
ML Math Derivations (17): Dimensionality Reduction and PCA
High-dimensional spaces are hostile to distance-based learning. This article derives PCA from two equivalent angles (max variance and min reconstruction error), and extends to kernel PCA, LDA, t-SNE, and ICA -- with figures that show what each method actually does to the same data.

What You Will Learn#
Feed a clustering algorithm $10{,}000$ -dimensional data and it will most likely fail — not because the algorithm is broken, but because high-dimensional space is a hostile environment for distance-based learning. Volumes evaporate into thin shells, the ratio of nearest- to farthest-neighbour distances tends to $1$ , and “closeness” stops carrying information. Dimensionality reduction is the response: project the data into a lower-dimensional space while keeping the structure that actually matters.
What you will learn:
- Why high-dimensional spaces behave counter-intuitively (the curse of dimensionality)
- PCA derived two equivalent ways: maximum variance and minimum reconstruction error
- How to choose $k$ in practice (scree plot, cumulative variance, reconstruction quality)
- Kernel PCA: PCA in an implicit feature space for nonlinear manifolds
- LDA: supervised dimensionality reduction via Fisher’s criterion
- t-SNE: a probabilistic neighbour-preserving embedding for visualisation
- ICA vs PCA: decorrelation is not independence
Prerequisites: linear algebra (eigenvalues / eigenvectors, the spectral theorem for symmetric matrices), basic probability (variance, covariance), and some familiarity with kernel methods .
The Curse of Dimensionality#
Two phenomena that break intuition#
$$V_d = \frac{\pi^{d/2}}{\Gamma(d/2 + 1)}$$ $$(0.99)^d \xrightarrow{d \to \infty} 0.$$For $d=100$ only about $36.6\%$ of the volume is inside that inner ball; for $d=1000$ it is essentially zero. Most of a high-dimensional ball is “skin”.
$$\frac{\max_{i \ne j}\|\mathbf{x}_i - \mathbf{x}_j\|}{\min_{i \ne j}\|\mathbf{x}_i - \mathbf{x}_j\|} \xrightarrow{d \to \infty} 1.$$When every point is roughly equidistant from every other, $k$ -NN, kernel density, and clustering algorithms lose their grip. This is why preprocessing high-dimensional data through a learned low-dimensional representation is so often the single most useful step in a pipeline.
What dimensionality reduction is for#
- Feature extraction. Strip out redundant directions to denoise, speed up downstream computation, and shrink storage.
- Visualisation. Project to $2$ D or $3$ D so a human can look at the cluster structure.
- Regularisation. Forcing models to live in a lower-dimensional subspace is itself a strong inductive bias.
We can categorise the methods we will study:
| Family | Method | Supervised? | Linear? |
|---|---|---|---|
| Variance | PCA | no | yes |
| Class separation | LDA | yes | yes |
| Independence | ICA | no | yes |
| Implicit feature space | Kernel PCA | no | no |
| Neighbour preservation | t-SNE / UMAP | no | no |
PCA: the Maximum-Variance Derivation#
Setup#
$$\mathbf{S} = \frac{1}{N} \mathbf{X}^\top \mathbf{X} \in \mathbb{R}^{d \times d}.$$$\mathbf{S}$ is symmetric and positive semidefinite, so by the spectral theorem it has an orthonormal eigenbasis with non-negative eigenvalues.
The first principal component#
$$\mathrm{Var}(z) = \frac{1}{N}\sum_{i=1}^{N} (\mathbf{w}_1^\top \mathbf{x}_i)^2 = \mathbf{w}_1^\top \mathbf{S}\, \mathbf{w}_1.$$ $$\mathcal{L}(\mathbf{w}_1, \lambda_1) = \mathbf{w}_1^\top \mathbf{S}\, \mathbf{w}_1 - \lambda_1 (\mathbf{w}_1^\top \mathbf{w}_1 - 1).$$ $$\mathbf{S}\, \mathbf{w}_1 = \lambda_1\, \mathbf{w}_1, \tag{1}$$and the projected variance is exactly $\mathbf{w}_1^\top \mathbf{S}\, \mathbf{w}_1 = \lambda_1$ . To maximise variance we therefore pick the eigenvector associated with the largest eigenvalue of $\mathbf{S}$ .
