
ML Math Derivations (11): Ensemble Learning
Derive why combining weak learners produces strong ones. Covers bias-variance decomposition, Bagging/Random Forest variance reduction, AdaBoost exponential loss, and GBDT gradient optimization in function space.
Why do mediocre classifiers in a committee outperform a single brilliant one? The answer is straightforward: averaging reduces variance, sequential reweighting reduces bias, and a bit of randomization breaks the correlation that would otherwise negate these benefits. This post delves into the math behind this — bias-variance decomposition, bootstrap aggregating, AdaBoost as forward stagewise minimization of exponential loss, and gradient boosting as gradient descent in function space.
By the end, you should be able to look at any ensemble method and say what it reduces, why it works, and when it fails.

What You Will Learn#
- Why averaging $T$ uncorrelated models cuts variance by a factor of $T$ , and what happens when they are correlated.
- How Bagging and Random Forest exploit randomisation to make trees nearly uncorrelated.
- The derivation that links AdaBoost to forward stagewise additive modelling under exponential loss.
- How GBDT performs gradient descent on the loss surface in function space.
- When to choose Bagging vs Boosting vs Stacking for a given problem.
Prerequisites#
- Bias–variance tradeoff (Parts 1–2 of this series).
- Decision tree basics (splits, impurity).
- Gradient descent.
- Probability: expectation and variance of sums of random variables.
Why ensembles work#
The variance reduction identity#
$$H(\mathbf{x}) = \frac{1}{T}\sum_{t=1}^T h_t(\mathbf{x}).$$ $$ \mathbb{E}[H(\mathbf{x})] = \mu, \qquad \operatorname{Var}[H(\mathbf{x})] = \rho\,\sigma^2 + \frac{1-\rho}{T}\,\sigma^2. $$This equation is the core reason ensembles exist. Read it carefully:
- Bias is preserved. $\mathbb{E}[H] = \mu$ , so averaging cannot fix systematic error of the base learner. If every tree underfits, the ensemble underfits too.
- Variance has two pieces. A floor of $\rho\sigma^2$ that we cannot remove by adding more learners, plus a $\sigma^2/T$ term that vanishes as $T \to \infty$ — but only when learners are uncorrelated ($\rho = 0$ ).
- Correlation is the enemy. Even with $T = \infty$ trees, variance bottoms out at $\rho\sigma^2$ . This is exactly why Random Forest randomises features, not just samples: feature randomisation is a direct attack on $\rho$ .
So every ensemble method is, at heart, an answer to two questions: how do I generate diverse learners (small $\rho$ ) without making them too weak (large bias)?
Bias–variance decomposition#
$$ \mathbb{E}[(y - \hat f(\mathbf{x}))^2] = \underbrace{(\mathbb{E}[\hat f] - f)^2}_{\text{bias}^2} \,+\, \underbrace{\mathbb{E}[(\hat f - \mathbb{E}[\hat f])^2]}_{\text{variance}} \,+\, \underbrace{\sigma_\epsilon^2}_{\text{irreducible noise}}. $$Complex models (deep trees, large neural nets) have low bias but high variance. Simple models (depth-1 stumps, linear regression) have low variance but significant bias. The two ensemble families address these issues differently:
- Bagging / Random Forest keeps a low-bias high-variance learner and averages away the variance.
- Boosting starts from a high-bias low-variance learner and adds capacity to drive down the bias.
That is the entire taxonomy in two sentences.
Why a committee of mediocre voters works#
$$P_{\text{ensemble}} = \sum_{k > T/2} \binom{T}{k}\,\epsilon^k(1-\epsilon)^{T-k}.$$With $T = 21$ and $\epsilon = 0.30$ this evaluates to about $0.026$ . A 30 % error rate is mediocre; a 2.6 % ensemble error rate is excellent. The catch is the word independent — which, again, points back to the decorrelation problem.
Bagging and Random Forest#
Bagging: the parallel recipe#
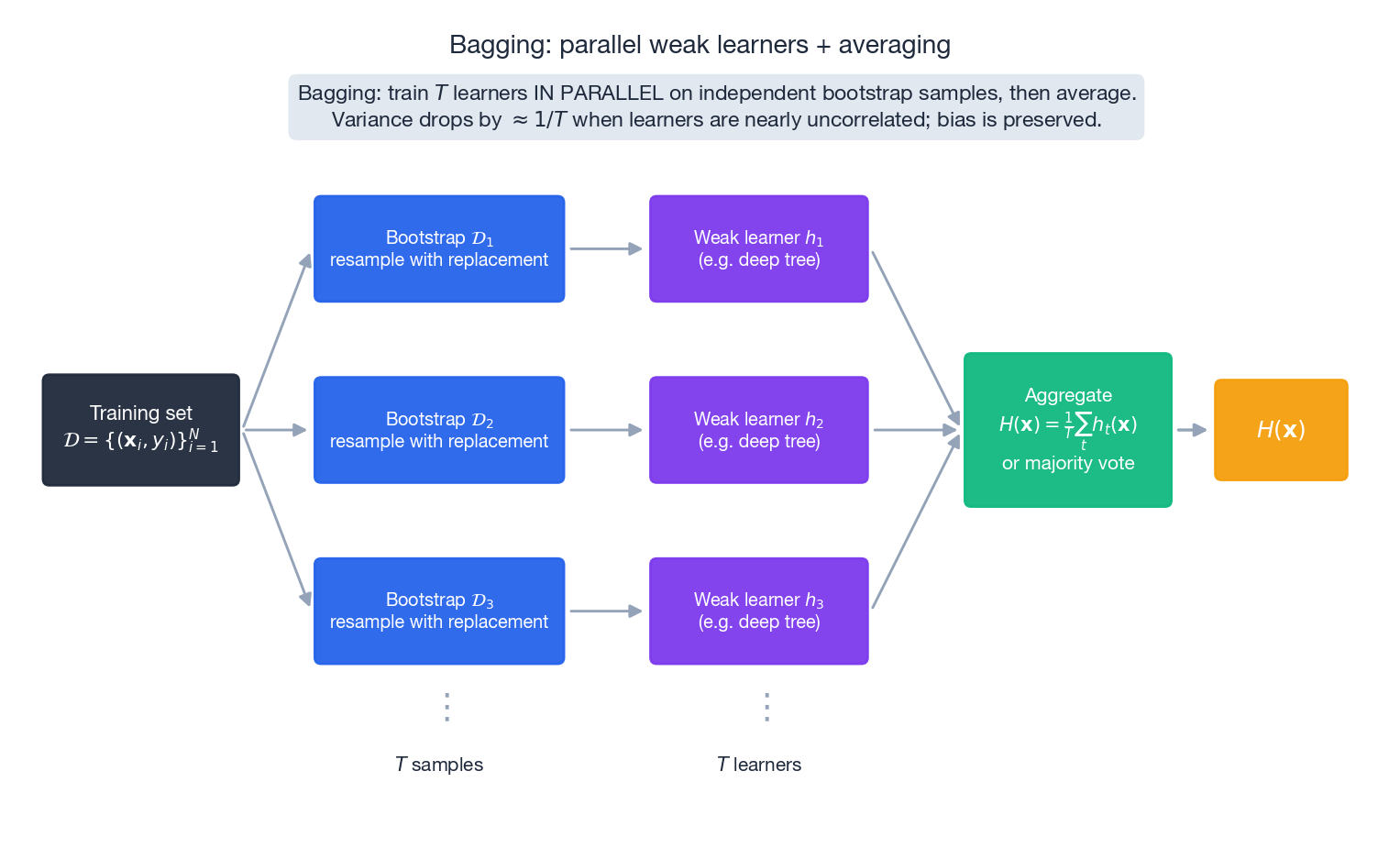
Bagging (Breiman, 1996) is the clearest way to apply the variance identity from §1.1.
