
ML Math Derivations (16): Conditional Random Fields
Why do CRFs outperform HMMs on sequence labeling? This article derives linear-chain CRF from the ground up -- potential functions, the forward-backward algorithm, gradient computation, and Viterbi decoding.

What You Will Learn#
Named entity recognition, POS tagging, information extraction — every one of these tasks asks you to label each element of a sequence. HMMs (Part 15 ) attack this problem generatively by modelling the joint distribution $P(\mathbf{X},\mathbf{Y})$ , but to make the joint factorise they pay a steep price: each observation is assumed independent of everything except its own hidden label. In real text, whether bank is a noun or a verb depends on the preceding and following words, the suffix, capitalization, and dictionary lookups — all these features together.
Conditional Random Fields (CRFs) drop the generative ambition entirely and model $P(\mathbf{Y}\mid\mathbf{X})$ directly. Once you no longer need a generative story for $\mathbf{X}$ , you can pile on as many overlapping features of $\mathbf{X}$ as you like.
What you will learn:
- Why CRF’s discriminative formulation beats HMM’s generative one for labeling tasks
- How transition and state feature functions define CRF’s scoring mechanism
- The forward-backward algorithm — now used to compute $Z(\mathbf{X})$ and marginals
- How the gradient of the log-likelihood reduces to empirical minus expected feature counts
- Viterbi decoding for finding the highest-scoring label sequence
Prerequisites: Probability basics (conditional probability, Bayes’ rule), familiarity with HMMs (Part 15 ), and comfort with matrix notation.
From HMM to CRF: Generative vs Discriminative#
What HMM forces you to assume#
$$P(\mathbf{X}, \mathbf{Y}) = P(y_1) \prod_{t=2}^{T} P(y_t \mid y_{t-1}) \prod_{t=1}^{T} P(x_t \mid y_t)$$To predict labels you go via Bayes’ rule: $\mathbf{Y}^* = \arg\max_{\mathbf{Y}} P(\mathbf{Y}\mid\mathbf{X})$ . Two assumptions make the joint tractable but also restrictive:
- Observation independence: $P(x_t \mid y_{1:T}, x_{\setminus t}) = P(x_t \mid y_t)$ . Each token sees only its own label.
- First-order Markov: $P(y_t \mid y_{1:t-1}) = P(y_t \mid y_{t-1})$ . Each label sees only the previous label.
The Markov assumption is mostly fine. The observation independence assumption, however, is the real issue: it forbids any feature that considers neighboring tokens, suffixes, or gazetteers based on the current label.
The CRF idea: model $P(\mathbf{Y}\mid\mathbf{X})$ directly#
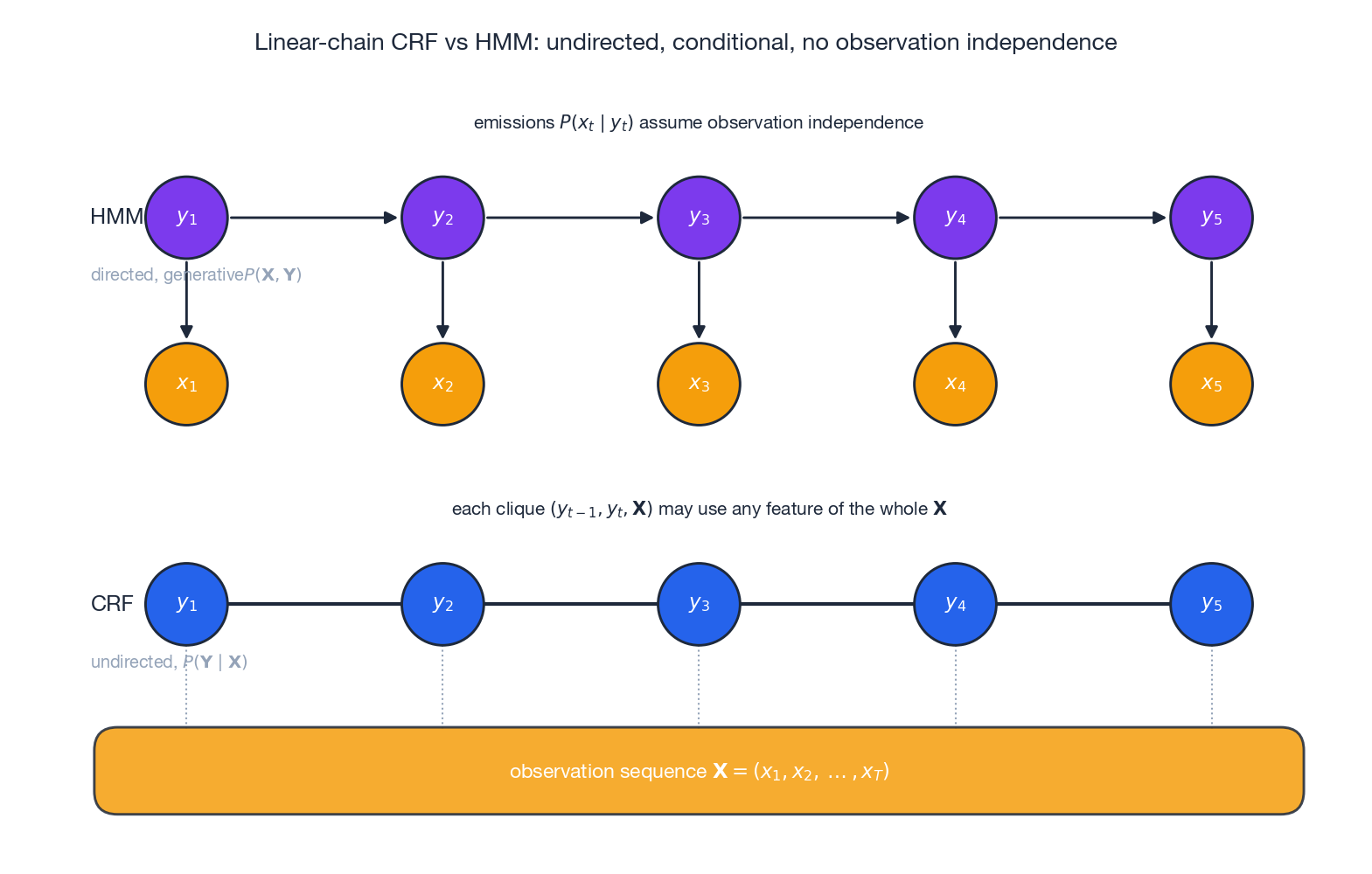
The figure above shows the structural difference. In HMM (top) every $x_t$ is a child of $y_t$ , so the model has to explain the observations — and to keep the joint factorisable each $x_t$ may depend only on $y_t$ . In CRF (bottom) the entire observation sequence $\mathbf{X}$ sits in a shared “context strip”; every clique $(y_{t-1}, y_t, \mathbf{X})$ is allowed to inspect any function of the whole $\mathbf{X}$ . We never write down $P(\mathbf{X})$ , so we never have to assume anything about how observations were generated.
