
ML Math Derivations (13): EM Algorithm and GMM
Derive the EM algorithm from Jensen's inequality and the ELBO, prove its monotone-ascent guarantee, and apply it to Gaussian Mixture Models with full E-step / M-step formulas, model selection via BIC/AIC, and the K-means correspondence.
When data has hidden structure — like an unobserved cluster label, a missing feature, or an unseen topic — maximum likelihood becomes challenging. The log of a sum has no closed form, and gradient methods get entangled with the latent variables. The EM algorithm sidesteps the difficulty with a deceptively simple idea: alternate between guessing the hidden variables under a posterior (E-step) and fitting the parameters as if those guesses were true (M-step). Each iteration is mathematically guaranteed to push the likelihood up. This post derives EM from first principles, proves the monotone-ascent property using Jensen’s inequality, and explores its most famous application: Gaussian Mixture Models (GMM) — the soft, elliptical generalization of K-means.

What You Will Learn#
- Why latent variables make MLE hard (the log-of-a-sum problem)
- How Jensen’s inequality builds the Evidence Lower Bound (ELBO)
- The EM algorithm as alternating maximisation of the ELBO in $(q, \boldsymbol{\theta})$
- A clean proof that $\ell(\boldsymbol{\theta}^{(t+1)}) \geq \ell(\boldsymbol{\theta}^{(t)})$
- Complete E-step / M-step formulas for GMM with full covariance
- How K-means is the hard, spherical limit of GMM
- Picking $K$ with BIC and AIC
Prerequisites#
- Maximum likelihood estimation
- Multivariate Gaussian density
- Jensen’s inequality and KL divergence
- K-means clustering
Latent variables and incomplete-data likelihood#
Setup#
$$ \ell(\boldsymbol{\theta}) \;=\; \sum_{i=1}^{N} \log p(\mathbf{x}_i \mid \boldsymbol{\theta}) \;=\; \sum_{i=1}^{N} \log \sum_{z} p(\mathbf{x}_i, z \mid \boldsymbol{\theta}). $$The summation inside the logarithm is the source of the problem. The log no longer factors over components, so the gradient doesn’t split into per-component pieces, and there is no closed-form maximizer.
The mixture example#
$$p(\mathbf{x}\mid \boldsymbol{\theta}) \;=\; \sum_{k=1}^{K} \pi_k\, \mathcal{N}(\mathbf{x}\mid \boldsymbol{\mu}_k, \boldsymbol{\Sigma}_k).$$If we knew which component each point came from, fitting would reduce to $K$ independent weighted-Gaussian MLEs — trivial. We do not know, and that is exactly what EM patches up.
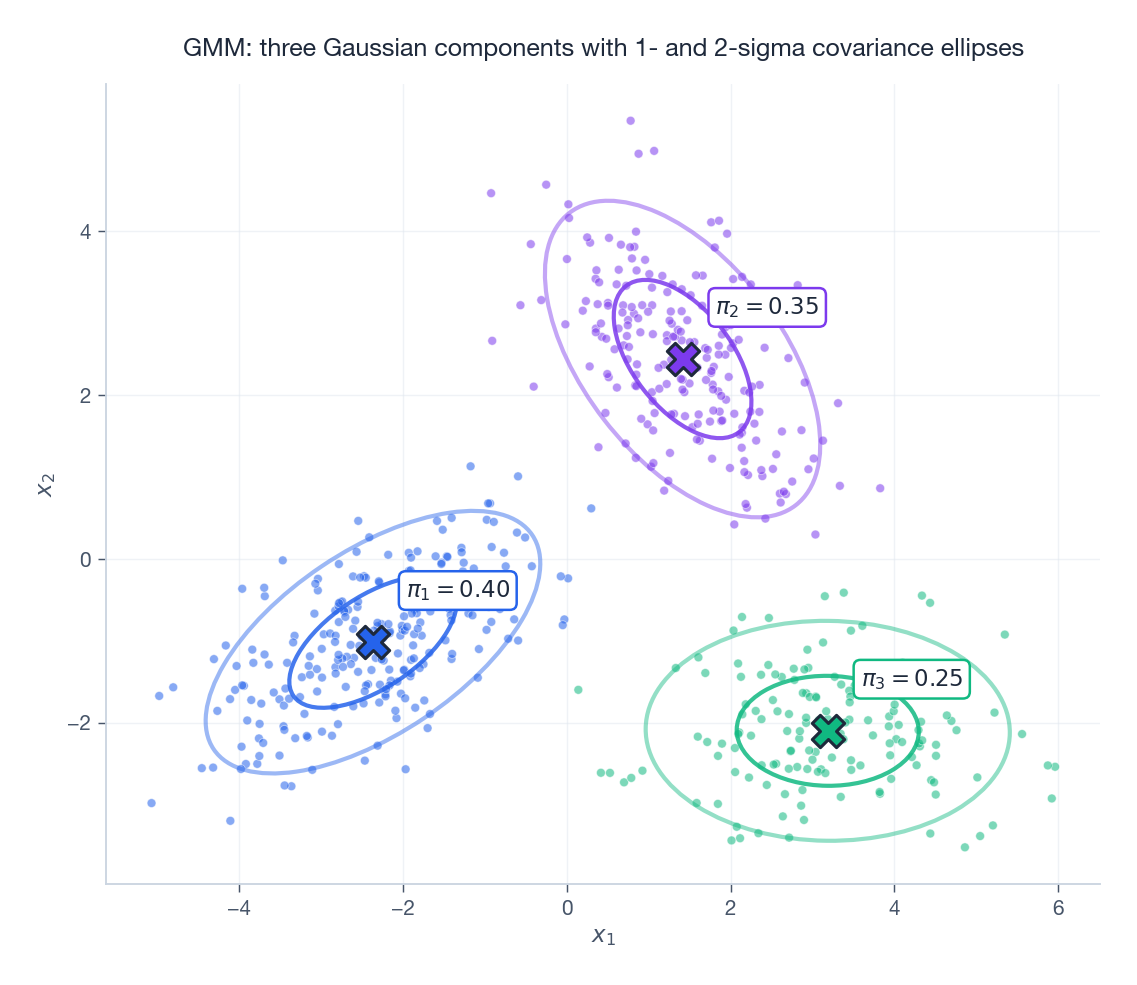
The figure above shows three Gaussian clusters fit by sklearn.mixture.GaussianMixture: each cross is a mean $\boldsymbol{\mu}_k$
, the inner ellipse is the 1-sigma contour, the outer is 2-sigma, and $\pi_k$
is the mixing weight.