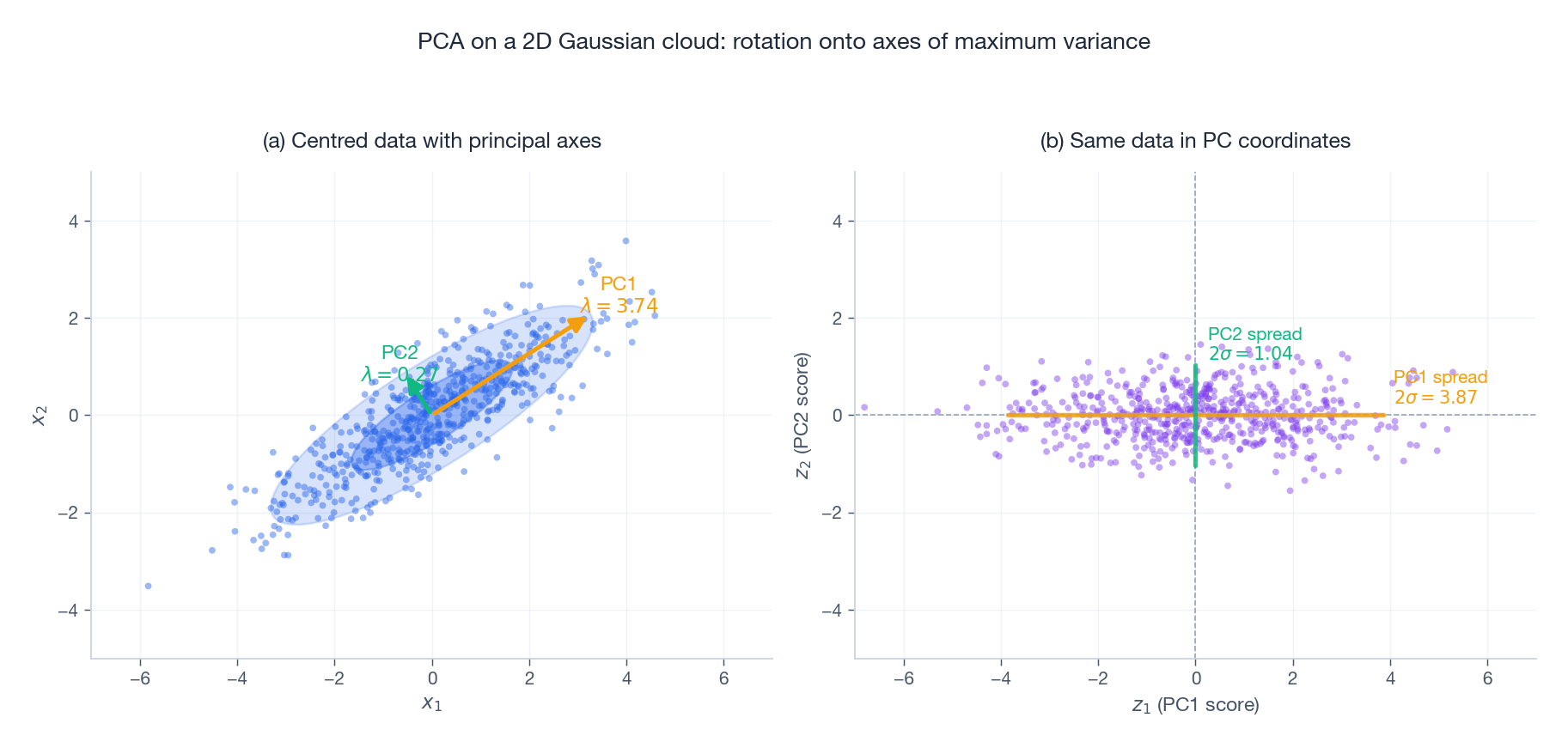
The figure makes the geometry concrete on a correlated $2$ D Gaussian cloud. In panel (a) the orange arrow (PC1) is the direction of maximum spread of the data; the green arrow (PC2) is orthogonal to it and absorbs whatever variance remains. The two ellipses are the $1\sigma$ and $2\sigma$ contours of the empirical Gaussian. Panel (b) shows the same points after the rigid rotation $\mathbf{z}_i = \mathbf{W}^\top \mathbf{x}_i$ : PC1 becomes the horizontal axis, PC2 the vertical axis, and the data is now decorrelated — its empirical covariance is the diagonal matrix $\mathrm{diag}(\lambda_1, \lambda_2)$ . PCA is, geometrically, just the rotation that aligns the coordinate axes with the data’s principal axes.
The full set of principal components#
$$\mathbf{W}_k = [\mathbf{w}_1, \dots, \mathbf{w}_k] \in \mathbb{R}^{d \times k},$$and the reduced representation of $\mathbf{x}_i$ is $\mathbf{z}_i = \mathbf{W}_k^\top \mathbf{x}_i \in \mathbb{R}^k$ .
Choosing $k$ in practice#
Two graphical tools cover almost all use cases.