What you gain:
- Arbitrary, overlapping features. Word identity, suffix, prefix, capitalisation, neighbour words, gazetteer matches — all simultaneously, without double counting penalties.
- Flexible feature engineering. Global features are first-class.
- Stronger empirical performance. On standard sequence-labeling benchmarks, CRF typically beats HMM by 5–10 F1 points.
What you pay: the partition function $Z(\mathbf{X})$ is a sum over all $L^T$ label sequences, and you have to compute it (and its gradients) every training iteration. The forward-backward algorithm does exactly this in $O(TL^2)$ .
The path from HMM through MEMM to CRF#
$$P(\mathbf{Y} \mid \mathbf{X}) = \prod_{t=1}^{T} P(y_t \mid y_{t-1}, \mathbf{X})$$MEMM is discriminative and allows arbitrary features of $\mathbf{X}$ , but it normalises locally at each position. This causes the label-bias problem: states with few outgoing transitions concentrate probability mass simply because their per-position softmax has fewer competitors — regardless of whether the observation supports them.
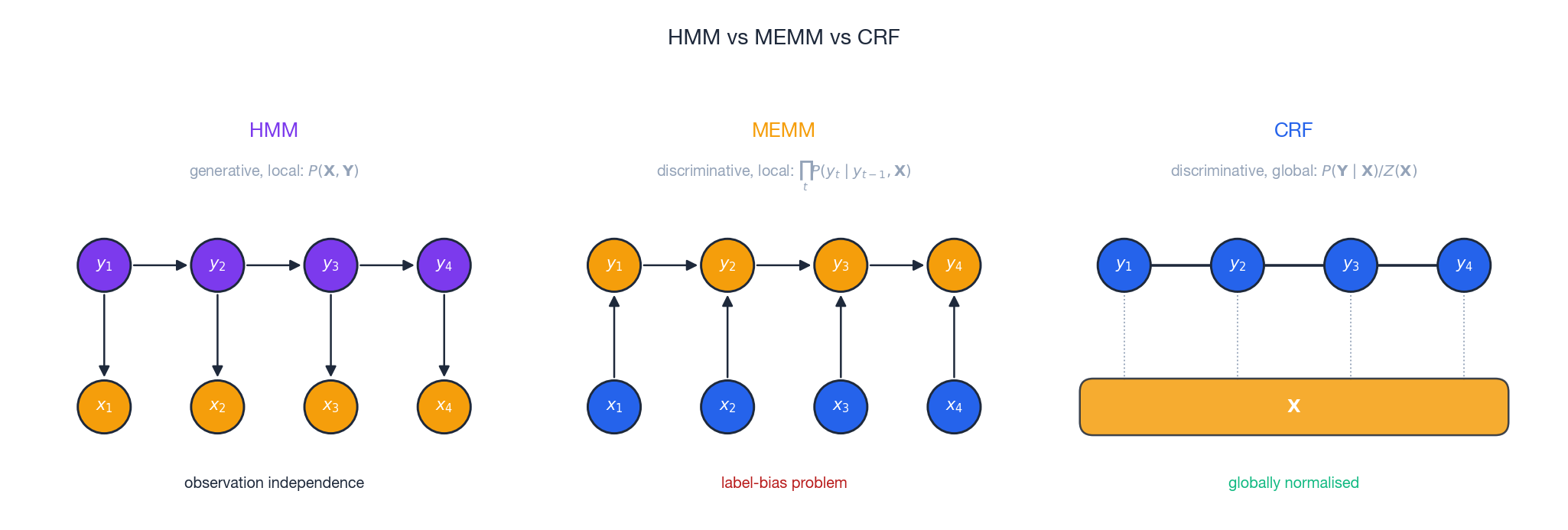
CRF fixes this with global normalisation: the single $Z(\mathbf{X})$ in the denominator forces all paths through the trellis to compete on the same playing field. Bottom line:
- HMM — generative, locally normalised, observation-independent.
- MEMM — discriminative, locally normalised, label-biased.
- CRF — discriminative, globally normalised, no label bias.