Algorithm.
- Bootstrap. From training set $\mathcal{D}$ of size $N$ , draw $T$ samples with replacement, each of size $N$ . Call them $\mathcal{D}_1,\dots,\mathcal{D}_T$ .
- Train. Fit base learner $h_t$ on $\mathcal{D}_t$ independently. Trees are usually grown deep (low bias, high variance — exactly what we want to average down).
- Aggregate. Regression: $H(\mathbf{x}) = \tfrac{1}{T}\sum_t h_t(\mathbf{x})$ . Classification: majority vote.
These leftover points are the out-of-bag (OOB) samples for tree $t$ .
$$ \widehat{\text{Err}}_{\text{OOB}} = \frac{1}{N}\sum_{i=1}^N L\!\left(y_i,\; \frac{1}{|\mathcal{S}_i|}\sum_{t \in \mathcal{S}_i} h_t(\mathbf{x}_i)\right),\qquad \mathcal{S}_i = \{t : (\mathbf{x}_i, y_i) \notin \mathcal{D}_t\}. $$This is an (almost) unbiased estimate of generalization error, computed for free as a side effect of training. No need for a held-out set or cross-validation loop.
Random Forest: decorrelating the trees#
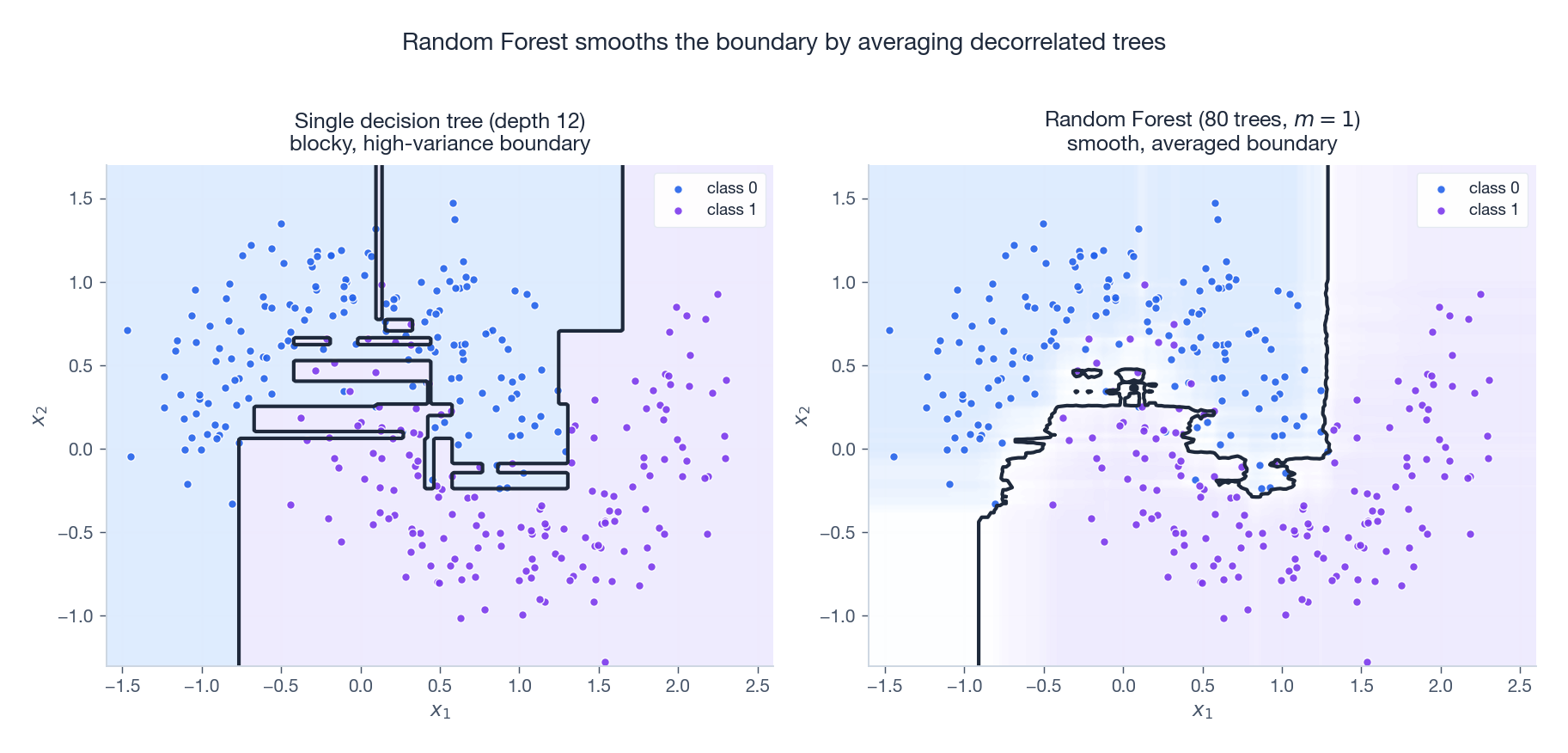
Bagging alone leaves a problem: bootstrap samples overlap heavily, so the trees end up making the same splits on the same dominant features. Their predictions are correlated, $\rho$ stays large, and the variance reduction in §1.1 stalls.
