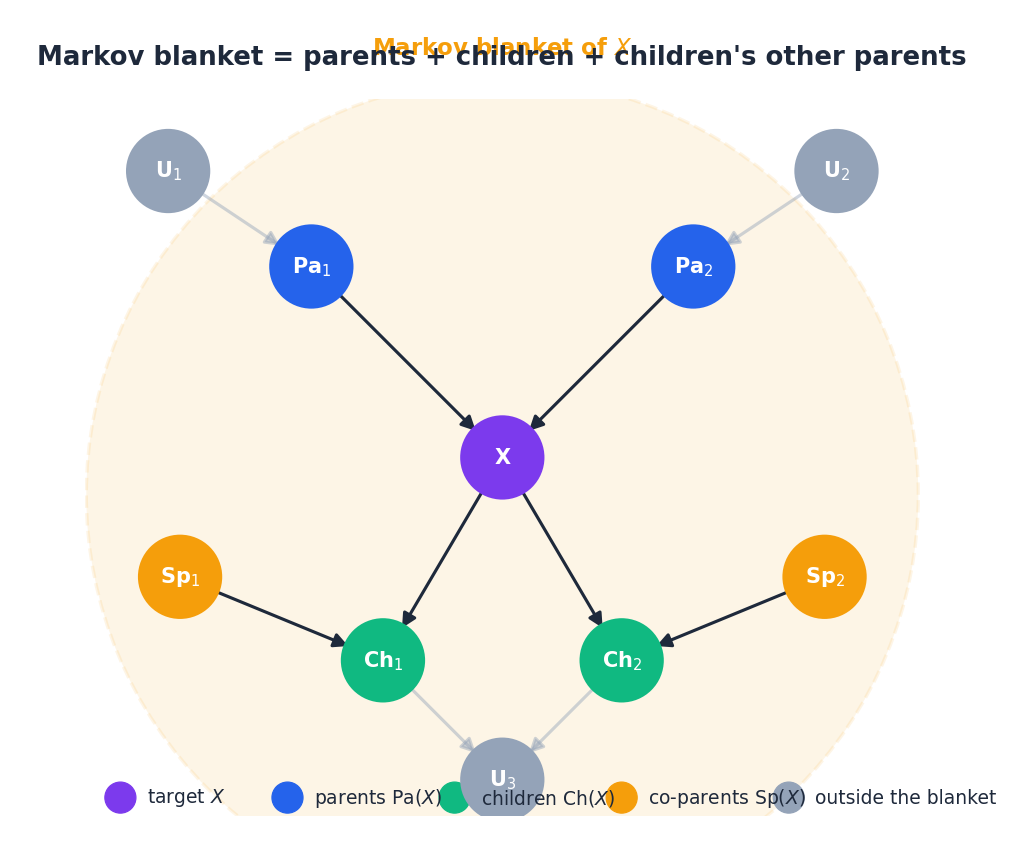
The co-parents are easy to forget but essential — they appear because, by the collider rule, conditioning on a child activates the path through that child to its other parents. The blanket is the minimal set of features a Gibbs sampler needs to resample $X$ , and in causal terms it is the smallest sufficient statistic for predicting $X$ from the rest of the network.