Generative vs discriminative more broadly#
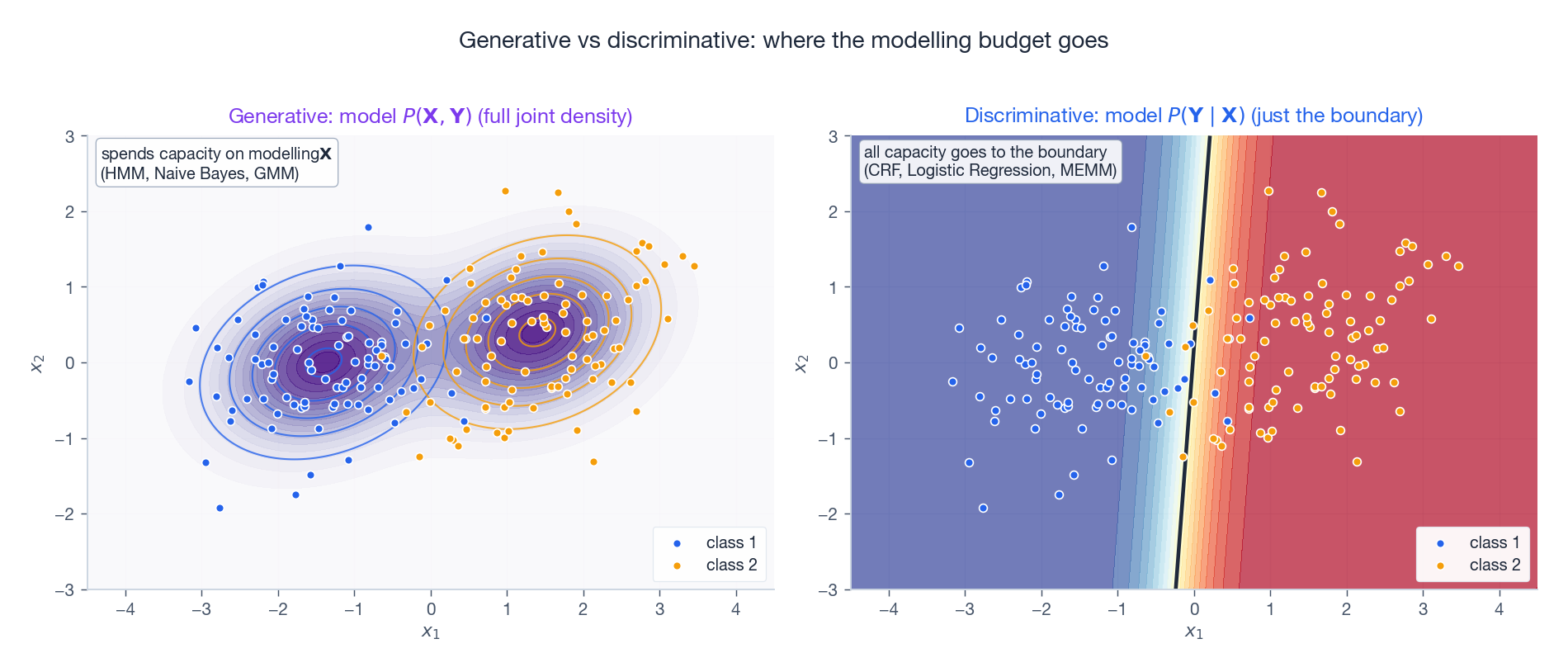
Even outside sequence problems the same trade-off applies. A generative model spends parameters on $P(\mathbf{X})$ — the smooth purple density on the left — which is wasted capacity if all you care about is the boundary. A discriminative model spends every parameter directly on the conditional decision boundary — the sharp diagonal contour on the right. Naive Bayes vs Logistic Regression and HMM vs CRF are exactly the same trade-off lifted to two scales.
Mathematical Framework of Linear-Chain CRF#
Basic definitions#
Input: observation sequence $\mathbf{X} = (x_1, x_2, \dots, x_T)$ . Output: label sequence $\mathbf{Y} = (y_1, y_2, \dots, y_T)$ with each $y_t \in \mathcal{Y} = \{1, 2, \dots, L\}$ .
$$P(\mathbf{Y} \mid \mathbf{X}) = \frac{1}{Z(\mathbf{X})} \prod_{t=1}^{T} \Psi_t(y_{t-1}, y_t, \mathbf{X}) \tag{1}$$where
- $\Psi_t(y_{t-1}, y_t, \mathbf{X}) > 0$ is the potential at position $t$ ,
- $Z(\mathbf{X}) = \sum_{\mathbf{Y}'} \prod_{t} \Psi_t(y'_{t-1}, y'_t, \mathbf{X})$ is the partition function that normalises across all label sequences.
By convention $y_0$
is a special START symbol so the first transition is well defined.
Feature function decomposition#
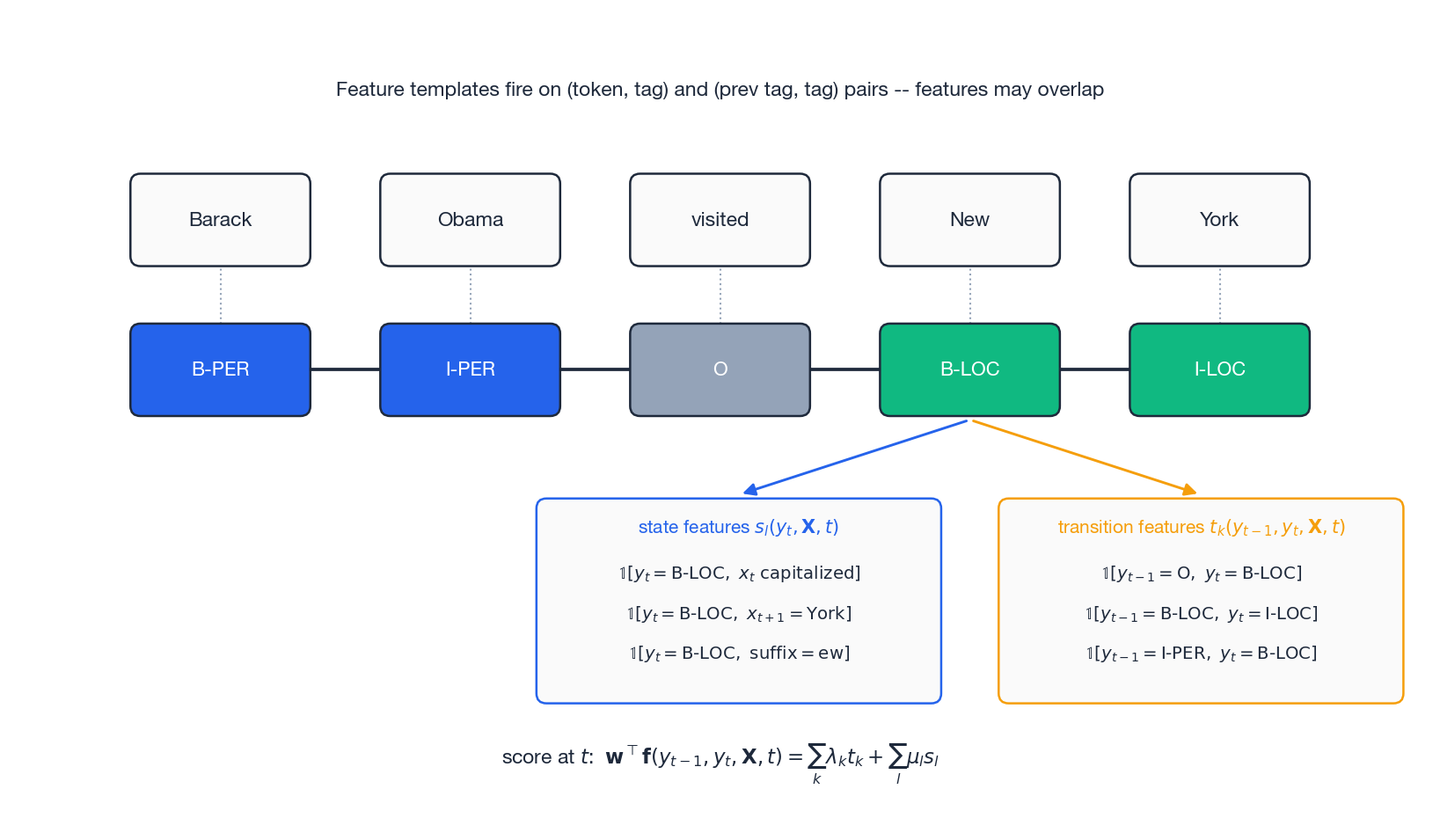
CRF parameterises the potential through two flavours of feature function:
$$t_k(y_{t-1}, y_t, \mathbf{X}, t), \qquad k = 1, \dots, K_1$$Example: $t_1 = \mathbb{1}[y_{t-1}=\text{B-PER},\, y_t=\text{I-PER}]$ — “person name continues”.
$$s_l(y_t, \mathbf{X}, t), \qquad l = 1, \dots, K_2$$Example: $s_1 = \mathbb{1}[y_t=\text{B-LOC},\, x_t \text{ is capitalized}]$ .
The figure shows what this looks like at one position on the sentence “Barack Obama visited New York”. For position $t=4$
(“New” $\to$
B-LOC), the state-feature panel collects every property of $\mathbf{X}$
that fires on label B-LOC at that position (capitalisation, the next word being “York”, the suffix ew), and the transition-feature panel records every (prev tag, current tag) configuration consistent with that decision. Crucially, these features can overlap freely.