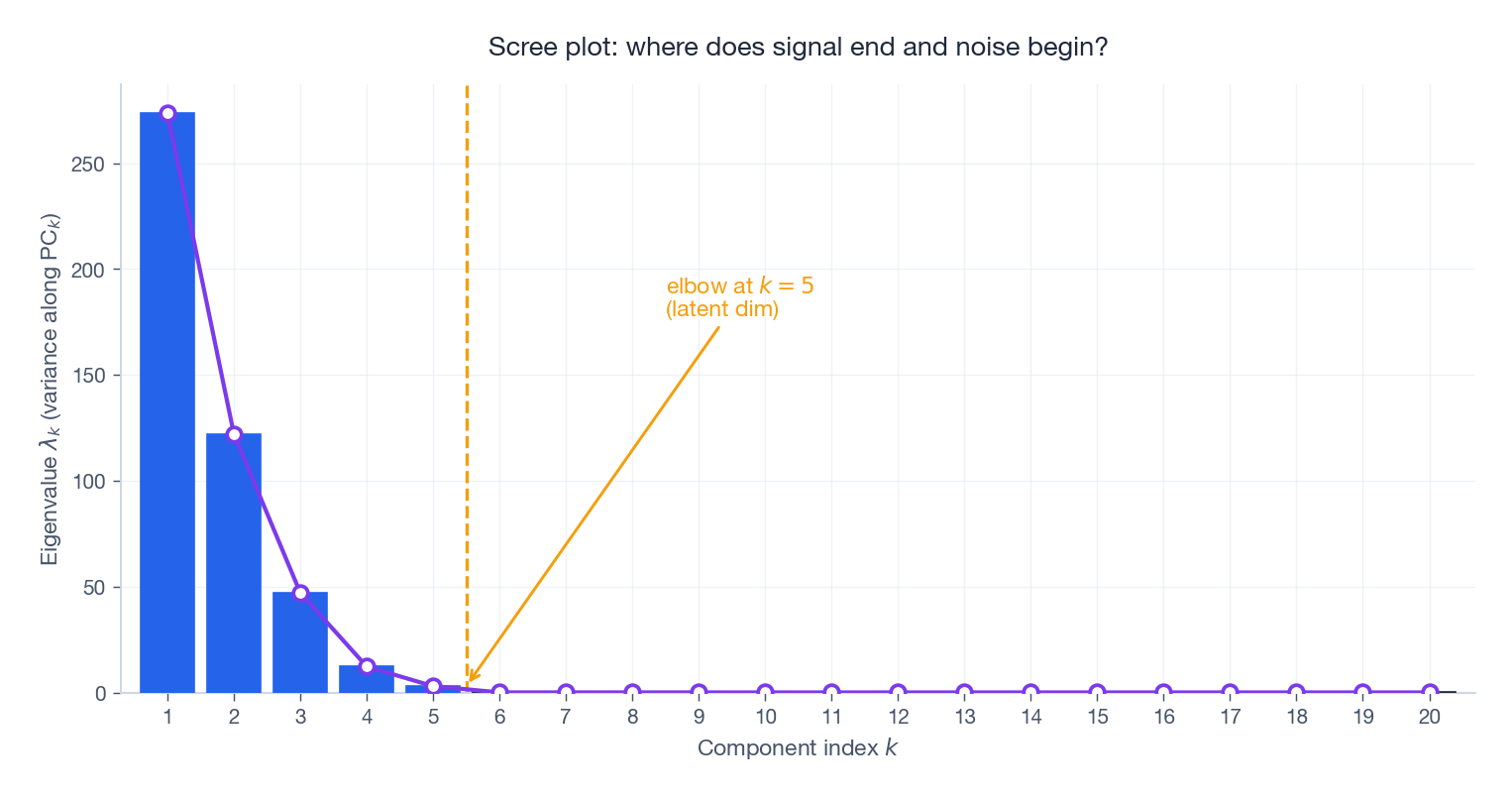
The scree plot shows the eigenvalues themselves. On data generated from a rank-$5$ latent signal plus isotropic noise, the spectrum has a clear elbow at $k=5$ : above it the eigenvalues are signal, below it they are noise. In real datasets the elbow is rarely so clean, but the plot is still the first thing to look at.
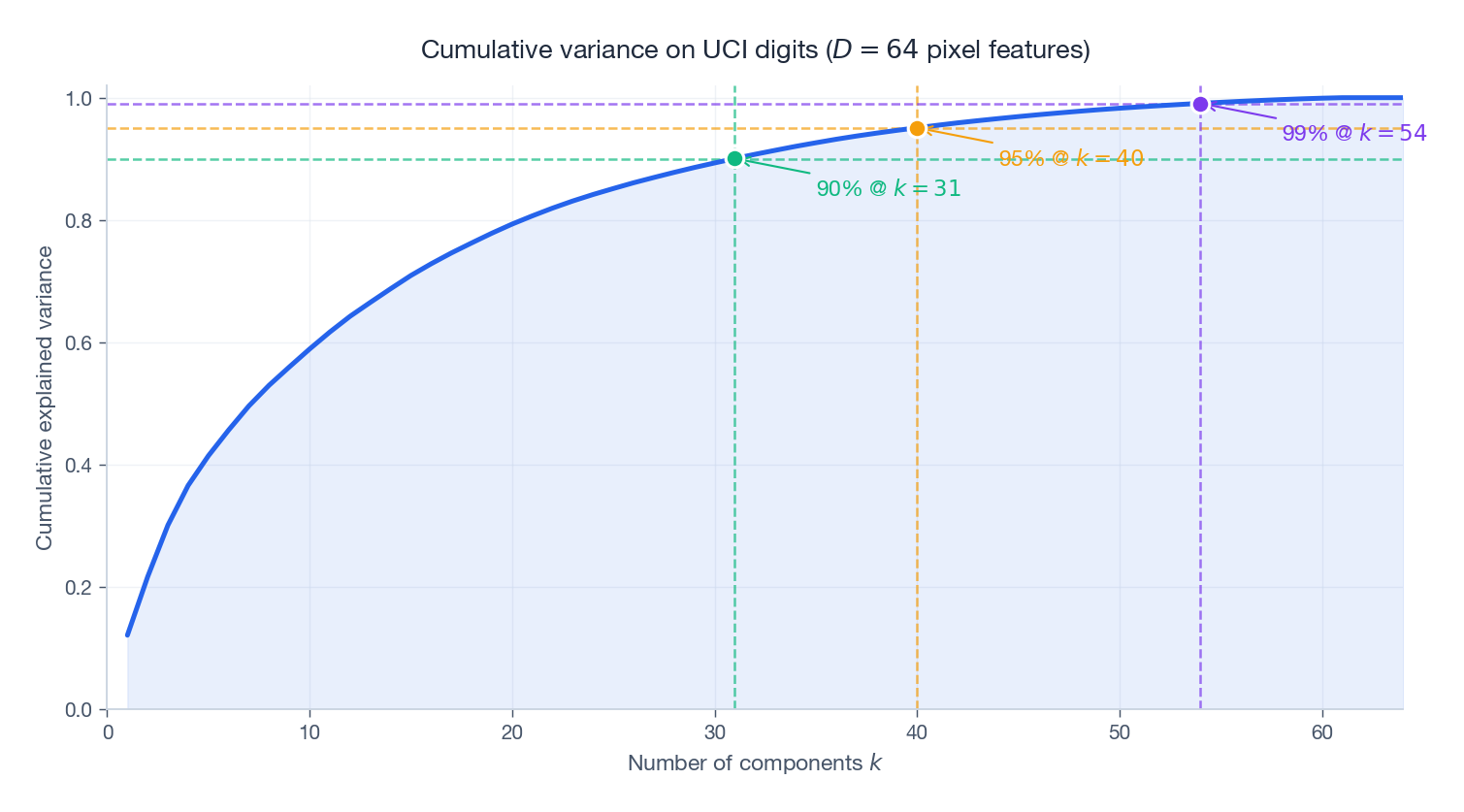
is more decision-friendly: pick the smallest $k$ that crosses your retention target. On the UCI digits dataset (above), $90\%$ retention needs only $k=21$ out of $D=64$ features, $95\%$ needs $k=29$ , and $99\%$ still only needs $k=41$ . Almost two-thirds of the original features are redundant for representing the digit images.
| |
PCA: the Minimum-Reconstruction-Error Derivation#
There is a second, equally natural way to derive PCA. Forget variance for a moment; instead ask: among all rank-$k$ projections, which one best approximates the original data?
$$\hat{\mathbf{x}}_i = \mathbf{W}_k\, \mathbf{W}_k^\top \mathbf{x}_i,$$ $$J_{\mathrm{recon}}(\mathbf{W}_k) = \frac{1}{N} \sum_{i=1}^{N} \big\| \mathbf{x}_i - \mathbf{W}_k \mathbf{W}_k^\top \mathbf{x}_i \big\|^2.