The ELBO and Jensen’s inequality#
Introducing an auxiliary distribution $q$ #
$$ \log p(\mathbf{x}\mid \boldsymbol{\theta}) = \log \sum_{z} q(z)\, \frac{p(\mathbf{x}, z\mid \boldsymbol{\theta})}{q(z)}. $$ $$ \boxed{\; \log p(\mathbf{x}\mid \boldsymbol{\theta}) \;\geq\; \sum_{z} q(z)\, \log \frac{p(\mathbf{x}, z\mid \boldsymbol{\theta})}{q(z)} \;\equiv\; \mathcal{L}(q,\boldsymbol{\theta}). \;} $$This $\mathcal{L}$ is the Evidence Lower Bound (ELBO). It depends on both the variational distribution $q$ and the parameters $\boldsymbol{\theta}$ .
The exact decomposition#
$$ \log p(\mathbf{x}\mid \boldsymbol{\theta}) \;=\; \mathcal{L}(q,\boldsymbol{\theta}) \;+\; \mathrm{KL}\bigl[q(z)\,\Vert\, p(z\mid \mathbf{x},\boldsymbol{\theta})\bigr]. $$Two consequences are immediate:
- The ELBO is always $\leq \log p(\mathbf{x}\mid\boldsymbol{\theta})$ because $\mathrm{KL}\geq 0$ .
- The bound becomes tight, $\mathcal{L} = \log p$ , iff $q(z) = p(z \mid \mathbf{x}, \boldsymbol{\theta})$ — the posterior.
This single identity is the entire engine of EM.
EM as coordinate ascent on the ELBO#

EM repeatedly raises $\mathcal{L}$ by alternating in its two arguments.
The two steps#
$$q^{(t)}(z) \;=\; p\bigl(z \mid \mathbf{x}, \boldsymbol{\theta}^{(t)}\bigr).$$After this step the bound is tight: $\mathcal{L}(q^{(t)},\boldsymbol{\theta}^{(t)}) = \log p(\mathbf{x}\mid \boldsymbol{\theta}^{(t)})$ .
$$ Q(\boldsymbol{\theta}\mid \boldsymbol{\theta}^{(t)}) \;=\; \mathbb{E}_{z\sim q^{(t)}}\!\bigl[\log p(\mathbf{x}, z\mid \boldsymbol{\theta})\bigr]. $$The monotone-ascent proof#
$$ \log p(\mathbf{x}\mid \boldsymbol{\theta}^{(t)}) \;\overset{(a)}{=}\; \mathcal{L}(q^{(t)},\boldsymbol{\theta}^{(t)}) \;\overset{(b)}{\leq}\; \mathcal{L}(q^{(t)},\boldsymbol{\theta}^{(t+1)}) \;\overset{(c)}{\leq}\; \log p(\mathbf{x}\mid \boldsymbol{\theta}^{(t+1)}). $$ $$\boxed{\;\ell(\boldsymbol{\theta}^{(t+1)}) \;\geq\; \ell(\boldsymbol{\theta}^{(t)})\;}$$at every iteration, with equality only at fixed points. EM converges to a stationary point of $\ell$ — typically a local maximum, occasionally a saddle point. It is not guaranteed to reach the global maximum, which is why multiple random restarts matter.