Unified parameterisation#
$$\mathbf{f}(y_{t-1}, y_t, \mathbf{X}, t) = (t_1, \dots, t_{K_1}, s_1, \dots, s_{K_2})^\top, \quad \mathbf{w} = (\lambda_1, \dots, \lambda_{K_1}, \mu_1, \dots, \mu_{K_2})^\top$$ $$\Psi_t(y_{t-1}, y_t, \mathbf{X}) = \exp\!\big(\mathbf{w}^\top \mathbf{f}(y_{t-1}, y_t, \mathbf{X}, t)\big) \tag{2}$$ $$\mathbf{F}(\mathbf{Y}, \mathbf{X}) = \sum_{t=1}^{T} \mathbf{f}(y_{t-1}, y_t, \mathbf{X}, t)$$ $$P(\mathbf{Y} \mid \mathbf{X}) = \frac{\exp\!\big(\mathbf{w}^\top \mathbf{F}(\mathbf{Y}, \mathbf{X})\big)}{Z(\mathbf{X})}, \quad Z(\mathbf{X}) = \sum_{\mathbf{Y}'} \exp\!\big(\mathbf{w}^\top \mathbf{F}(\mathbf{Y}', \mathbf{X})\big) \tag{3}$$Matrix form#
$$[\mathbf{M}_t(\mathbf{X})]_{i,j} = \exp\!\big(\mathbf{w}^\top \mathbf{f}(y_{t-1}=i, y_t=j, \mathbf{X}, t)\big)$$ $$Z(\mathbf{X}) = \mathbf{1}^\top \!\left(\prod_{t=1}^{T} \mathbf{M}_t(\mathbf{X})\right)\! \mathbf{1}$$This is exactly $T$ multiplications of $L\times L$ matrices, hence $O(TL^2)$ — the same shape as forward-backward.
Forward-Backward for CRF#
Forward recursion#
Define the forward variable $\alpha_t(j)$ as the unnormalised total score of all partial paths ending in label $j$ at position $t$ .
$$\alpha_1(j) = \Psi_1(y_0 = \text{START},\, y_1 = j,\, \mathbf{X})$$ $$\alpha_t(j) = \sum_{i=1}^{L} \alpha_{t-1}(i) \cdot \Psi_t(y_{t-1}=i,\, y_t=j,\, \mathbf{X}) \tag{4}$$ $$Z(\mathbf{X}) = \sum_{j=1}^{L} \alpha_T(j)$$Intuitively, $\alpha_t(j)$ accumulates the (unnormalised) probability mass of every partial sequence of length $t$ that lands on label $j$ . The recursion just says: to land on $j$ at time $t$ , you must have come from some label $i$ at time $t-1$ .
Complexity: $O(TL^2)$ — at each of $T$ steps you sum over $L \times L$ transitions. Compared to the brute-force $O(L^T)$ , this is the difference between practical and impossible.
Backward recursion#
Symmetrically, $\beta_t(i)$ is the unnormalised total score of all partial paths starting in label $i$ at position $t$ and going to the end. The recursion runs from $t = T$ backwards to $t = 1$ , with the same $O(TL^2)$ cost.