Inference: variable elimination#
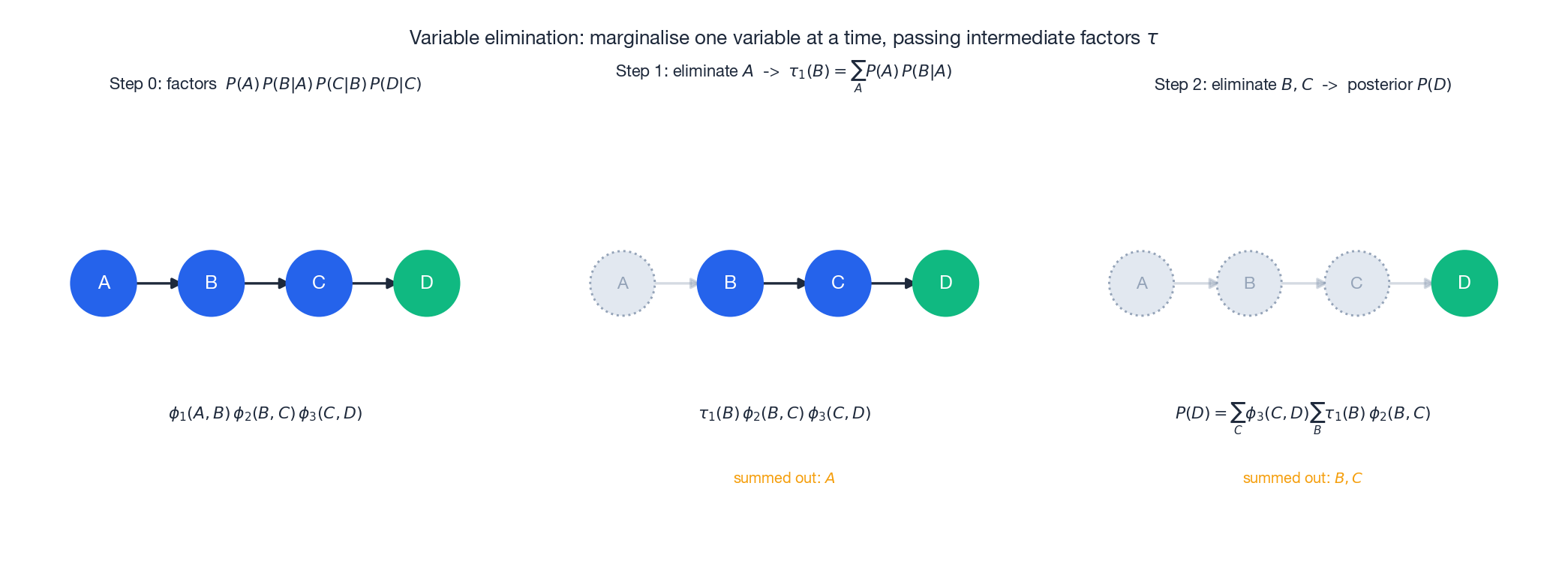

To compute a posterior such as $P(D\mid \text{evidence})$ , the brute-force sum is exponential in $n$ . Variable elimination sums variables out one at a time, reusing intermediate factors:
- Pick an elimination order, e.g. $A,B,C$ for a query about $D$ .
- Multiply all factors that mention $A$ , then $\sum_A$ to obtain a new factor $\tau_1(B)$ .
- Repeat: multiply factors that mention $B$ , sum to get $\tau_2(C)$ , and so on.
For a chain this is $O(nS^{2})$ instead of $O(S^{n})$ . For a general graph the complexity is governed by the treewidth of the graph — equivalently, by the largest intermediate factor produced — which is why dense graphs remain hard even after elimination.
The junction tree#
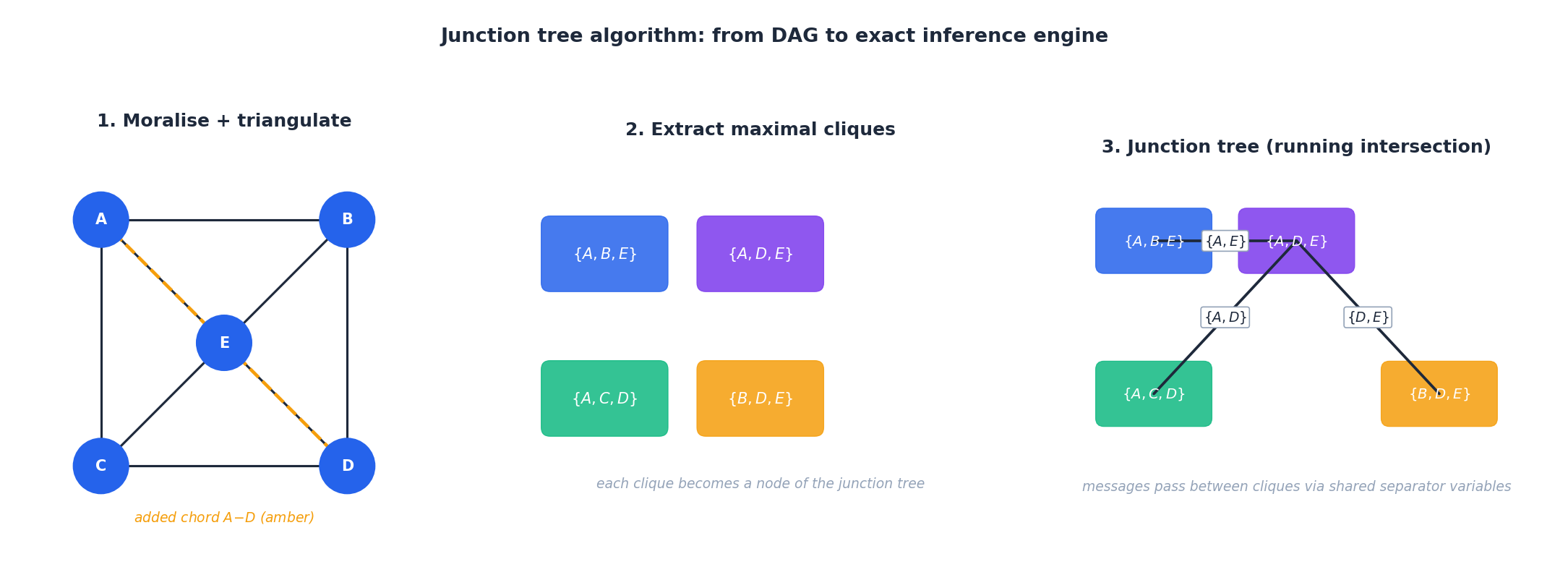
The junction tree algorithm globalises variable elimination so that all marginals can be computed by two message-passing sweeps over a single auxiliary structure:
- Moralise. Add an undirected edge between every pair of co-parents and drop edge directions.
- Triangulate. Add chords until every cycle of length $\geq 4$ has a chord. This step controls treewidth and is NP-hard in the optimal case — heuristics such as minimum fill-in work well in practice.
- Extract maximal cliques of the triangulated graph.
- Build a tree of cliques in which adjacent cliques share a separator set, satisfying the running-intersection property: any variable appears in a connected sub-tree.
- Pass messages along the tree (collect-then-distribute). After two sweeps, every clique holds the correct marginal over its variables.
The cost is exponential in the largest clique size, i.e. treewidth $+\,1$ — the same complexity bound as variable elimination, but amortised across all queries.
Structure learning#
The DAG itself can be learned. Two main families:
- Score + search. Hill-climb over DAGs by adding/deleting/reversing edges; score each candidate with a likelihood-plus-penalty criterion. The standard one is BIC:
The first term rewards fit, the second penalises parameter count — BIC is consistent (selects the true graph as $N\to\infty$ ) under standard assumptions.
- Constraint-based (PC algorithm, FCI). Run conditional-independence tests, build the skeleton, then orient v-structures using the collider rule.
A hard fact: exact structure learning is NP-hard (Chickering, Heckerman & Meek 2004). All practical algorithms cap the number of parents per node, use random restarts, or exploit prior knowledge about edge directions.
Inference complexity in Bayesian networks#
The structural decomposition $P(\mathbf{x}) = \prod_i P(x_i \mid \mathrm{pa}(x_i))$ buys you compact storage, but inference — answering $P(X_q \mid \mathbf{x}_e)$ — is in general #P-hard. The complexity hides in how nicely the network factorises along an elimination order.
$$\phi'(\mathbf{Y}) = \sum_{x_k} \prod_{f \in \mathcal{F}_k} f(x_k, \mathbf{Y}).$$The cost of one elimination step is exponential in the width — the size of the largest scope $\mathbf{Y} \cup \{x_k\}$ encountered. The optimal elimination order minimises the induced treewidth, which is in general NP-hard to compute. In practice, min-fill or min-neighbour heuristics work well on networks with hundreds of nodes.
For tree-structured networks (no undirected cycles in the moralised graph), treewidth is 1 and exact inference is $O(N \cdot K^2)$ for $N$ variables of cardinality $K$ — exactly the forward-backward cost of HMMs, which are the chain-structured special case. For dense networks like fully connected medical diagnosis BNs, treewidth grows linearly with $N$ and exact inference becomes infeasible by $N \approx 30$ .