Visualising the two views#
The ELBO view makes the dynamics very concrete. After every E-step the KL gap closes; the M-step then raises both the log-likelihood and the ELBO together, and the gap re-opens until the next E-step.
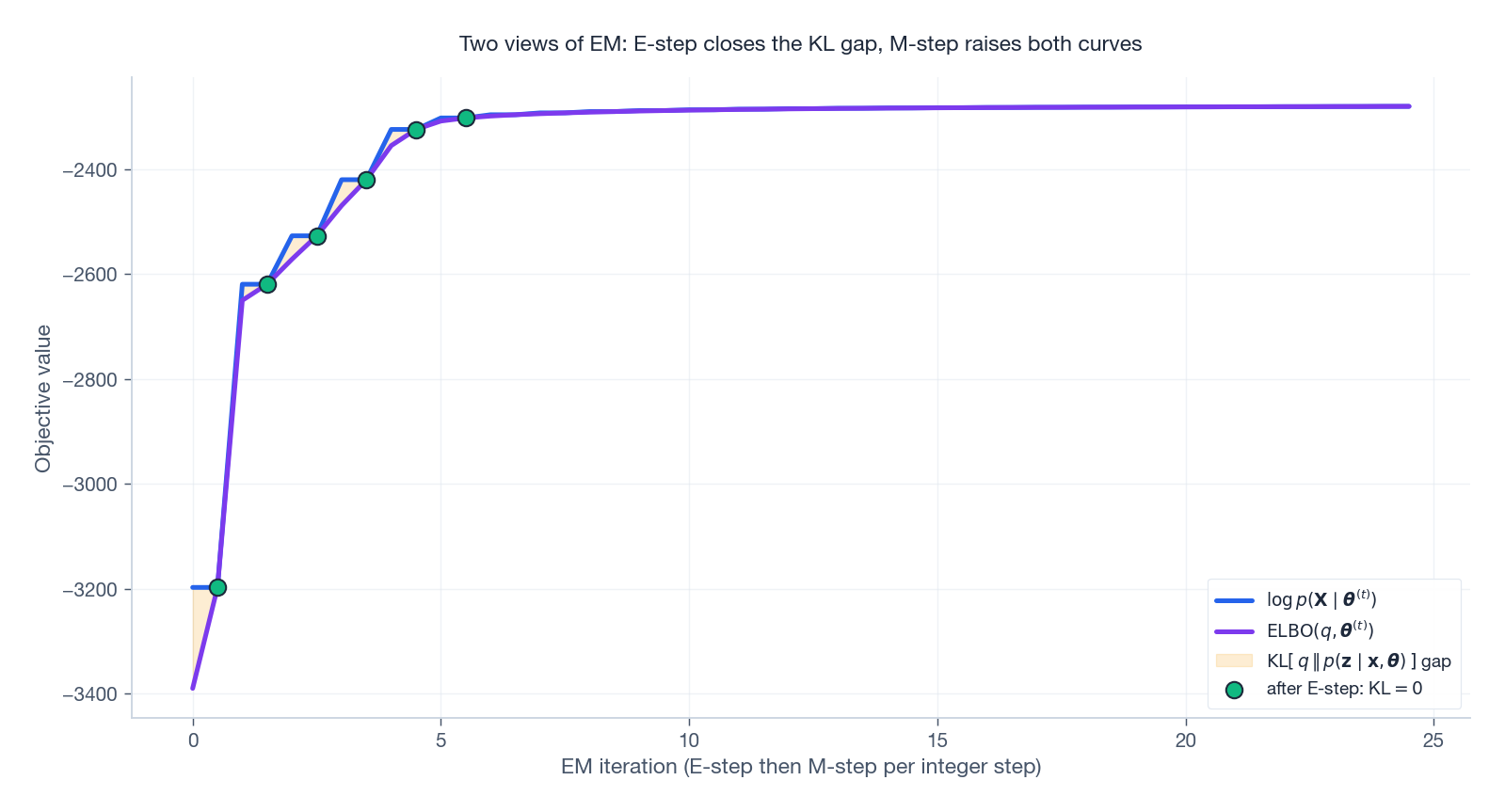
The shaded amber band is exactly the KL divergence $\mathrm{KL}\bigl[q\,\Vert\, p(z\mid\mathbf{x},\boldsymbol{\theta})\bigr]$ . Green dots mark the post-E-step instants where the gap is zero by construction.
EM for Gaussian Mixture Models#
The model#
Generative process for one observation:
- Sample a component label $z_i \sim \mathrm{Categorical}(\pi_1,\dots,\pi_K)$ .
- Sample $\mathbf{x}_i \mid z_i = k \;\sim\; \mathcal{N}(\boldsymbol{\mu}_k, \boldsymbol{\Sigma}_k)$ .
The parameters are $\boldsymbol{\theta} = \{\pi_k, \boldsymbol{\mu}_k, \boldsymbol{\Sigma}_k\}_{k=1}^{K}$ , with $\sum_k \pi_k = 1$ and each $\boldsymbol{\Sigma}_k \succ 0$ .
The E-step: responsibilities#
$$ \boxed{\; \gamma_{ik} \;=\; p\bigl(z_i = k \mid \mathbf{x}_i, \boldsymbol{\theta}^{(t)}\bigr) \;=\; \frac{\pi_k\,\mathcal{N}(\mathbf{x}_i\mid \boldsymbol{\mu}_k, \boldsymbol{\Sigma}_k)}{\sum_{j=1}^{K} \pi_j\,\mathcal{N}(\mathbf{x}_i\mid \boldsymbol{\mu}_j, \boldsymbol{\Sigma}_j)}. \;} $$Each row $(\gamma_{i1},\dots,\gamma_{iK})$ sums to 1 — the soft cluster membership.
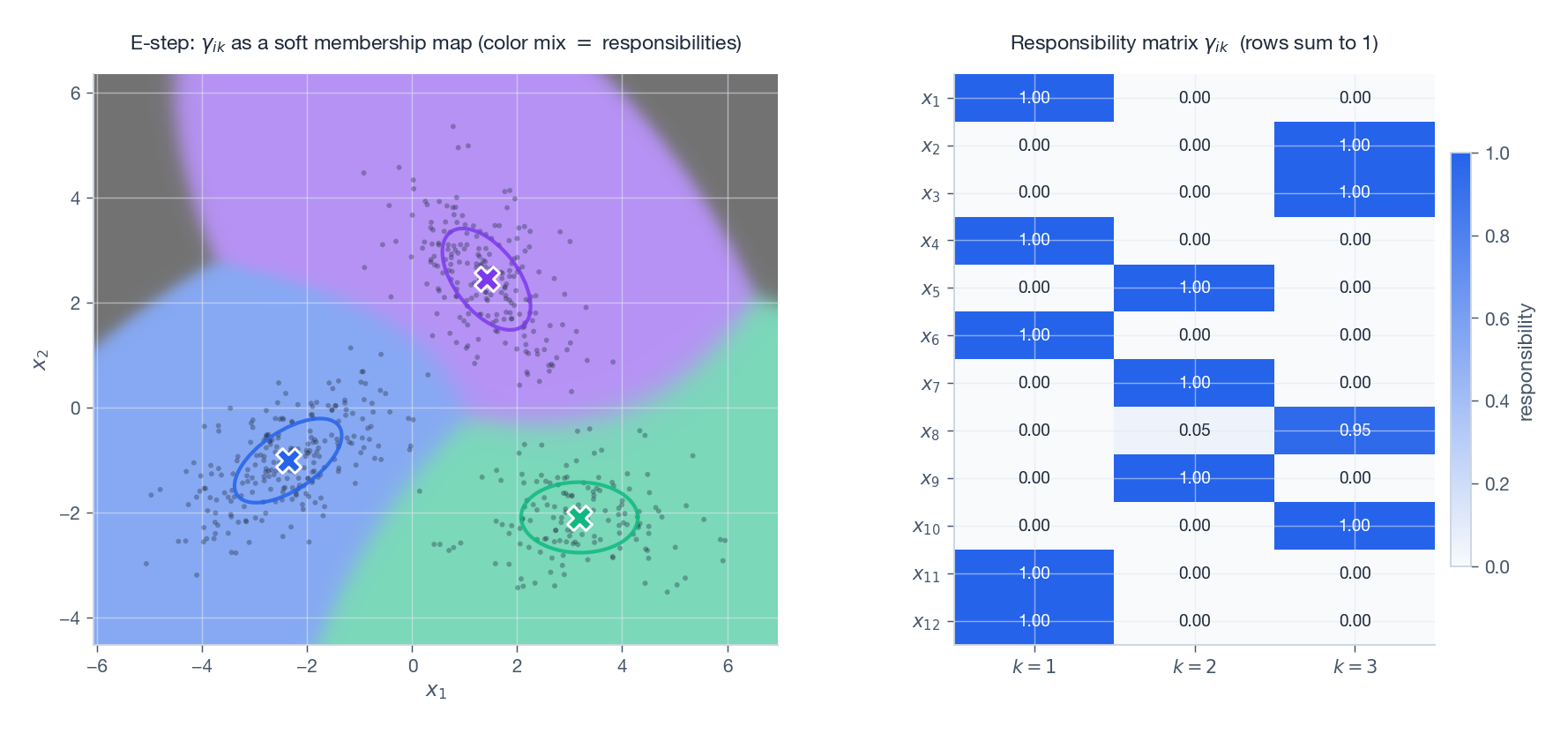
On the left every grid point is coloured by mixing the three component colours according to $\gamma_{ik}$ : pure colour where one component dominates, blended colours along the boundaries. On the right, the responsibility matrix $\gamma_{ik}$ for twelve sample points — rows sum to 1.