Marginals from $\alpha$ and $\beta$ #
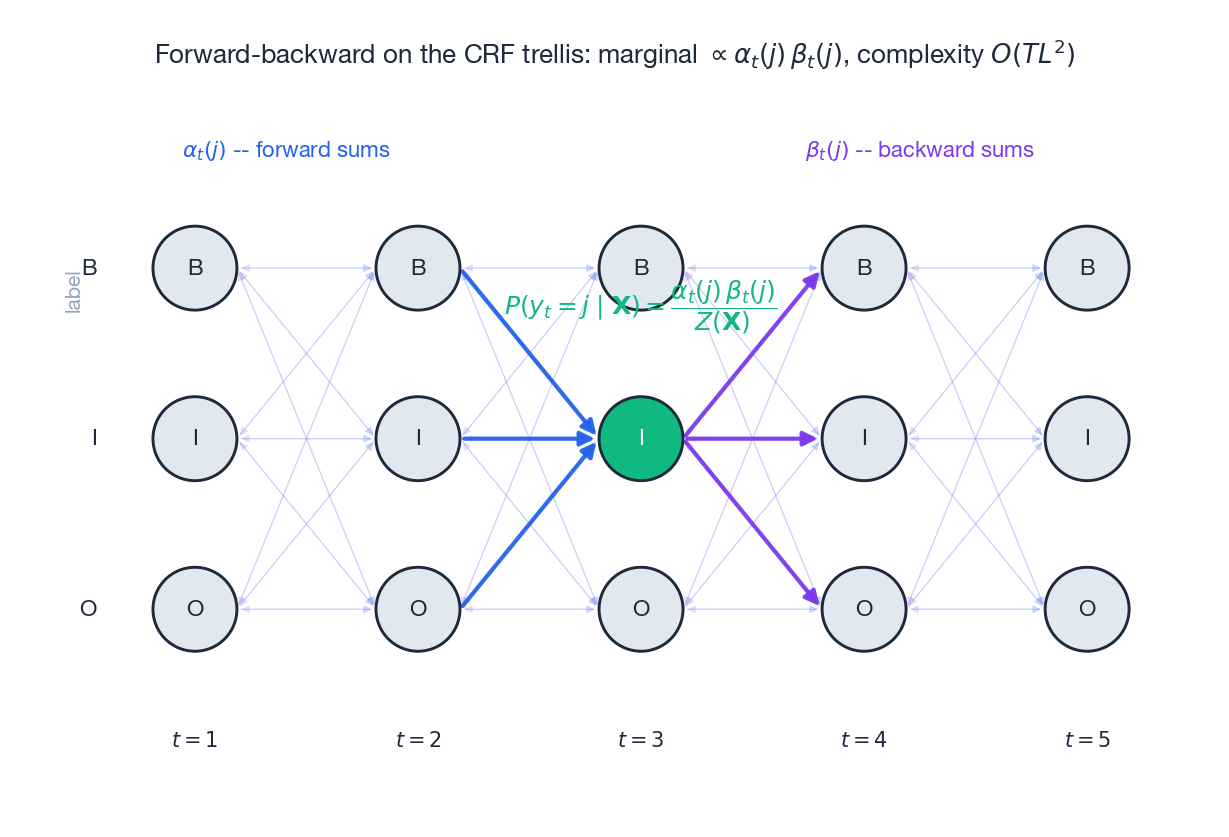
Once you have both passes, marginal probabilities fall out for free:
$$P(y_t = j \mid \mathbf{X}) = \frac{\alpha_t(j) \cdot \beta_t(j)}{Z(\mathbf{X})}$$ $$P(y_{t-1} = i, y_t = j \mid \mathbf{X}) = \frac{\alpha_{t-1}(i) \cdot \Psi_t(i, j, \mathbf{X}) \cdot \beta_t(j)}{Z(\mathbf{X})} \tag{5}$$The trellis figure shows the geometry: every cell $(t, j)$ collects forward arrows from the left (blue) and backward arrows from the right (purple). Their product, normalised by $Z(\mathbf{X})$ , is the per-position marginal that we are about to plug into the gradient.
Numerical stability: log-space#
$$\log\!\sum_i e^{x_i} = \max_i x_i + \log\!\sum_i e^{x_i - \max_i x_i}$$This is the same trick that makes softmax stable.
Parameter Learning: Maximum Likelihood#
Objective#
$$\ell(\mathbf{w}) = \sum_{n=1}^{N} \log P(\mathbf{Y}^{(n)} \mid \mathbf{X}^{(n)}; \mathbf{w}) = \sum_{n=1}^{N}\!\left[\mathbf{w}^\top \mathbf{F}(\mathbf{Y}^{(n)}, \mathbf{X}^{(n)}) - \log Z(\mathbf{X}^{(n)})\right] \tag{6}$$The first term is linear in $\mathbf{w}$ and trivial; all the difficulty is in $\log Z$ .
$$\ell_{\text{reg}}(\mathbf{w}) = \ell(\mathbf{w}) - \tfrac{\lambda}{2} \|\mathbf{w}\|^2$$Gradient: empirical minus expected#
$$\nabla_{\mathbf{w}} \ell = \sum_{n=1}^{N}\!\left[\underbrace{\mathbf{F}(\mathbf{Y}^{(n)}, \mathbf{X}^{(n)})}_{\text{empirical feature counts}} - \underbrace{\mathbb{E}_{P(\mathbf{Y}'\mid \mathbf{X}^{(n)})}\!\big[\mathbf{F}(\mathbf{Y}', \mathbf{X}^{(n)})\big]}_{\text{model-expected feature counts}}\right] \tag{7}$$This is the same shape as the gradient of any maximum-entropy model: how often the feature actually fired in the data, minus how often the current model thinks it should fire. Training pushes the model’s expectations onto the empirical ones; at convergence they match exactly (the maximum-entropy condition).