Approximate alternatives. When treewidth is too large, the standard fallbacks are loopy belief propagation (run BP on the cyclic graph anyway, hope it converges; works empirically on many real networks but with no guarantee), Gibbs sampling (cycle through variables, sample each from its conditional given current values of others), and variational methods (Part 14 ). For a dense BN with binary variables and treewidth 25, a single Gibbs chain of 100k samples typically gives 2-3 digit accuracy on marginals — slow, but at least it returns.
Structure learning: where the hard problem actually is#
Given fixed structure $G$ , MLE for the conditional probability tables is trivial — count and divide. The genuinely hard problem is learning $G$ itself from data. Two families dominate.
$$\mathrm{BIC}(G, \mathcal{D}) = \log P(\mathcal{D} \mid \hat\theta_G, G) - \frac{|\theta_G|}{2}\log N,$$and BDeu (Bayesian Dirichlet equivalent uniform), the marginal likelihood under a uniform Dirichlet prior. Both are decomposable — the score factorises over families $(x_i, \mathrm{pa}(x_i))$ — so local edge changes only require rescoring affected families. Even so, the space of DAGs on $n$ nodes is super-exponential ($\sim n! \cdot 2^{\binom{n}{2}}$ ), so search is greedy: hill-climbing with random restarts, tabu search, or for small $n$ ($\lesssim 25$ ) the dynamic-programming exact algorithm of Silander-Myllymäki.
Constraint-based search. Test conditional independencies in the data and orient edges accordingly. The PC algorithm starts from the complete graph, removes edge $(X, Y)$ whenever it finds a separating set $\mathbf{Z}$ such that $X \perp Y \mid \mathbf{Z}$ , then orients colliders using the Meek rules. Cleaner theoretical guarantees than score-based search (under faithfulness, recovers the Markov equivalence class), but very sensitive to the conditional independence test’s Type-I error on finite samples. I have only ever seen PC work well on toy datasets; on real noisy data, score-based hill-climbing wins.
The TAN advantage revisited. TAN restricts the search to trees augmented over the class node, which is solvable exactly by Chow-Liu in $O(n^2 N)$ time using the maximum-weight spanning tree on conditional mutual information. This is why TAN is the practitioner’s compromise — almost all of Naive Bayes’s robustness, much of a full BN’s expressiveness, polynomial structure learning.
Worked exercises#
Exercise 1 — SPODE parameter estimate#
$$\hat{P}\bigl(x^{(j)}{=}1\mid c_1, x^{(p)}{=}1\bigr) \;=\; \frac{18+1}{30+1\cdot 3} \;=\; \frac{19}{33} \;\approx\; 0.576.$$Exercise 2 — TAN parameter count#
For binary features and $K$ classes:
- Naive Bayes: $K\cdot d$ parameters (one per class-feature combination).
- TAN: each non-root feature has the class and one binary parent, hence $2$ extra entries per class-feature pair — total $2Kd$ .
TAN doubles the parameter budget and gains the ability to model pairwise dependencies. With $S$ -valued features the multiplier is $S$ instead of $2$ .
Exercise 3 — Chow-Liu edge selection#
Suppose $I(X_1;X_2\mid Y)=0.8$ and $I(X_1;X_3\mid Y)=0.5$ . The Chow-Liu MST greedily picks the heavier edge first, so $X_1\!-\!X_2$ enters the tree before $X_1\!-\!X_3$ . The lighter edge enters only if it does not close a cycle.
Exercise 4 — d-separation#
In $A\to C\leftarrow B$ :
- (a) $A\perp B$ ? Yes. $C$ is a collider, and unobserved colliders block the path.
- (b) $A\perp B\mid D$ where $D$ is unrelated? Yes. $D$ is not a descendant of $C$ , so the collider stays blocked.
- (c) $A\perp B\mid C$ ? No. Conditioning on a collider activates the path — explaining away couples $A$ and $B$ .
Exercise 5 — AODE vs TAN with limited data#
- Large $d$ , plenty of data: TAN’s $O(d)$ prediction beats AODE’s $O(d^{2})$ .
- Small $N$ : AODE’s averaging acts as regulariser, often more robust than TAN’s structure search which can overfit MST edges from noisy CMI estimates.
A reasonable default policy: start with AODE; switch to TAN once $N$ comfortably exceeds the number of CMI cells you need to estimate.
FAQ#
Why not always use a full Bayesian network?#
Structure learning is NP-hard and the search space of DAGs is super-exponential. TAN and AODE constrain the structure to keep learning tractable while capturing the most informative dependencies.