Random Forest (Breiman, 2001) adds a second source of randomness. At every node split, sample a random subset of $m$ features out of $d$ and only consider splits on those. Typical defaults:
- Classification: $m = \lfloor\sqrt{d}\rfloor$ .
- Regression: $m = \lfloor d/3 \rfloor$ .
Forcing each tree to use different features at each split breaks the dominant-feature trap and pushes $\rho$ down. The figure above makes the effect visible: a single deep tree carves the input space into rectangular blocks and overfits the noise; a Random Forest with $m=1$ averages 80 such blocky surfaces and produces a smooth, generalising boundary.
$$\text{Generalisation error} \;\le\; \frac{\bar\rho\,(1 - s^2)}{s^2}.$$This is the variance-reduction identity dressed up. To improve a forest you have exactly two knobs:
- Strengthen each tree (raise $s$ ): grow deeper, use more features per split, use more samples.
- Decorrelate the trees (lower $\bar\rho$ ): reduce $m$ , vary depths, add more trees.
The two knobs trade off, which is why $m$ is a tuning parameter, not a constant.
Feature importance#
Two ways to score features:
- Mean decrease in impurity. Sum the impurity drop $\Delta\text{Gini}$ at every node that splits on feature $j$ , averaged across trees. Cheap, but biased toward high-cardinality features.
- Permutation importance. Take an OOB sample, shuffle column $j$ , and measure how much the OOB accuracy drops. Slower but unbiased, and it correctly handles correlated features.
When two features carry similar signal, mean-decrease-in-impurity splits the importance between them; permutation importance attributes the same drop to each, which is usually what you want for interpretability.
Bagging vs Boosting — the bias–variance picture#
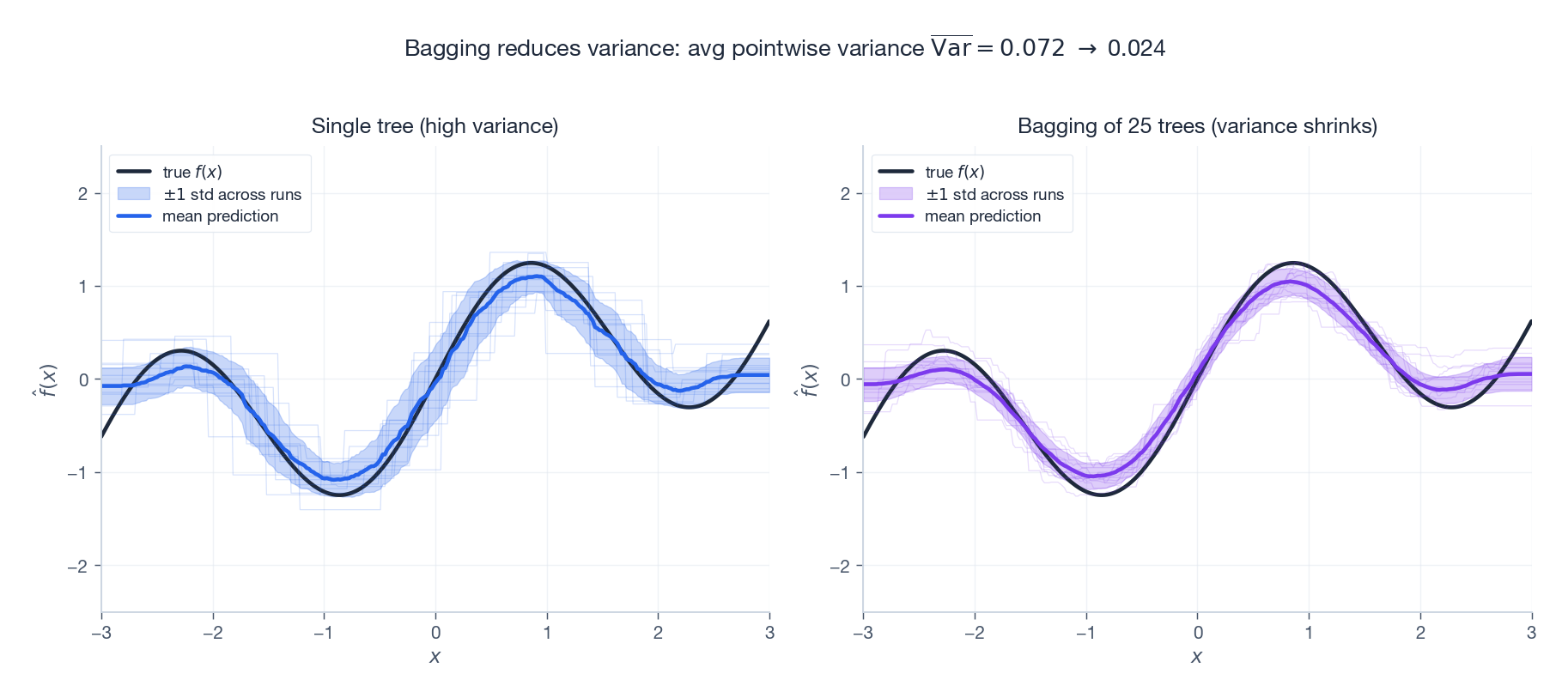
The figure above runs the same fitting procedure on 120 freshly drawn datasets and plots the predictions. Look at the spread:
- Single tree (left). The mean prediction is close to the truth (low bias), but each individual fit wiggles wildly (high variance). The shaded $\pm 1$ standard-deviation band is wide.
- Bagging of 25 trees (right). Same mean, dramatically narrower band. The variance number drops by roughly a factor of $T$ , exactly as §1.1 predicts.
Bagging cannot do anything about systematic bias — if the base learner is fundamentally too weak, all the averaging in the world will not help. That is the job of boosting.