The M-step: weighted MLE#
$$ \boxed{\; \pi_k = \frac{N_k}{N},\qquad \boldsymbol{\mu}_k = \frac{1}{N_k}\sum_{i=1}^{N} \gamma_{ik}\,\mathbf{x}_i,\qquad \boldsymbol{\Sigma}_k = \frac{1}{N_k}\sum_{i=1}^{N} \gamma_{ik}\,(\mathbf{x}_i - \boldsymbol{\mu}_k)(\mathbf{x}_i - \boldsymbol{\mu}_k)^{\!\top}. \;} $$These are exactly the standard Gaussian MLE formulas, but with each sample re-weighted by its responsibility.
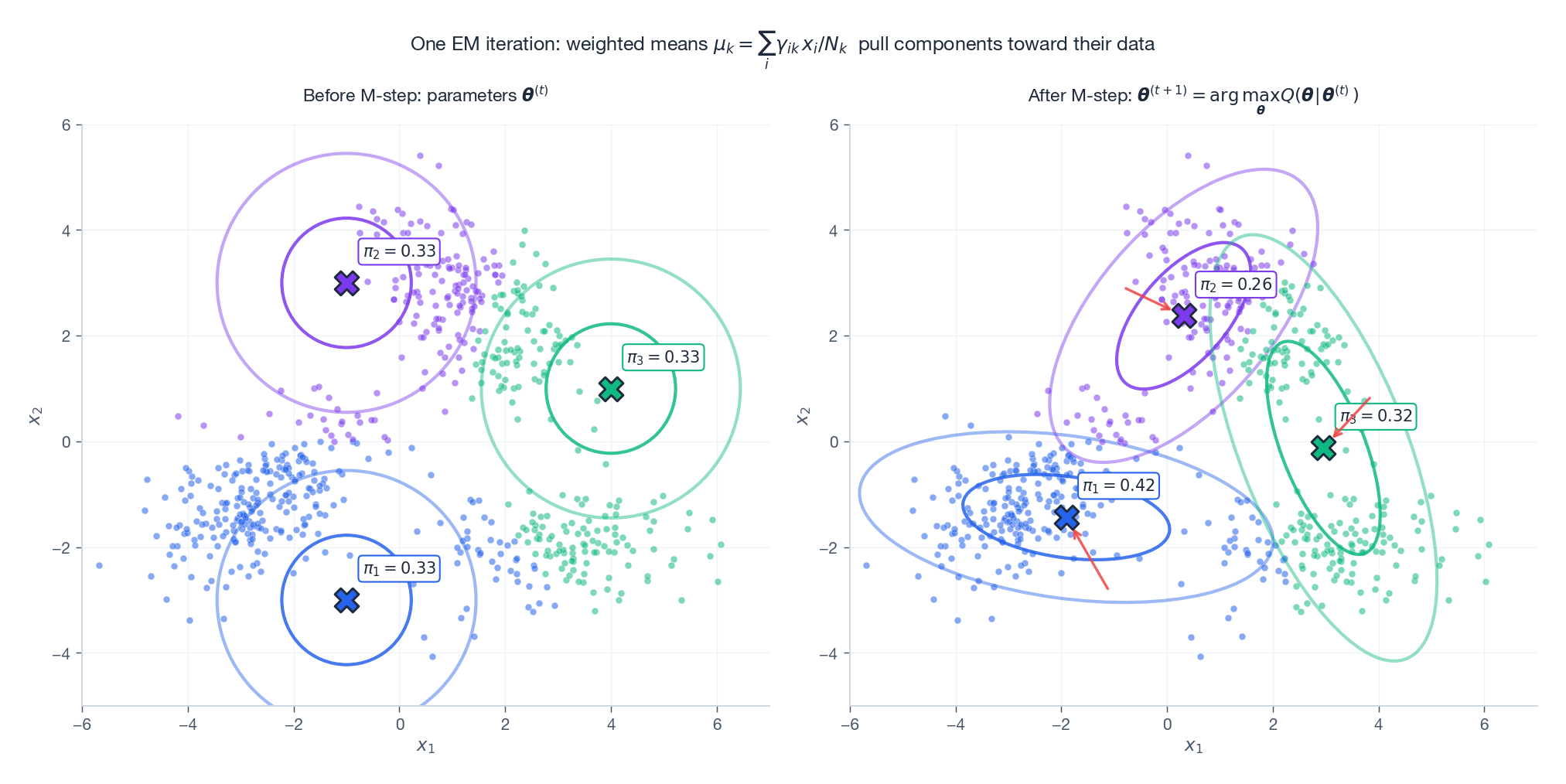
Starting from a deliberately bad initialisation, a single M-step pulls the means (red arrows) onto the data and stretches the covariance ellipses to match the observed scatter. After only a handful of E-M cycles the fit is essentially correct.