$$ $$\underbrace{\mathrm{tr}(\mathbf{S})}_{\text{total variance}} \;=\; \underbrace{\mathrm{tr}(\mathbf{W}_k^\top \mathbf{S}\, \mathbf{W}_k)}_{\text{captured variance}} \;+\; \underbrace{J_{\mathrm{recon}}(\mathbf{W}_k)}_{\text{lost = reconstruction error}}. \tag{3}$$The total is fixed, so maximising captured variance is exactly the same problem as minimising reconstruction error. The two derivations land on the same eigenvalue equation (1), and the optimal $\mathbf{W}_k$ is the same matrix in both cases.
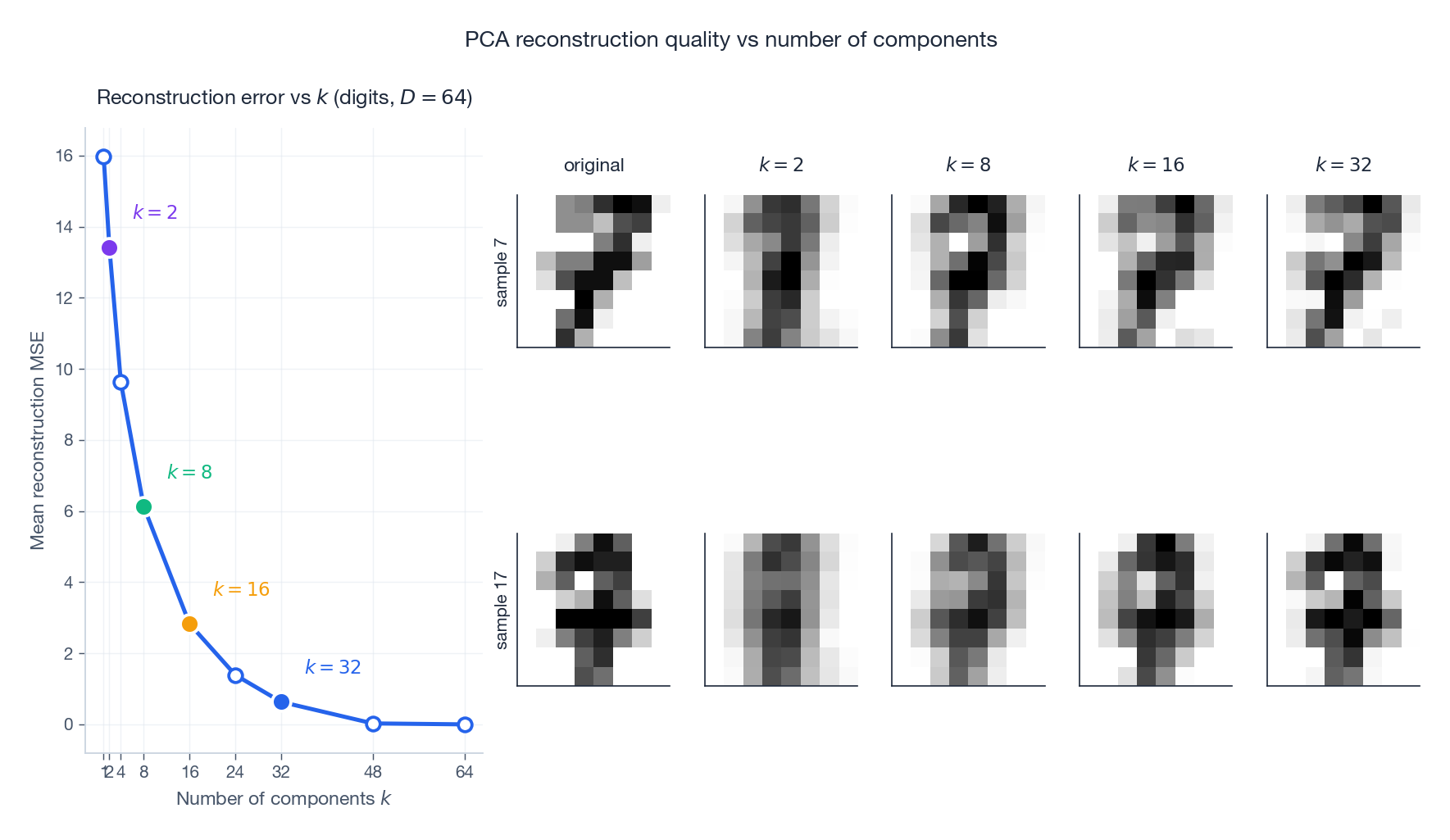
The figure shows what (3) means visually on the digits dataset. The MSE curve falls fast and then plateaus: most of the error is killed by the first handful of components. The image grid on the right shows reconstructions of two specific digits (the original is in the leftmost column). With $k=2$ the reconstructions are blurry blobs; by $k=8$ the digit identity is recognisable; by $k=16$ the strokes are sharp; by $k=32$ the reconstruction is visually indistinguishable from the original. This is the practical statement of (3): how many components you need depends on what level of fidelity you require downstream.
Why this matters. PCA is the unique linear method that simultaneously decorrelates the data and gives the best rank-$k$ approximation in the Frobenius norm. Any other linear “feature extractor” gives up at least one of those properties.