Do directed edges encode causation?#
No. Two DAGs can encode the same conditional-independence structure (an equivalence class) and be statistically indistinguishable. Causal claims require interventional or counterfactual assumptions — Pearl’s do-calculus, instrumental variables, etc.
Continuous features in TAN?#
Three options: (i) discretise (equal-width or equal-frequency bins), (ii) assume conditional Gaussians (a Conditional Gaussian Bayesian network), (iii) use kernel density estimates for the local CPDs.
AODE or TAN — which?#
AODE: simpler, more robust on small samples, no structure-learning step. TAN: faster prediction at high $d$ , captures heterogeneous parent choices. In production text classifiers AODE is the more common default; in dense numeric tables, TAN often wins.
Where do deep generative models fit?#
Modern latent-variable models (VAE, normalising flows, autoregressive models like PixelCNN) are continuous Bayesian networks where the CPTs are neural conditional densities. The factorisation idea is identical; only the parametric form of $P(X_i\mid \mathrm{Pa}(X_i))$ has changed.
What’s next#
Bayesian networks let me draw the dependence structure explicitly, but they expose a new problem: however accurate one model is, it can still have high variance from data noise, initialization, feature sampling. The next chapter switches strategy entirely — ensemble learning.
The ensemble bet is: rather than tune one model to perfection, train a crowd of base learners and cancel their errors out by voting (Bagging) or weighting (Boosting). Bagging cuts variance by resampling; random forests further decorrelate by feature subsampling; Boosting stacks weak learners along the gradient direction to eat away the bias. Both lines rest on a single algebraic fact — the variance of a weighted average is controlled by the correlation $\rho$ between members. Internalising that is more important than memorising any specific algorithm.
References#
- Friedman, N., Geiger, D. & Goldszmidt, M. (1997). Bayesian network classifiers. Machine Learning, 29(2-3), 131-163.
- Webb, G. I., Boughton, J. R. & Wang, Z. (2005). Not so naive Bayes: Aggregating one-dependence estimators. Machine Learning, 58(1), 5-24.
- Chow, C. & Liu, C. (1968). Approximating discrete probability distributions with dependence trees. IEEE Transactions on Information Theory, 14(3), 462-467.
- Chickering, D. M., Heckerman, D. & Meek, C. (2004). Large-sample learning of Bayesian networks is NP-hard. JMLR, 5, 1287-1330.
- Koller, D. & Friedman, N. (2009). Probabilistic Graphical Models: Principles and Techniques. MIT Press.
- Pearl, J. (1988). Probabilistic Reasoning in Intelligent Systems. Morgan Kaufmann.
ML Mathematical Derivations Series
< Part 9: Naive Bayes | Part 10: Semi-Naive Bayes & Bayesian Networks | Part 11: Ensemble Learning >
ML Math Derivations 20 parts
- 01 ML Math Derivations (1): Introduction and Mathematical Foundations
- 02 ML Math Derivations (2): Linear Algebra and Matrix Theory
- 03 ML Math Derivations (3): Probability Theory and Statistical Inference
- 04 ML Math Derivations (4): Convex Optimization Theory
- 05 ML Math Derivations (5): Linear Regression
- 06 ML Math Derivations (6): Logistic Regression and Classification
- 07 ML Math Derivations (7): Decision Trees
- 08 ML Math Derivations (8): Support Vector Machines
- 09 ML Math Derivations (9): Naive Bayes
- 10 ML Math Derivations (10): Semi-Naive Bayes and Bayesian Networks you are here
- 11 ML Math Derivations (11): Ensemble Learning
- 12 ML Math Derivations (12): XGBoost and LightGBM
- 13 ML Math Derivations (13): EM Algorithm and GMM
- 14 ML Math Derivations (14): Variational Inference and Variational EM
- 15 ML Math Derivations (15): Hidden Markov Models
- 16 ML Math Derivations (16): Conditional Random Fields
- 17 ML Math Derivations (17): Dimensionality Reduction and PCA
- 18 ML Math Derivations (18): Clustering Algorithms
- 19 ML Math Derivations (19): Neural Networks and Backpropagation
- 20 ML Math Derivations (20): Regularization and Model Selection