Convergence in practice#
Run EM for several random restarts and watch the log-likelihood:
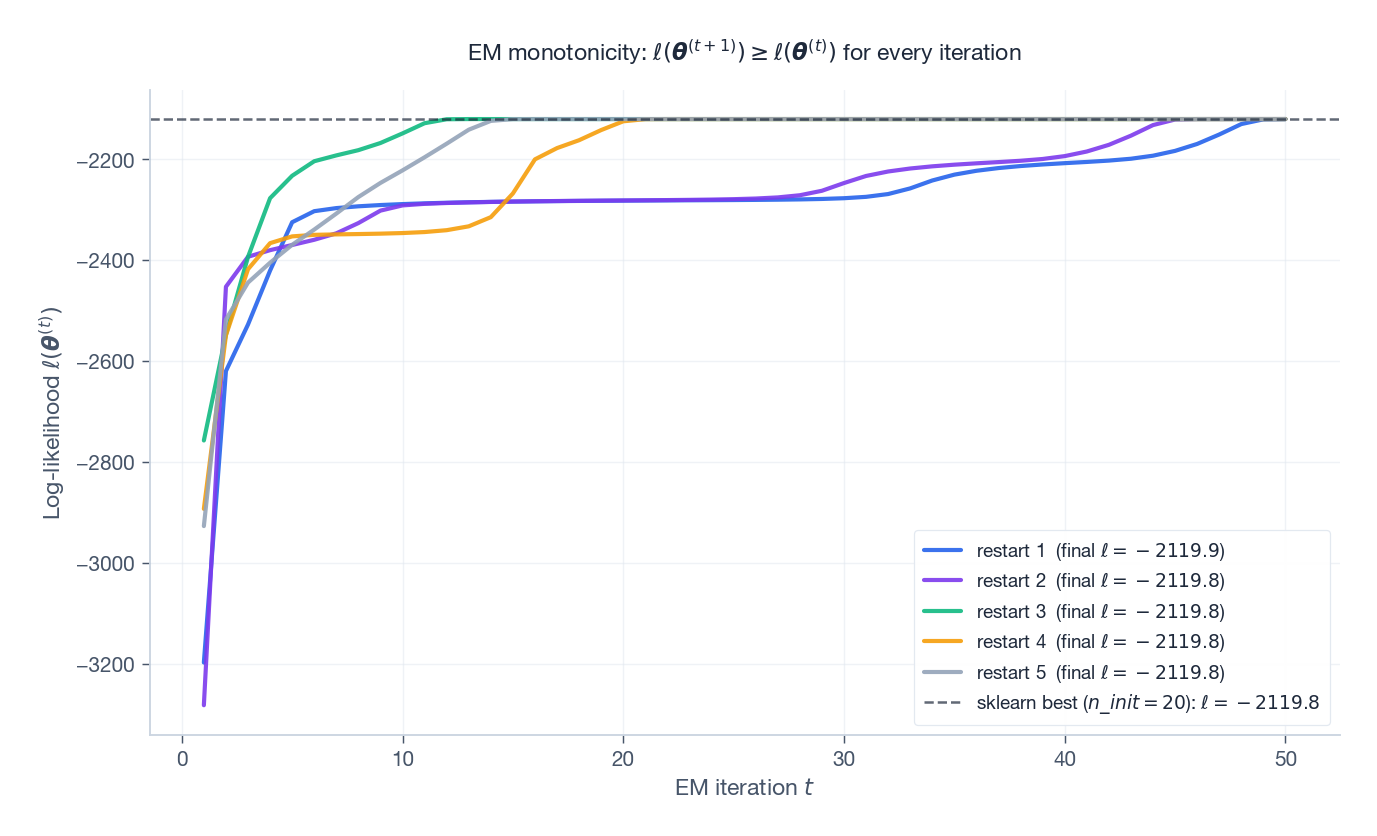
Every restart curve is non-decreasing — this is the algorithmic guarantee. Different restarts plateau at different basins; the dashed line is the best value found by sklearn.mixture.GaussianMixture with n_init=20. Use multiple random or K-means initialisations and keep the best.
K-means is the hard, spherical limit of GMM#
Let $\boldsymbol{\Sigma}_k = \epsilon \mathbf{I}$ for all $k$ and let $\epsilon \to 0$ . The Gaussian density becomes infinitely peaked; the responsibility for the closest mean tends to 1 and the others to 0. The E-step degenerates to hard assignment and the M-step to averaging the assigned points — exactly K-means.