Kernel PCA: Nonlinear Structure via the Kernel Trick#
Where linear PCA fails#
Linear PCA can only ever rotate and project; it cannot bend. If the data lies on a curved manifold — two concentric circles, a Swiss roll, a sphere — then no linear projection can preserve its structure.
PCA in feature space#
$$k(\mathbf{x}_i, \mathbf{x}_j) = \phi(\mathbf{x}_i)^\top \phi(\mathbf{x}_j).$$Common choices:
- Polynomial: $k(\mathbf{x}, \mathbf{y}) = (\mathbf{x}^\top \mathbf{y} + c)^p$
- RBF (Gaussian): $k(\mathbf{x}, \mathbf{y}) = \exp(-\gamma \|\mathbf{x} - \mathbf{y}\|^2)$
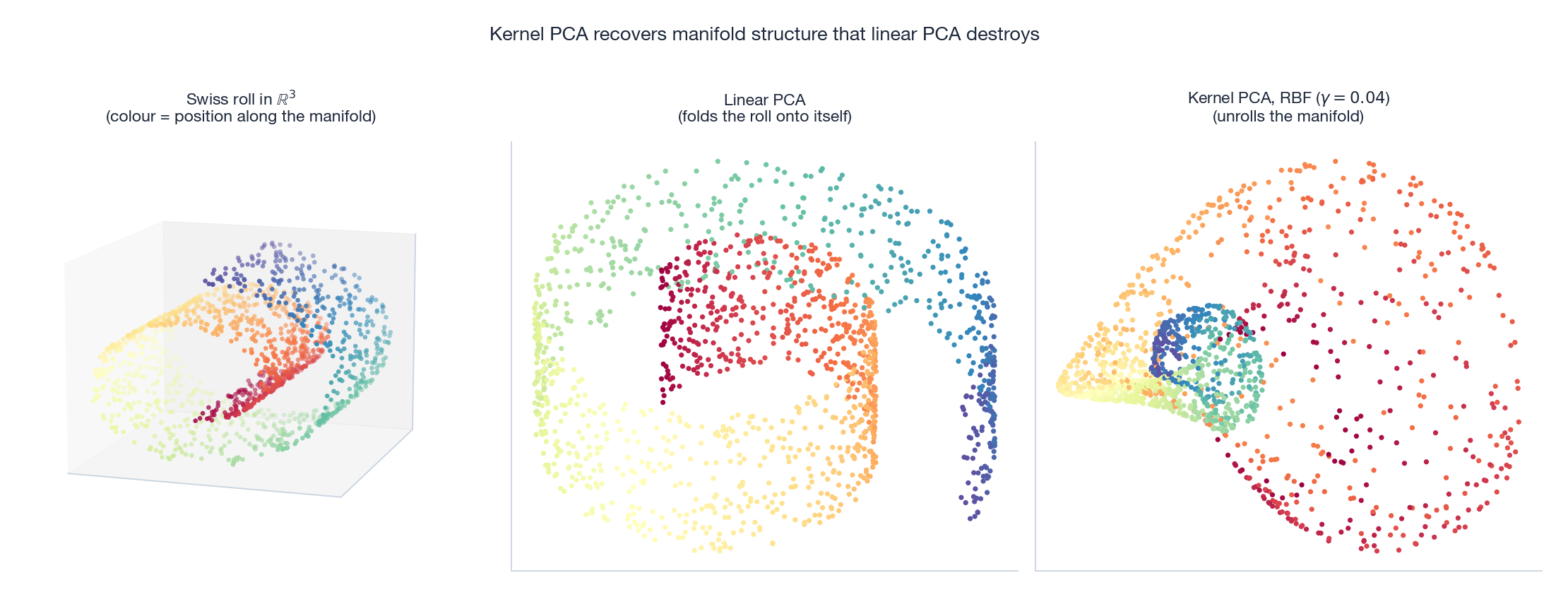
The Swiss roll is the canonical illustration. The original $3$ D manifold is a $2$ D sheet rolled up in space; colour encodes position along the sheet. Linear PCA (middle) projects the roll onto a flat plane, folding distant parts of the manifold on top of each other — yellow ends up next to red. Kernel PCA with an RBF kernel (right) effectively performs PCA in the high-dimensional feature space where the roll becomes more linear, and unrolls the sheet so neighbouring colours end up next to each other in $2$ D.