Boosting: sequential bias reduction#
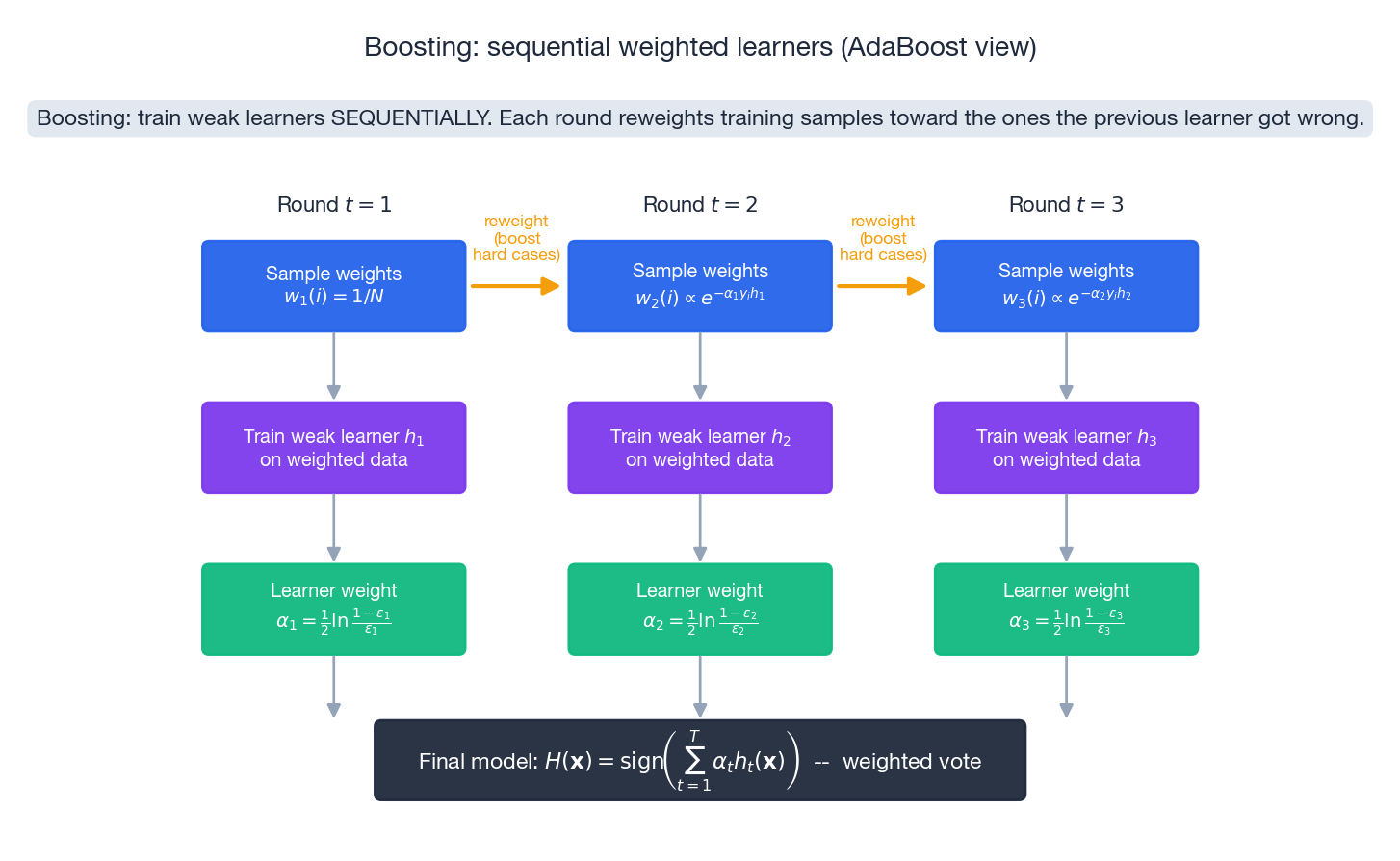
Boosting flips the picture. Instead of training $T$ learners in parallel and averaging, boosting trains them sequentially, each one focused on the mistakes of its predecessor. Each learner is intentionally weak (e.g. a depth-1 stump), which makes it high bias, low variance. The sequence then drives the bias down.
AdaBoost: the algorithm#
For binary labels $y_i \in \{-1, +1\}$ , initialise weights $w_1(i) = 1/N$ . For $t = 1,\dots,T$ :
- Train weak learner $h_t$ on the weighted data.
- Compute weighted error $\epsilon_t = \sum_i w_t(i)\,\mathbf{1}[h_t(\mathbf{x}_i) \neq y_i]$ .
- Compute learner weight $\alpha_t = \tfrac{1}{2}\ln\tfrac{1-\epsilon_t}{\epsilon_t}$ .
- Update sample weights $w_{t+1}(i) = w_t(i)\exp(-\alpha_t y_i h_t(\mathbf{x}_i)) / Z_t$ , where $Z_t$ normalises.
Output $H(\mathbf{x}) = \operatorname{sign}\bigl(\sum_t \alpha_t h_t(\mathbf{x})\bigr)$ .
Three things to notice in the formulas:
- $\alpha_t$ is a log-odds. A learner with $\epsilon_t = 0.1$ gets $\alpha_t \approx 1.10$ ; one with $\epsilon_t = 0.49$ gets $\alpha_t \approx 0.02$ . Better learners get larger votes.
- $\alpha_t > 0 \iff \epsilon_t < 1/2$ . A learner worse than random gets a negative weight — AdaBoost flips its predictions and keeps it. Nothing is wasted.
- Weight update is multiplicative. Correctly classified samples ($y_i h_t = +1$ ) shrink by $e^{-\alpha_t}$ ; misclassified ones grow by $e^{+\alpha_t}$ . The next round literally cannot ignore the hard cases.