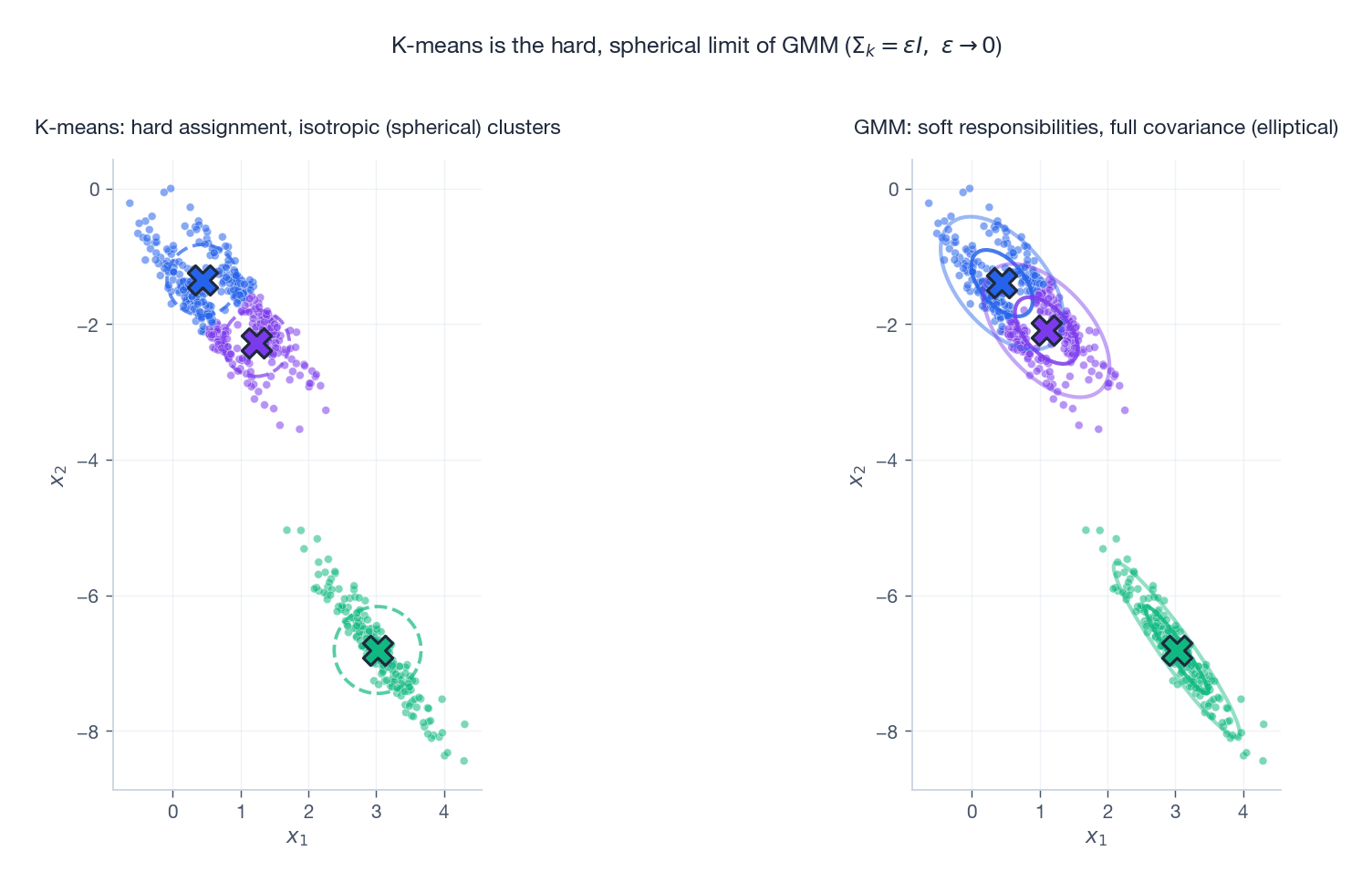
On anisotropic data the difference is stark: K-means (left) imposes spherical Voronoi cells and slices the elongated cluster awkwardly; GMM (right) fits an ellipse along the actual axis of variation. GMM should be your default whenever clusters are not isotropic or you want soft membership probabilities.
Choosing the number of components#
$$ \mathrm{BIC}(K) = -2\,\hat{\ell}(K) + p_K\,\log N, \qquad \mathrm{AIC}(K) = -2\,\hat{\ell}(K) + 2\,p_K, $$where $p_K$ is the parameter count: for full-covariance GMM in $d$ dimensions, $p_K = (K-1) + Kd + K\frac{d(d+1)}{2}$ .
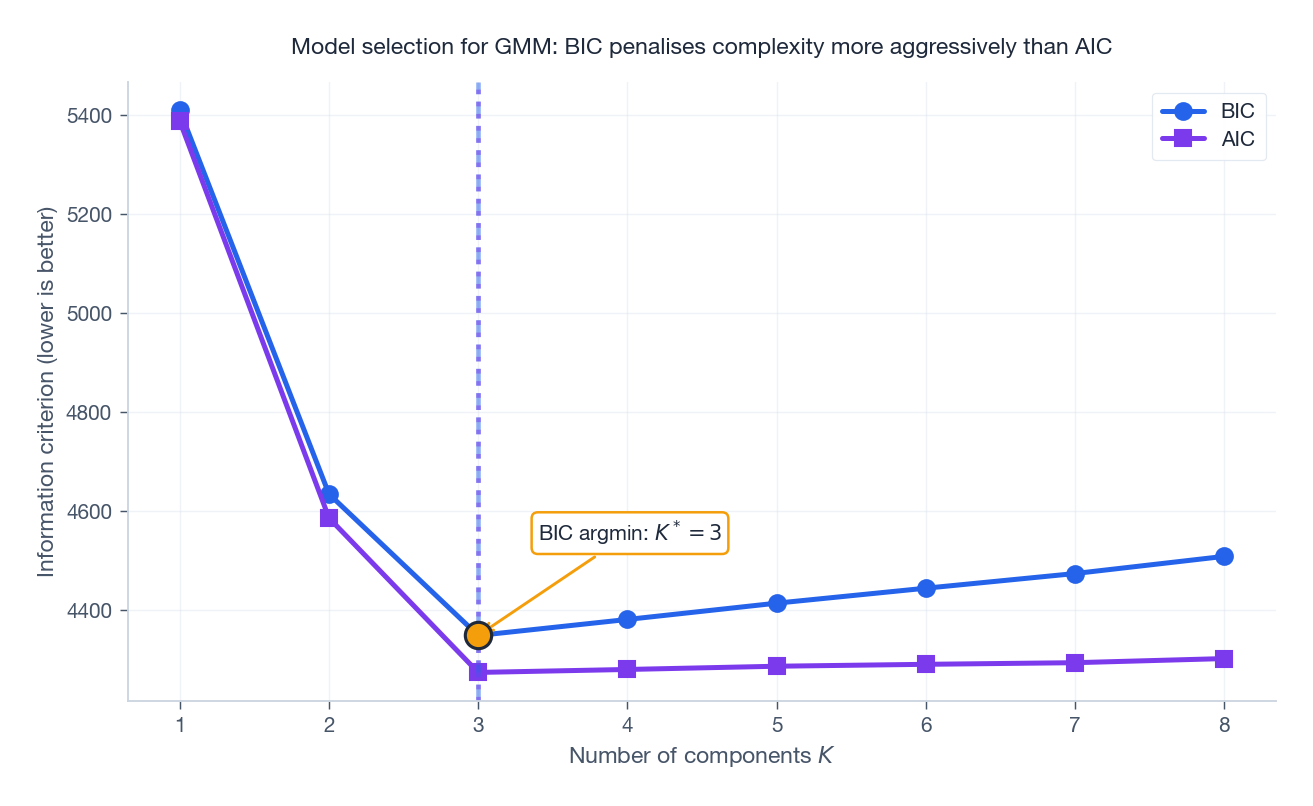
Both curves drop sharply going from $K=1$ to the true $K=3$ and then flatten or rise. BIC penalises complexity more aggressively (an extra $\log N$ factor), so it tends to prefer smaller $K$ than AIC. Either gives the right answer here.