Computing the expectation in $O(TL^2)$ #
$$\mathbb{E}\big[\mathbf{F}(\mathbf{Y}, \mathbf{X})\big] = \sum_{t=1}^{T} \sum_{i,j} P(y_{t-1}=i, y_t=j \mid \mathbf{X}) \cdot \mathbf{f}(i, j, \mathbf{X}, t) \tag{8}$$The pair-marginals on the right are exactly the ones we computed in (5), at $O(TL^2)$ cost. So one sweep of forward-backward gives us $\log Z$ and every gradient component at once.
Optimisation: L-BFGS#
The objective is concave (and strictly concave once you add L2), so any first-order method converges to the global optimum. In practice L-BFGS is the standard CRF optimiser:
- Quasi-Newton, so it approximates the inverse Hessian and has near-quadratic convergence.
- The “limited-memory” flavour stores only the last $m$ gradient differences — crucial when the feature space has $10^6$ + dimensions.
- Typically converges in 50–200 iterations, where each iteration is one forward-backward pass per training sequence.
scipy.optimize.fmin_l_bfgs_b is the canonical implementation.
Viterbi Decoding#
The decoding problem#
$$\mathbf{Y}^* = \arg\max_{\mathbf{Y}} P(\mathbf{Y} \mid \mathbf{X}) = \arg\max_{\mathbf{Y}} \mathbf{w}^\top \mathbf{F}(\mathbf{Y}, \mathbf{X})$$The denominator $Z(\mathbf{X})$ doesn’t depend on $\mathbf{Y}$ , so we never need to compute it for decoding.
Dynamic programming#
$$\delta_t(j) = \max_{i \in \{1,\dots,L\}}\!\left[\delta_{t-1}(i) + \mathbf{w}^\top \mathbf{f}(i, j, \mathbf{X}, t)\right] \tag{9}$$ $$\psi_t(j) = \arg\max_{i}\!\left[\delta_{t-1}(i) + \mathbf{w}^\top \mathbf{f}(i, j, \mathbf{X}, t)\right]$$so that after reaching position $T$ , $y_T^* = \arg\max_j \delta_T(j)$ , and tracing back $y_{t-1}^* = \psi_t(y_t^*)$ recovers the full best sequence.
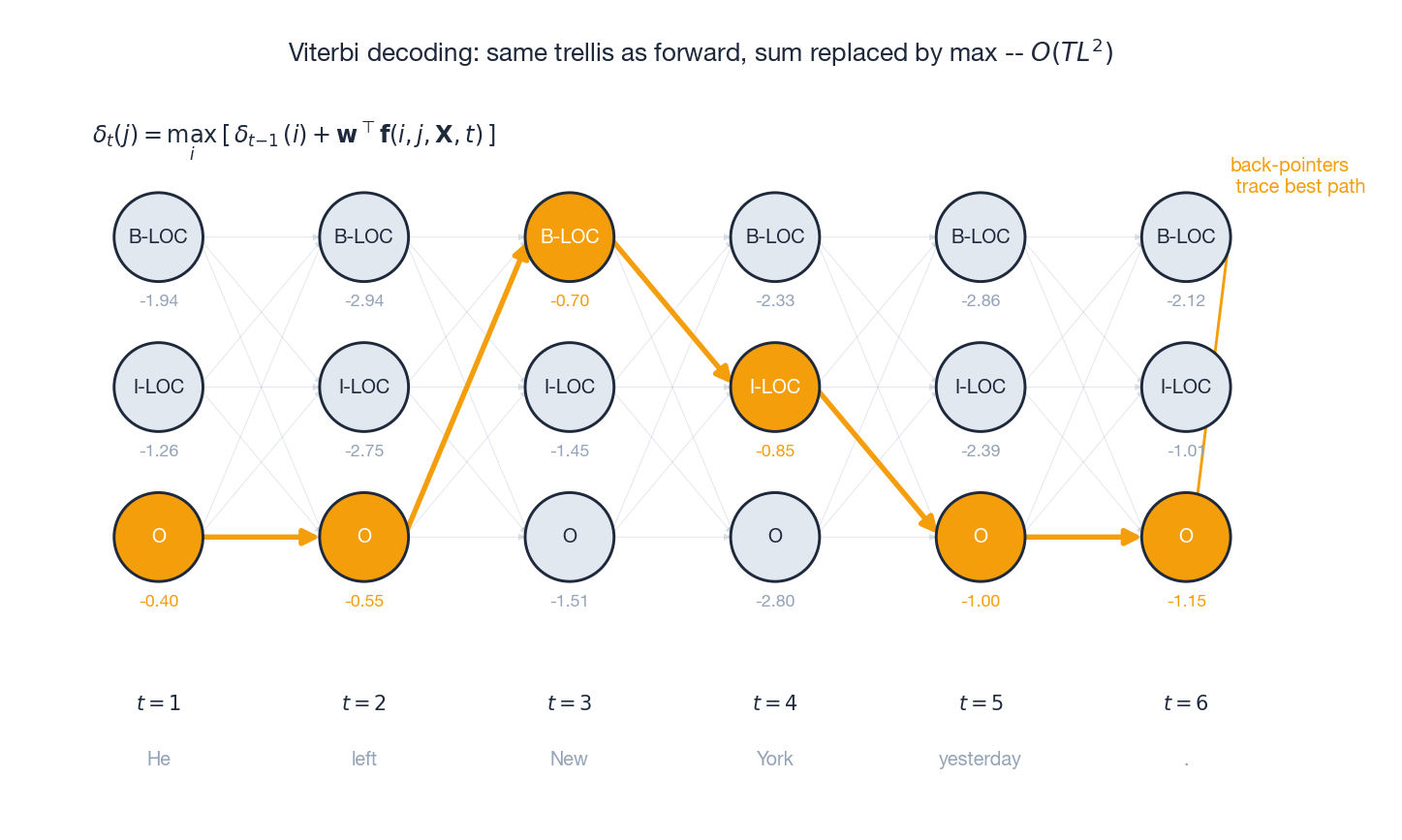
The trellis above shows it visually: faint grey arrows are all candidate transitions; orange arrows mark the surviving max at each step; the highlighted nodes form the back-pointer trace and decode to O O B-LOC I-LOC O O for the toy sentence “He left New York yesterday .”. Same $O(TL^2)$
cost as forward.