Cost. Linear PCA is $O(d^3 + Nd^2)$ ; kernel PCA is $O(N^3 + N^2 d)$ . When $N \ll d$ (e.g. genomics with thousands of features and hundreds of samples) kernel PCA can actually be cheaper.
LDA: Supervised Dimensionality Reduction#
PCA ignores labels. If your downstream task is classification, this is wasteful: a direction can have huge variance and yet be perfectly useless for separating classes (and conversely, a tiny-variance direction can be a perfect class separator). Linear Discriminant Analysis uses the labels to find directions that maximise class spread relative to within-class spread.
Two-class derivation#
$$\mathbf{S}_B = (\boldsymbol{\mu}_1 - \boldsymbol{\mu}_2)(\boldsymbol{\mu}_1 - \boldsymbol{\mu}_2)^\top \quad \text{(between-class scatter)}$$ $$\mathbf{S}_W = \sum_{c=1}^{2} \sum_{i \in C_c}(\mathbf{x}_i - \boldsymbol{\mu}_c)(\mathbf{x}_i - \boldsymbol{\mu}_c)^\top \quad \text{(within-class scatter)}$$ $$J(\mathbf{w}) = \frac{\mathbf{w}^\top \mathbf{S}_B\, \mathbf{w}}{\mathbf{w}^\top \mathbf{S}_W\, \mathbf{w}}. \tag{4}$$ $$\mathbf{w}^* \propto \mathbf{S}_W^{-1}(\boldsymbol{\mu}_1 - \boldsymbol{\mu}_2). \tag{5}$$Multi-class#
$$\mathbf{S}_B = \sum_{c=1}^{C} N_c\, (\boldsymbol{\mu}_c - \boldsymbol{\mu})(\boldsymbol{\mu}_c - \boldsymbol{\mu})^\top.$$Solve $\mathbf{S}_B \mathbf{w} = \lambda\, \mathbf{S}_W \mathbf{w}$ for the top eigenvectors. Because $\mathbf{S}_B$ is a sum of $C$ rank-$1$ matrices whose summands are constrained by $\sum_c N_c (\boldsymbol{\mu}_c - \boldsymbol{\mu}) = 0$ , $\mathrm{rank}(\mathbf{S}_B) \le C - 1$ — so LDA can output at most $C - 1$ components. This is a hard ceiling that is easy to forget in practice.
PCA vs LDA at a glance#
| Aspect | PCA | LDA |
|---|---|---|
| Uses labels? | no | yes |
| Objective | max variance | max class-separation |
| Output dim | up to $d$ | up to $C - 1$ |
| Best for | visualisation, denoising | classifier preprocessing |
t-SNE: Neighbour-Preserving Visualisation#

PCA preserves global structure (variance, distances); for $2$ D visualisation that is often the wrong objective. A user looking at a scatter plot wants to see clusters — which usually means preserving local neighbourhoods. t-SNE does exactly this with a probabilistic formulation.
High-dimensional similarities#
$$p_{j \mid i} = \frac{\exp(-\|\mathbf{x}_i - \mathbf{x}_j\|^2 / 2\sigma_i^2)}{\sum_{k \ne i} \exp(-\|\mathbf{x}_i - \mathbf{x}_k\|^2 / 2\sigma_i^2)}, \qquad p_{ij} = \frac{p_{j \mid i} + p_{i \mid j}}{2N}.$$The bandwidth $\sigma_i$ is set per point by binary search so that the perplexity, $2^{H(P_i)}$ , matches a user-chosen value — effectively the number of effective neighbours each point should have.