The exponential training-error bound#
$$ \frac{1}{N}\sum_{i=1}^N \mathbf{1}[H(\mathbf{x}_i) \neq y_i] \;\le\; \prod_{t=1}^T Z_t. $$ $$ \text{train err}(H) \;\le\; \prod_{t=1}^T \sqrt{1 - 4\gamma_t^2} \;\le\; \exp\!\left(-2\sum_{t=1}^T \gamma_t^2\right). $$Training error decays exponentially in $T$ . The figure below shows this on a synthetic problem with a handful of intentionally hard borderline points:
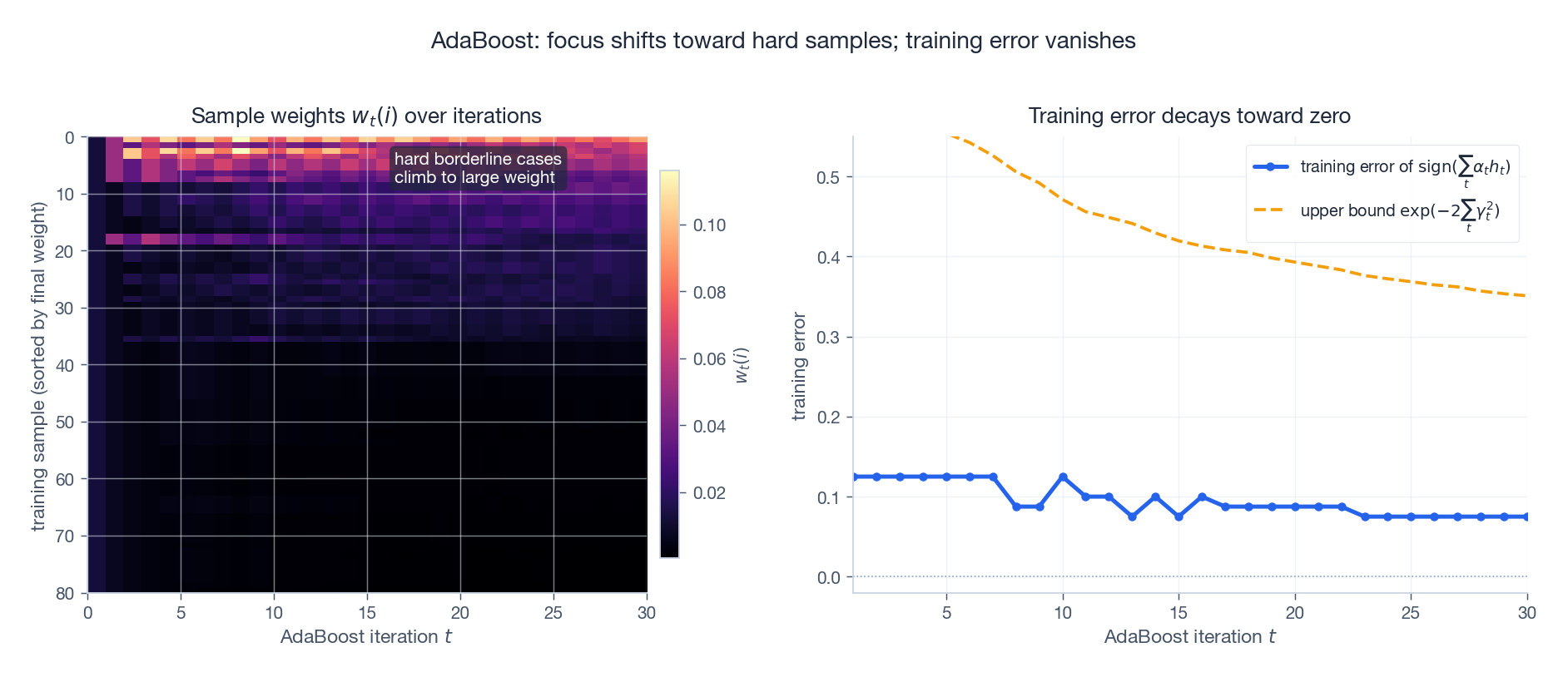
The left heatmap traces every sample’s weight across iterations: most points fade to near zero while the borderline cases light up brightly — the algorithm is literally diverting all its attention to the points the previous learners cannot solve. The right panel tracks the cumulative classifier’s training error against the theoretical $\exp(-2\sum_t \gamma_t^2)$ bound; the bound is loose but always above the truth.
This is also AdaBoost’s failure mode: with genuinely mislabelled points the same dynamic dumps all of the model’s capacity on the noise. GBDT with a robust loss (e.g. Huber) handles that case much more gracefully.
Why exponential weights? The forward stagewise view#
$$F(\mathbf{x}) = \sum_{t=1}^T \alpha_t\, h_t(\mathbf{x}),$$ $$(\alpha_t, h_t) \;=\; \arg\min_{\alpha, h}\sum_{i=1}^N \exp\bigl(-y_i\,(F_{t-1}(\mathbf{x}_i) + \alpha\, h(\mathbf{x}_i))\bigr).$$ $$\min_{\alpha, h}\sum_{i=1}^N w_t(i)\exp(-\alpha\, y_i\, h(\mathbf{x}_i)).$$Solving for the optimal $h_t$ first (it minimises weighted error) and then for $\alpha_t$ (one-dimensional calculus) recovers exactly AdaBoost’s update formulas. Nothing is heuristic; it is coordinate descent in function space, with one new basis function added per iteration.
Gradient Boosting Decision Trees (GBDT)#
Boosting as gradient descent in function space#
Exponential loss is fragile. The key insight of Friedman (2001) was that the forward-stagewise idea works for any differentiable loss if we recast it as gradient descent.
$$r_{ti} \;=\; -\!\left[\frac{\partial L(y_i, F)}{\partial F}\right]_{F = F_{t-1}(\mathbf{x}_i)},\qquad i = 1,\dots,N.$$These $r_{ti}$ are the pseudo-residuals. A single regression tree fit to $\{(\mathbf{x}_i, r_{ti})\}$ is a finite-dimensional approximation of the steepest-descent direction in function space.
Algorithm (Friedman, 2001).
- Initialise $F_0(\mathbf{x}) = \arg\min_c \sum_i L(y_i, c)$ (e.g. the mean for squared loss).
- For $t = 1,\dots,T$
:
- Compute pseudo-residuals $r_{ti}$ .
- Fit regression tree $h_t$ to $\{(\mathbf{x}_i, r_{ti})\}$ .
- Line search the step size $\rho_t = \arg\min_\rho \sum_i L(y_i, F_{t-1}(\mathbf{x}_i) + \rho\, h_t(\mathbf{x}_i))$ .
- Update $F_t = F_{t-1} + \eta\,\rho_t\, h_t$ , where $\eta \in (0, 1]$ is the learning rate.