Reference implementation#
A minimal NumPy implementation that mirrors the formulas above. The actual experiments and figures use sklearn.mixture.GaussianMixture for verification.
| |
Numerical considerations#
The EM iteration above is mathematically clean but numerically dangerous. Three failure modes hit me repeatedly in production.
$$\log \gamma_{ik} = \log\pi_k + \log\mathcal{N}_k(\mathbf{x}_i) - \mathrm{logsumexp}_j\big(\log\pi_j + \log\mathcal{N}_j(\mathbf{x}_i)\big),$$then $\gamma_{ik} = \exp(\log \gamma_{ik})$ . Always work in log-space until the very last subtraction.
Singular covariance matrices. When a component captures a single point, $\boldsymbol{\Sigma}_k$ collapses toward the rank-zero matrix and the determinant goes to zero. The likelihood then explodes to $+\infty$ . This is not a bug — it is the correct MLE — but it is useless. The two practical mitigations are (1) add a ridge $\boldsymbol{\Sigma}_k \leftarrow \boldsymbol{\Sigma}_k + \lambda \mathbf{I}$ with $\lambda \approx 10^{-6} \cdot \mathrm{tr}(\boldsymbol{\Sigma}_k)/D$ , and (2) re-initialise any component whose effective count $N_k = \sum_i \gamma_{ik}$ falls below some threshold (I use $N_k < 1$ ).
Conditioning of $\boldsymbol{\Sigma}_k^{-1}$ . When you compute the Mahalanobis distance $(\mathbf{x} - \boldsymbol{\mu})^\top \boldsymbol{\Sigma}^{-1} (\mathbf{x} - \boldsymbol{\mu})$ , never form $\boldsymbol{\Sigma}^{-1}$ explicitly. Take the Cholesky factorisation $\boldsymbol{\Sigma} = \mathbf{L}\mathbf{L}^\top$ and solve $\mathbf{L}\mathbf{y} = \mathbf{x} - \boldsymbol{\mu}$ by forward substitution; the squared distance is $\Vert \mathbf{y}\Vert^2$ , and the log-determinant is $2\sum_d \log L_{dd}$ . This costs $O(D^2)$ per sample instead of $O(D^3)$ and is numerically stable when $\boldsymbol{\Sigma}$ has large condition number.
A useful sanity check during iteration: the ELBO must be monotonically non-decreasing. If you ever see a decrease larger than $10^{-6}$ , your numerics are wrong, not your math.
What this looks like in scikit-learn#
| |
Two flags worth knowing. covariance_type='diag' drops the off-diagonals of $\boldsymbol{\Sigma}_k$
, reducing parameters from $K D(D+1)/2$
to $KD$
and per-iteration cost from $O(NKD^2)$
to $O(NKD)$
. On $D=128$
embeddings this is the difference between minutes and hours. init_params='k-means++' runs K-means first to seed the means; without it, EM on more than four components routinely converges to a degenerate solution where one mixture absorbs everything.
The BIC score $\mathrm{BIC} = -2\log\hat L + p\log N$
(where $p$
is the parameter count) is the cheapest principled $K$
-selection. Sweep $K \in \{1, \dots, K_{\max}\}$
, pick the elbow. Do not trust a single fit — always use n_init >= 5 because EM finds local optima.
FAQ#
Does EM reach the global optimum?#
No. Monotone ascent guarantees only a stationary point — typically a local maximum, sometimes a saddle. Defend yourself with multiple restarts (random or K-means seeded) and keep the run with the highest final log-likelihood.
GMM vs K-means — when does it actually matter?#
Use GMM when (i) clusters are clearly elliptical / anisotropic, (ii) you want soft membership probabilities for downstream calibration, or (iii) you need a generative density model for sampling or anomaly scoring. K-means is faster and fine for roughly spherical, well-separated clusters.
Singular covariance / collapsed component?#
Add a ridge $\boldsymbol{\Sigma}_k + \epsilon \mathbf{I}$ (the reference implementation does this), tie covariances across components, restrict to diagonal covariance, or restart on detection.
Why does the E-step “make the bound tight”?#
Because $\log p = \mathcal{L}(q,\boldsymbol{\theta}) + \mathrm{KL}[q\Vert p(z\mid\mathbf{x},\boldsymbol{\theta})]$ and the KL is zero exactly when $q$ equals the posterior.
What is generalised EM?#
Replace the M-step’s full maximisation with any update that increases $Q$ (e.g. one gradient step). The monotone ascent argument still goes through.