Low-dimensional similarities and the heavy tail#
$$q_{ij} = \frac{(1 + \|\mathbf{y}_i - \mathbf{y}_j\|^2)^{-1}}{\sum_{k \ne l}(1 + \|\mathbf{y}_k - \mathbf{y}_l\|^2)^{-1}}. \tag{6}$$Why a heavy tail? In $2$ D there is far less “room” than in the original space, so moderately distant points have to crowd together. The Cauchy distribution decays as $\|y\|^{-2}$ instead of the Gaussian’s $\exp(-\|y\|^2)$ , which gives those distant points more space to spread out — this is the crowding problem fix.
Optimisation#
$$\frac{\partial C}{\partial \mathbf{y}_i} = 4\sum_{j}(p_{ij} - q_{ij})(1 + \|\mathbf{y}_i - \mathbf{y}_j\|^2)^{-1}(\mathbf{y}_i - \mathbf{y}_j). \tag{7}$$If $p_{ij} > q_{ij}$ the term is attractive (the points are similar in the original space but too far apart in the embedding); if $p_{ij} < q_{ij}$ it is repulsive.
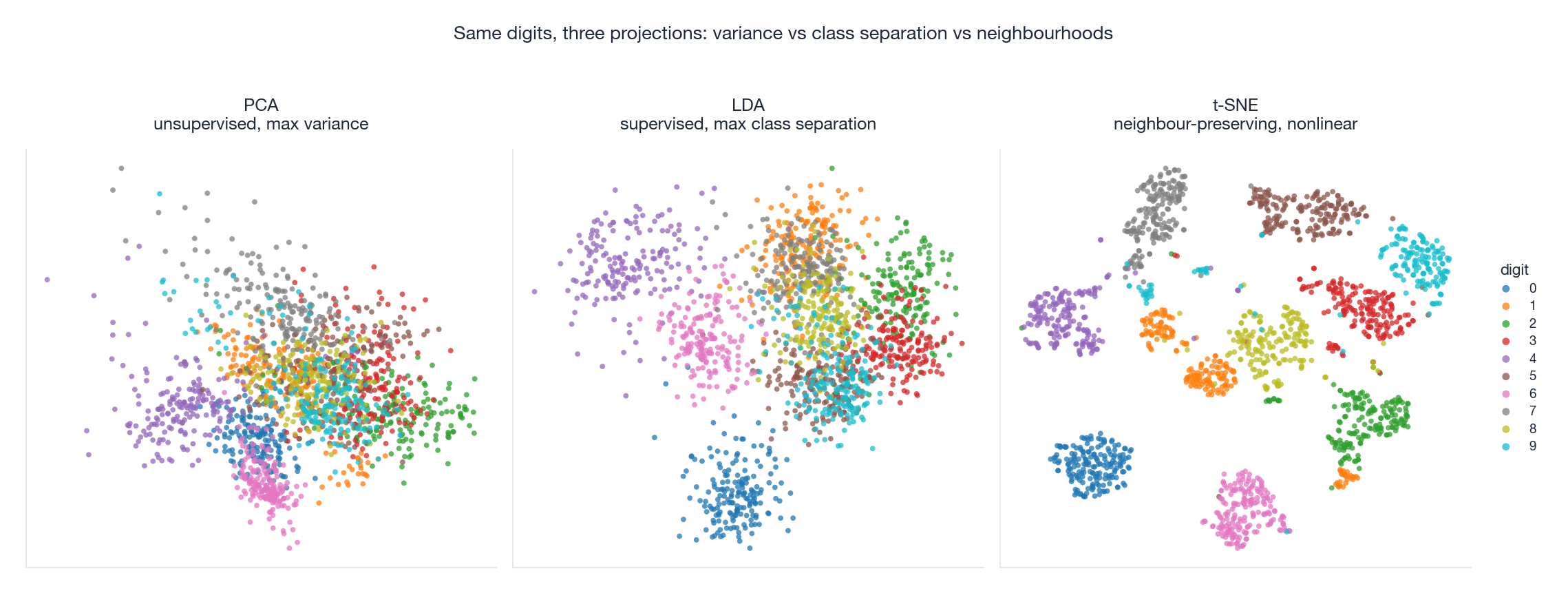
The figure runs all three methods on the digits dataset. PCA (left) packs the variance into two axes but most digit classes are heavily overlapped — variance is not class identity. LDA (middle) explicitly maximises class separation and produces visible class-coloured stripes, but it is restricted to $C - 1 = 9$ output dimensions. t-SNE (right) ignores variance and global geometry entirely; it warps the embedding to keep local neighbours together, producing the crisp, well-separated cluster blobs that you have probably seen in many machine-learning papers.
Caveat for practice. t-SNE is a visualisation tool, not a clustering tool. Inter-cluster distances and cluster sizes in a t-SNE plot are not meaningful, and the result depends on the random seed, perplexity, and number of iterations. Cluster in the original space (or in PCA space); use t-SNE only to show the result.
ICA: When You Need Independence, Not Just Decorrelation#
PCA finds orthogonal directions of maximum variance. Two PCA components are uncorrelated — but uncorrelated is much weaker than independent. If the underlying signals are truly independent and non-Gaussian, you can recover them with Independent Component Analysis (ICA).
$$\mathbf{x} = A\, \mathbf{s},$$and you want to recover $\mathbf{s}$ given only $\mathbf{x}$ and the assumption that the $s_j$ are mutually independent and at most one of them is Gaussian. ICA solves this by finding an unmixing matrix $W$ that maximises the non-Gaussianity of the recovered components $\mathbf{y} = W\mathbf{x}$ — typically via a contrast function such as negentropy or kurtosis. The intuition is the central limit theorem: any linear mixture of independent variables is “more Gaussian” than the variables themselves, so the un-mixed components should be the least Gaussian directions you can find.