What the pseudo-residuals look like for common losses#
| Loss | $L(y, F)$ | Pseudo-residual $r_i$ | Notes |
|---|---|---|---|
| Squared (regression) | $\tfrac{1}{2}(y - F)^2$ | $y_i - F_{t-1}(\mathbf{x}_i)$ | Plain residual fitting. |
| Absolute (robust) | $\lvert y - F \rvert$ | $\operatorname{sign}(y_i - F_{t-1}(\mathbf{x}_i))$ | Robust to outliers, ignores magnitude. |
| Huber (robust) | piecewise | clipped residual | Best of both: smooth near zero, robust at the tails. |
| Logistic (binary) | $\log(1 + e^{-yF})$ | $y_i / (1 + e^{y_i F_{t-1}(\mathbf{x}_i)})$ | $F$ is log-odds; predict $\sigma(F)$ . |
For squared loss the algorithm reduces to “fit the residuals”, which was the original boosting intuition. For everything else the gradient-in-function-space picture is the only clean way to see what is happening.
Regularisation: the three knobs that actually matter#
GBDT will overfit aggressively if you let it. The standard defences:
- Shrinkage. $F_t = F_{t-1} + \eta\,h_t$ with small $\eta$ (typically $0.01$ to $0.1$ ). Smaller $\eta$ requires more trees but generalises better, just like a small SGD step size avoids overshooting. Always pair small $\eta$ with large $T$ .
- Stochastic gradient boosting. Each round, fit $h_t$ on a random subsample (e.g. 50–80 % of training rows). Acts like SGD on the function-space objective, decorrelates trees, and cheapens each round.
- Tree complexity caps. Limit depth (typically 3–6), minimum samples per leaf, or the number of leaves outright. Each tree stays a weak learner — the whole point of boosting.
XGBoost (next post) adds an explicit $L_2$ penalty on leaf weights and a leaf-count penalty, baked into a closed-form leaf optimisation. It is worth understanding plain GBDT first.
Stacking: meta-learning over heterogeneous models#
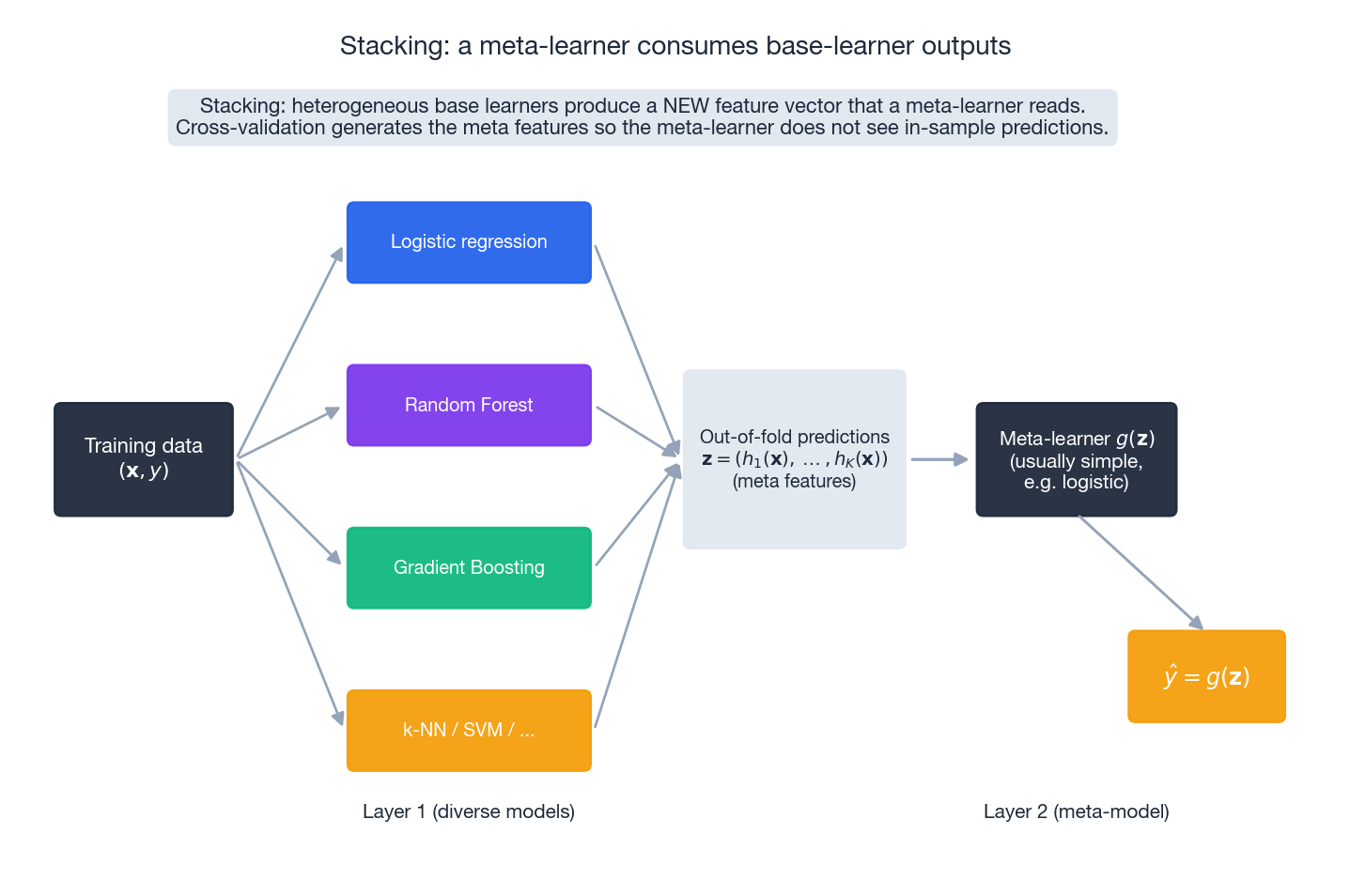
Bagging and Boosting use a single family of base learners. Stacking uses several:
- Layer 1. Fit $K$ base models (e.g. logistic regression, random forest, GBDT, k-NN) on the training set. To avoid leakage, generate predictions via out-of-fold cross-validation: every training row is predicted by a model that did not see it.
- Layer 2. Treat the $K$ out-of-fold predictions as a new feature vector $\mathbf{z} \in \mathbb{R}^K$ and fit a meta-learner $g(\mathbf{z}) \to \hat y$ .
The meta-learner is usually simple — logistic regression or a shallow GBDT — because the base learners have already done the heavy lifting. The hard part is the cross-validation plumbing; if any base prediction is in-sample, the meta-learner will learn to trust it blindly and overfit catastrophically.
When does stacking pay off? When the base learners make different kinds of mistakes. Stacking a deep model with a tree model with a linear model often beats any single one because their errors decorrelate.