How is EM related to variational inference?#
Variational EM relaxes the E-step from the exact posterior to a tractable family $q \in \mathcal{Q}$ . The decomposition $\log p = \mathcal{L} + \mathrm{KL}$ is identical; the algorithm now alternately minimises KL inside $\mathcal{Q}$ (E) and maximises $\mathcal{L}$ in $\boldsymbol{\theta}$ (M). See Part 14 .
Exercises#
E1 (E-step). Consider a 1D GMM with $K=2$ , equal priors $\pi_1 = \pi_2 = 1/2$ , $\mu_1 = 0,\, \mu_2 = 3,\, \sigma^2 = 1$ . Compute $\gamma_{i1}$ for $x_i = 1.5$ .
Solution. By symmetry the two component densities at $x=1.5$ are equal, so $\gamma_{i1} = 1/2$ .
E2 (M-step). Two samples $x_1 = 1,\; x_2 = 4$ with responsibilities $\gamma_{11} = 0.8,\; \gamma_{21} = 0.3$ . Compute $\mu_1$ after the M-step.
Solution. $N_1 = 0.8 + 0.3 = 1.1$ , so $\mu_1 = (0.8 \cdot 1 + 0.3 \cdot 4) / 1.1 = 2.0 / 1.1 \approx 1.82$ .
E3 (Monotonicity). Where in the chain $\log p(\boldsymbol{\theta}^{(t)}) = \mathcal{L}(q^{(t)},\boldsymbol{\theta}^{(t)}) \leq \mathcal{L}(q^{(t)},\boldsymbol{\theta}^{(t+1)}) \leq \log p(\boldsymbol{\theta}^{(t+1)})$ does the M-step’s optimality enter, and where does the ELBO inequality enter?
Solution. The middle $\leq$ is by the M-step (definition of $\boldsymbol{\theta}^{(t+1)}$ ); the right $\leq$ is the ELBO inequality applied at the new parameters.
E4 (Singular limit). Show that if you fix $\boldsymbol{\Sigma}_k = \epsilon \mathbf{I}$ for all $k$ and let $\epsilon \to 0^+$ , the EM updates for the means converge to the K-means updates.
Sketch. Write $\mathcal{N}(\mathbf{x}\mid\boldsymbol{\mu}_k, \epsilon\mathbf{I}) \propto \exp(-\Vert \mathbf{x} - \boldsymbol{\mu}_k\Vert^2 / (2\epsilon))$ . As $\epsilon \to 0$ the soft-max over $-\Vert\mathbf{x}-\boldsymbol{\mu}_k\Vert^2$ becomes a hard argmin, $\gamma_{ik}\in\{0,1\}$ , and the M-step mean reduces to the cluster-mean update of K-means.
E5 (Missing data). Suppose feature $j$ of $\mathbf{x}_i$ is missing. Outline an EM scheme that treats the missing value as an additional latent variable and updates parameters and missing entries jointly.
Sketch. Add the missing entry $x_{ij}$ to $z$ . The E-step computes $\mathbb{E}[x_{ij} \mid \text{observed},\boldsymbol{\theta}^{(t)}]$ (and second moments). The M-step uses these expectations as plug-ins in the weighted-MLE updates above.
What’s next#
EM is smooth on models like GMM where the E-step has a closed form, but it has a hidden prerequisite: the posterior $p(z\mid x,\theta)$ has to be computable exactly. As soon as the model gets slightly more complicated — LDA, deep Bayesian networks, Bayesian neural nets — that posterior becomes intractable. The next chapter swaps in a far more general tool: variational inference.
The VI idea is: if the true posterior cannot be computed, pick the closest member $q^\star$ from a tractable family $\mathcal{Q}$ and turn inference into optimization. ELBO is the workhorse objective, mean field is the most common family, CAVI is the most direct solver. Reread EM as “E-step with the true posterior, M-step over $\theta$ ”; variational EM then becomes “E-step with an approximate posterior, M-step over $\theta$ ” — the single most important step between classical EM and modern deep generative models, of which the VAE is just amortised variational EM.
References#
- Dempster, A. P., Laird, N. M., & Rubin, D. B. (1977). Maximum likelihood from incomplete data via the EM algorithm. Journal of the Royal Statistical Society: Series B, 39(1), 1-22.
- Bishop, C. M. (2006). Pattern Recognition and Machine Learning. Springer. Chapter 9.
- Murphy, K. P. (2012). Machine Learning: A Probabilistic Perspective. MIT Press. Chapter 11.
- McLachlan, G., & Krishnan, T. (2007). The EM Algorithm and Extensions (2nd ed.). Wiley.
- Neal, R. M., & Hinton, G. E. (1998). A view of the EM algorithm that justifies incremental, sparse, and other variants. Learning in Graphical Models, 89, 355-368.
ML Math Derivations 20 parts
- 01 ML Math Derivations (1): Introduction and Mathematical Foundations
- 02 ML Math Derivations (2): Linear Algebra and Matrix Theory
- 03 ML Math Derivations (3): Probability Theory and Statistical Inference
- 04 ML Math Derivations (4): Convex Optimization Theory
- 05 ML Math Derivations (5): Linear Regression
- 06 ML Math Derivations (6): Logistic Regression and Classification
- 07 ML Math Derivations (7): Decision Trees
- 08 ML Math Derivations (8): Support Vector Machines
- 09 ML Math Derivations (9): Naive Bayes
- 10 ML Math Derivations (10): Semi-Naive Bayes and Bayesian Networks
- 11 ML Math Derivations (11): Ensemble Learning
- 12 ML Math Derivations (12): XGBoost and LightGBM
- 13 ML Math Derivations (13): EM Algorithm and GMM you are here
- 14 ML Math Derivations (14): Variational Inference and Variational EM
- 15 ML Math Derivations (15): Hidden Markov Models
- 16 ML Math Derivations (16): Conditional Random Fields
- 17 ML Math Derivations (17): Dimensionality Reduction and PCA
- 18 ML Math Derivations (18): Clustering Algorithms
- 19 ML Math Derivations (19): Neural Networks and Backpropagation
- 20 ML Math Derivations (20): Regularization and Model Selection