CRF in the Deep Learning Era: BiLSTM-CRF#
- BiLSTM (or a Transformer encoder) learns a contextual representation of every token, replacing handcrafted state features with learned ones.
- CRF layer keeps a learnable transition matrix $A_{ij}$ over labels. This matters because constraints like “I-PER must follow B-PER, never O” are sequential and a token-wise softmax cannot enforce them.
- Training jointly optimises both halves end-to-end via the same negative log-likelihood objective.
- Inference still uses Viterbi.
A minimal PyTorch sketch:
| |
The loss is exactly $\log Z(\mathbf{X}) - \text{score}(\mathbf{Y}_{\text{gold}}, \mathbf{X})$ , i.e. the negative of equation (6) for one example. Backpropagation through the forward algorithm computes the same “empirical minus expected” gradient automatically — the chain rule rediscovers (7).
End-to-end NER with confidence#
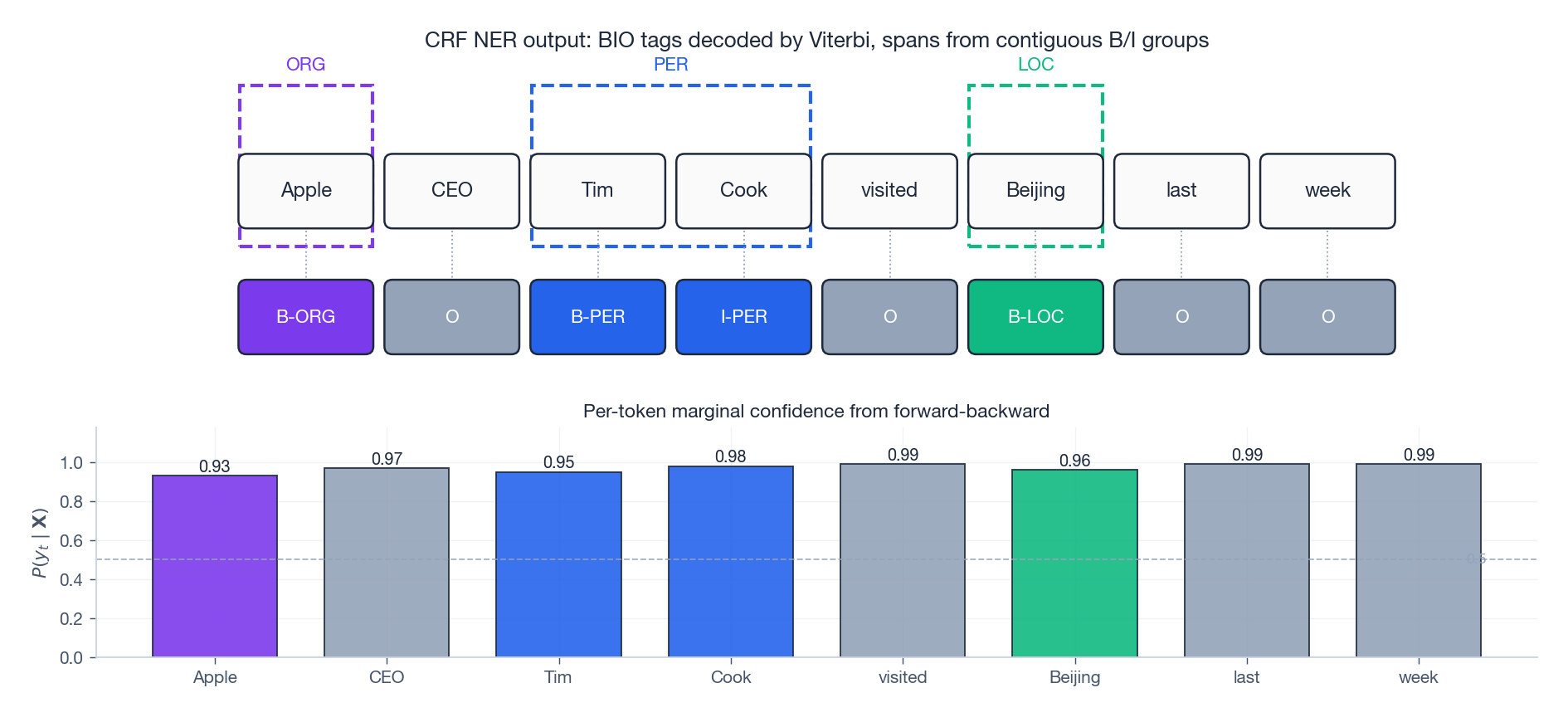
The figure above shows what a trained CRF actually outputs at inference time on the sentence “Apple CEO Tim Cook visited Beijing last week”:
- Top: the Viterbi-decoded BIO tags. Three contiguous B/I groups are decoded as spans (ORG, PER, LOC). The dashed boxes around groups are the entity spans you would emit to a downstream consumer.
- Bottom: the per-token marginal $P(y_t \mid \mathbf{X})$ from forward-backward, which acts as a calibrated confidence score. These marginals are not the same as the decoded path probability; they are computed independently per position and are useful for active learning, abstention, or downstream Bayesian aggregation.
Exercises#
Exercise 1: CRF vs HMM features. In a POS-tagging task, explain why CRF can use the feature “the next word is a verb” but HMM cannot.
Solution: HMM’s observation-independence assumption means $P(x_t \mid y_t)$ may only inspect $x_t$ itself. CRF has no such restriction — its feature functions $\mathbf{f}(y_{t-1}, y_t, \mathbf{X}, t)$ take the entire observation sequence $\mathbf{X}$ as input, so a feature that fires when $x_{t+1}$ is a verb is perfectly legal.