Ensemble size: how many learners is enough?#
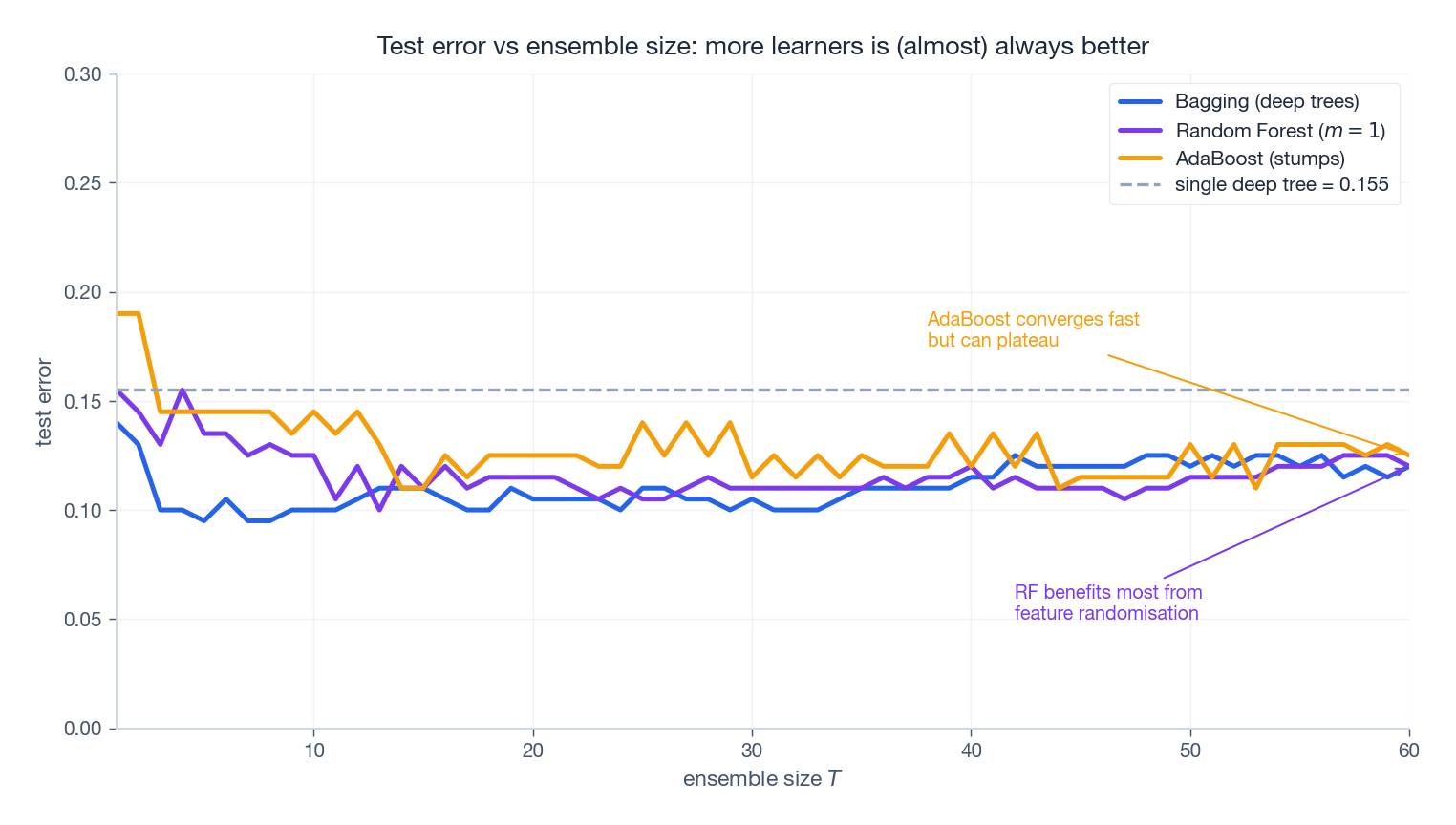
The figure trains all three ensembles on the same problem and tracks test error as $T$ grows. A few patterns to internalise:
- All three crush the single deep tree within the first dozen learners.
- Bagging and Random Forest plateau gracefully. More trees never hurt; they just stop helping. The flat tail is a direct consequence of the variance floor $\rho\sigma^2$ in §1.1.
- AdaBoost converges fastest but can increase test error in some regimes once it starts overfitting noise. Watch for it.
- Random Forest’s plateau is lower than vanilla Bagging because feature randomisation drops $\bar\rho$ .
Practical takeaway: for Bagging/RF, choose $T$ as large as your compute budget allows. For Boosting, choose $T$ via early stopping on a validation set.
Reference implementations#
The code below is intentionally short and dependency-free. It is not as fast as scikit-learn, but every step maps directly onto the formulas above.
| |
FAQ#
Q1. Bagging or Boosting — which is “better”? Wrong question. Bagging reduces variance; Boosting reduces bias. If your base learner overfits (deep tree on a small dataset), bag it. If your base learner underfits (a stump), boost it. They live on opposite sides of the bias–variance dilemma.
Q2. Why exactly the exponential weight update in AdaBoost? It is the closed-form solution of forward-stagewise minimisation of exponential loss. There is nothing magical about it — different losses give different update rules. Logistic loss gives LogitBoost; squared loss gives gradient-boosted residual fitting.
Q3. Why does a small learning rate help GBDT? Same reason a small step size helps SGD: each tree is a noisy estimate of the true descent direction. A small $\eta$ trusts no single tree too much, so individual mistakes get diluted. The cost is more trees; the benefit is better generalisation. Pair $\eta = 0.05$ with $T = 1000$ over $\eta = 0.5$ with $T = 100$ almost every time.
Q4. How do I choose $m$ for Random Forest? Defaults of $\sqrt{d}$ (classification) and $d/3$ (regression) are excellent starting points. If the OOB error is high and trees look too similar, decrease $m$ to inject diversity. If the OOB error is high and individual trees look too weak, increase $m$ .
Q5. Can Boosting be parallelised? The outer loop (rounds $t = 1, 2, \dots$ ) cannot be parallelised — each round depends on the previous. The inner loop (finding the best split inside a tree) is embarrassingly parallel across features and across data shards. XGBoost and LightGBM exploit this aggressively.