Exercise 2: Designing feature templates. Design three feature functions for named entity recognition.
Solution: (1) Transition: $\mathbb{1}[y_{t-1} = \text{B-ORG},\, y_t = \text{I-ORG}]$ (organisation continuation). (2) State (capitalisation): $\mathbb{1}[y_t = \text{B-LOC},\, x_t \text{ is capitalized}]$ . (3) Context: $\mathbb{1}[y_t = \text{B-PER},\, x_{t-1} = \text{Mr.}]$ (previous word is a personal title).
Exercise 3: Forward-algorithm complexity. A CRF has $T = 100$ and $L = 50$ . How many operations does the forward algorithm need? Compare with brute force.
Solution: At each of $T$ positions you compute $L$ forward values, each summing over $L$ predecessors: $O(TL^2) = O(100 \cdot 50^2) = O(2.5 \times 10^5)$ . Brute force enumerates all paths: $O(L^T) = O(50^{100}) \approx 10^{170}$ . Dynamic programming wins by roughly 165 orders of magnitude.
Exercise 4: Gradient interpretation. State in words what $\nabla_\mathbf{w}\ell = \mathbf{F}_{\text{empirical}} - \mathbb{E}_{\text{model}}[\mathbf{F}]$ means and what holds at convergence.
Solution: If a feature fires more often in the training data than the current model expects, its weight is pushed up; if the model over-predicts a feature, the weight is pushed down. At convergence (gradient = 0), empirical and expected feature counts match exactly. This is the maximum-entropy condition: among all distributions consistent with the empirical feature averages, the CRF picks the one with maximum entropy.
Exercise 5: CRF Viterbi vs HMM Viterbi. Both use Viterbi. What is the only real difference?
Solution: The trellis and the $O(TL^2)$ DP recursion are identical. The difference is the per-edge score: HMM uses local probabilities $\log P(x_t \mid y_t) + \log P(y_t \mid y_{t-1})$ ; CRF uses an inner product $\mathbf{w}^\top \mathbf{f}(y_{t-1}, y_t, \mathbf{X}, t)$ where the feature vector may inspect any function of the whole $\mathbf{X}$ .
Exercise 6: Why label bias bites MEMM but not CRF. Show with a tiny two-state example why local normalisation can ignore the observation, and explain why global normalisation does not.
Solution: Suppose state $A$ has only one outgoing transition (to $B$ ) while state $C$ has 50. After local softmax, the $A \to B$ probability is 1 regardless of the observation — the model has nothing to be uncertain about. CRF’s single denominator $Z(\mathbf{X})$ sums over whole paths, so a high-scoring path through $A$ must out-score all whole paths through $C$ — and observation evidence does enter that comparison.
What’s next#
The full supervised line, sequences included, ends here. From the next chapter on I move to unsupervised representation learning, starting with the most classical dimensionality-reduction method — principal component analysis (PCA).
PCA looks like nothing more than “find the direction of largest variance”, but four equivalent views sit underneath it: maximum-variance projection, minimum reconstruction error, eigendecomposition of the covariance, and SVD of the data matrix. Each view is more natural in different problems, and together they form the semantic skeleton of modern dimensionality reduction — kernel PCA is “PCA in a different inner-product space”, probabilistic PCA is “PCA with Gaussian noise”, t-SNE / UMAP are “PCA with a different similarity metric”. Understanding the four-fold equivalence pays off more than learning any single nonlinear method individually — it is the parent body of the whole family.
References#
[1] Lafferty, J., McCallum, A., & Pereira, F. C. (2001). Conditional random fields: Probabilistic models for segmenting and labeling sequence data. ICML.
[2] Sutton, C., & McCallum, A. (2012). An introduction to conditional random fields. Foundations and Trends in Machine Learning, 4(4), 267-373.
[3] Huang, Z., Xu, W., & Yu, K. (2015). Bidirectional LSTM-CRF models for sequence tagging. arXiv:1508.01991 .
[4] Lample, G., Ballesteros, M., Subramanian, S., Kawakami, K., & Dyer, C. (2016). Neural architectures for named entity recognition. NAACL-HLT.
[5] Ma, X., & Hovy, E. (2016). End-to-end sequence labeling via bi-directional LSTM-CNNs-CRF. ACL.
This is Part 16 of the ML Mathematical Derivations series. Next: Part 17 — Dimensionality Reduction and PCA . Previous: Part 15 — Hidden Markov Models .
ML Math Derivations 20 parts
- 01 ML Math Derivations (1): Introduction and Mathematical Foundations
- 02 ML Math Derivations (2): Linear Algebra and Matrix Theory
- 03 ML Math Derivations (3): Probability Theory and Statistical Inference
- 04 ML Math Derivations (4): Convex Optimization Theory
- 05 ML Math Derivations (5): Linear Regression
- 06 ML Math Derivations (6): Logistic Regression and Classification
- 07 ML Math Derivations (7): Decision Trees
- 08 ML Math Derivations (8): Support Vector Machines
- 09 ML Math Derivations (9): Naive Bayes
- 10 ML Math Derivations (10): Semi-Naive Bayes and Bayesian Networks
- 11 ML Math Derivations (11): Ensemble Learning
- 12 ML Math Derivations (12): XGBoost and LightGBM
- 13 ML Math Derivations (13): EM Algorithm and GMM
- 14 ML Math Derivations (14): Variational Inference and Variational EM
- 15 ML Math Derivations (15): Hidden Markov Models
- 16 ML Math Derivations (16): Conditional Random Fields you are here
- 17 ML Math Derivations (17): Dimensionality Reduction and PCA
- 18 ML Math Derivations (18): Clustering Algorithms
- 19 ML Math Derivations (19): Neural Networks and Backpropagation
- 20 ML Math Derivations (20): Regularization and Model Selection