Q6. Does ensembling prevent overfitting? Bagging/RF: almost yes — adding trees rarely hurts, because variance is the only thing being reduced. Boosting: no — you can drive training error to zero while test error climbs. Cure with early stopping on a validation set, learning-rate shrinkage, subsampling, and tree-complexity caps.
Q7. Why is GBDT the foundation for XGBoost / LightGBM / CatBoost? Because the function-space view generalises: any differentiable loss, any tree learner, any second-order trick. XGBoost adds a Newton-step approximation and explicit regularisation; LightGBM adds histogram binning and leaf-wise growth; CatBoost adds ordered boosting for categorical features. All three are GBDT plus engineering.
Exercises#
Exercise 1 — Bias–variance arithmetic#
Three independent regression models each have $\text{bias}^2 = 4$ and $\text{Var} = 9$ . Compute the expected MSE of (i) a single model and (ii) the simple-average ensemble. Ignore noise.
Solution. Single: $4 + 9 = 13$ . Ensemble: bias unchanged at $4$ , variance reduced to $9/3 = 3$ , total $7$ . Improvement: $\approx 46\%$ .
Exercise 2 — Majority-vote error#
Twenty-one independent binary classifiers, each with error rate $\epsilon = 0.30$ . Compute the majority-vote error.
$$P_{\text{ensemble}} = \sum_{k=11}^{21} \binom{21}{k}(0.3)^k(0.7)^{21-k} \approx 0.026.$$A 30 % individual error becomes 2.6 %.
Exercise 3 — AdaBoost weight update by hand#
After round $t$ , learner $h_t$ has $\epsilon_t = 0.2$ . Sample $i$ is correctly classified with current weight $w_t(i) = 0.05$ . Compute (i) the learner weight $\alpha_t$ and (ii) the unnormalised next weight $w_{t+1}(i)$ .
Solution. $\alpha_t = \tfrac{1}{2}\ln(0.8/0.2) = \tfrac{1}{2}\ln 4 \approx 0.693$ . Correctly classified, so $w_{t+1}(i) = 0.05 \cdot e^{-0.693} = 0.025$ before normalisation. Misclassified samples would be multiplied by $e^{0.693} = 2$ .
Exercise 4 — AdaBoost vs GBDT cheat sheet#
| Aspect | AdaBoost | GBDT |
|---|---|---|
| Loss | Exponential (fixed) | Any differentiable |
| Base learner | Decision stumps | CART regression trees |
| Update | Reweight samples | Fit negative gradient |
| Noise robustness | Poor (exp. penalty) | Tunable (Huber etc.) |
| Native regression | No | Yes |
Bottom line: AdaBoost is a beautiful special case; GBDT is the framework you want for production work.
Exercise 5 — Why feature randomisation matters#
A Random Forest with $m = d$ (i.e. consider all features at every split) is just bagging. Explain why this typically performs worse than a forest with smaller $m$ , even though each individual tree is stronger.
Solution. With $m = d$ every tree greedily picks the same dominant features at the top of the tree. The pairwise correlation $\rho$ stays large, so the variance floor $\rho\sigma^2$ in §1.1 is high. Reducing $m$ slightly weakens each tree (lower $s$ ) but dramatically lowers $\rho$ , and the bound $\bar\rho(1 - s^2)/s^2$ tightens. The forest as a whole generalises better.
What’s next#
The algebraic skeleton of Bagging and Boosting ends here. What actually wins on tabular data in industry, though, are two engineering culminations of the Boosting line — XGBoost and LightGBM. The next chapter reads them as “Boosting + system optimization” rather than as two new algorithms.
XGBoost’s second-order Taylor expansion turns “find the optimal leaf weight” into a closed form, and the regularization term enters the split-gain formula directly; LightGBM’s histograms, GOSS sampling, and leaf-wise growth shave training time by an order of magnitude. The math under both libraries is not deep — the core idea is still “add one tree that maximizes second-order loss decrease” — but they push that simple idea to the systems, memory, and concurrency limit. The gap between “the math of gradient boosting” and “XGBoost in production” is not new theory; it is having polished every line of the loop.
References#
- Breiman, L. (1996). Bagging predictors. Machine Learning, 24(2), 123–140.
- Breiman, L. (2001). Random forests. Machine Learning, 45(1), 5–32.
- Freund, Y., & Schapire, R. E. (1997). A decision-theoretic generalization of on-line learning and an application to boosting. Journal of Computer and System Sciences, 55(1), 119–139.
- Friedman, J. H. (2001). Greedy function approximation: A gradient boosting machine. Annals of Statistics, 29(5), 1189–1232.
- Friedman, J. H. (2002). Stochastic gradient boosting. Computational Statistics & Data Analysis, 38(4), 367–378.
- Schapire, R. E., Freund, Y., Bartlett, P., & Lee, W. S. (1998). Boosting the margin: A new explanation for the effectiveness of voting methods. Annals of Statistics, 26(5), 1651–1686.
- Hastie, T., Tibshirani, R., & Friedman, J. (2009). The Elements of Statistical Learning (2nd ed.). Springer. Chapter 10 .
ML Math Derivations 20 parts
- 01 ML Math Derivations (1): Introduction and Mathematical Foundations
- 02 ML Math Derivations (2): Linear Algebra and Matrix Theory
- 03 ML Math Derivations (3): Probability Theory and Statistical Inference
- 04 ML Math Derivations (4): Convex Optimization Theory
- 05 ML Math Derivations (5): Linear Regression
- 06 ML Math Derivations (6): Logistic Regression and Classification
- 07 ML Math Derivations (7): Decision Trees
- 08 ML Math Derivations (8): Support Vector Machines
- 09 ML Math Derivations (9): Naive Bayes
- 10 ML Math Derivations (10): Semi-Naive Bayes and Bayesian Networks
- 11 ML Math Derivations (11): Ensemble Learning you are here
- 12 ML Math Derivations (12): XGBoost and LightGBM
- 13 ML Math Derivations (13): EM Algorithm and GMM
- 14 ML Math Derivations (14): Variational Inference and Variational EM
- 15 ML Math Derivations (15): Hidden Markov Models
- 16 ML Math Derivations (16): Conditional Random Fields
- 17 ML Math Derivations (17): Dimensionality Reduction and PCA
- 18 ML Math Derivations (18): Clustering Algorithms
- 19 ML Math Derivations (19): Neural Networks and Backpropagation
- 20 ML Math Derivations (20): Regularization and Model Selection