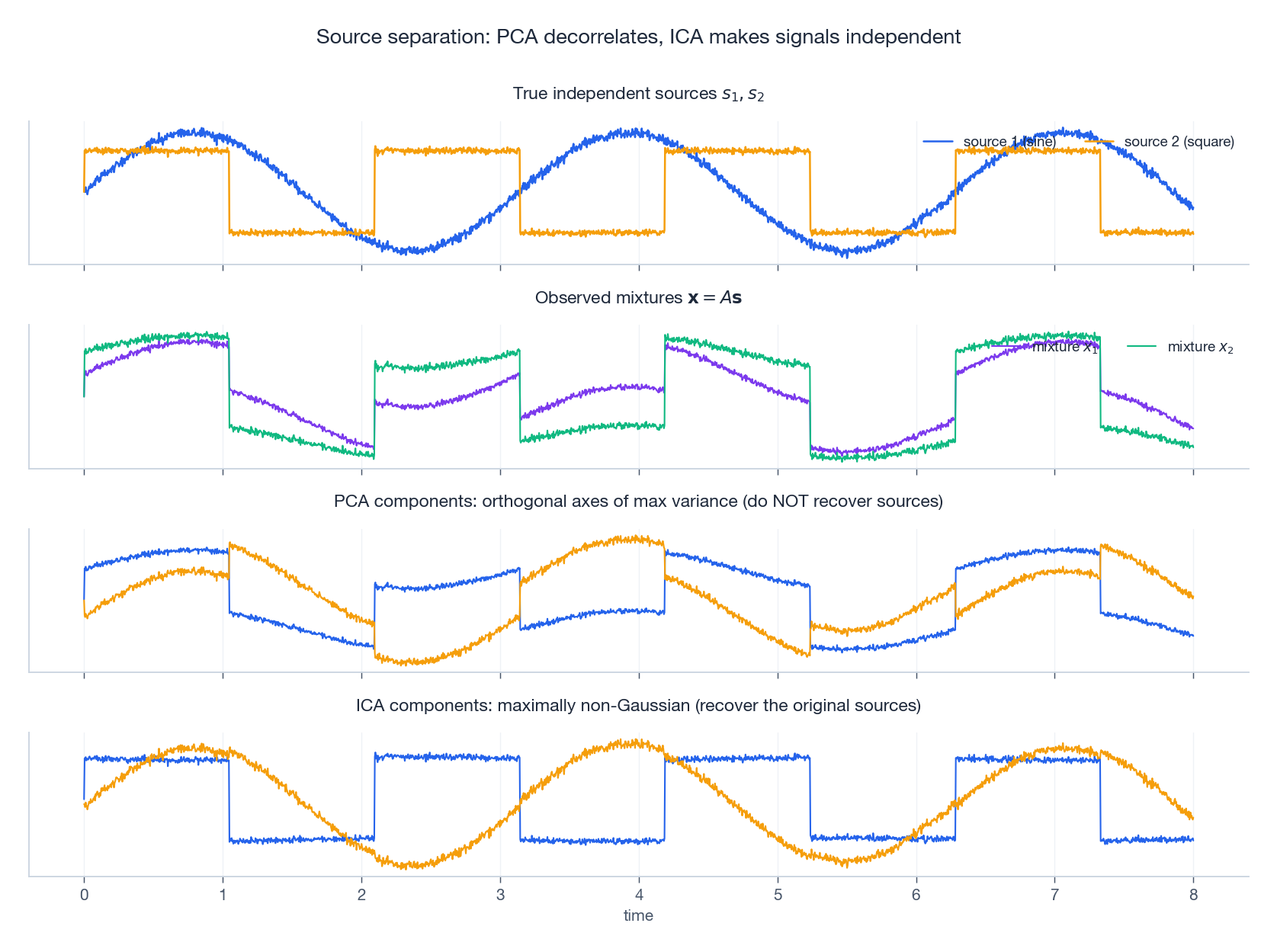
The figure shows what this means concretely. The top row are two true independent sources — a sine and a square wave. The second row are two observed mixtures (each microphone hears both sources). PCA in the third row finds the orthogonal directions of maximum variance, but these axes are not the original sources — you can see the sine and square shapes interfering inside each component. ICA in the bottom row recovers the original sources almost exactly (up to sign and scale, which are inherent ambiguities of the ICA model).
This is the cleanest way to remember the difference: PCA decorrelates, ICA separates. Use PCA for compression and visualisation; use ICA when you have reason to believe your data is a mixture of independent generative sources (audio separation, EEG denoising, fMRI component analysis).
Putting It Together#
| Method | Optimises | Output dim | Linear? | Uses labels? | Use it for |
|---|---|---|---|---|---|
| PCA | variance / reconstruction MSE | up to $d$ | yes | no | denoising, compression, preprocessing |
| Kernel PCA | variance in feature space | up to $N$ | no | no | nonlinear manifolds |
| LDA | class separation (Fisher) | up to $C - 1$ | yes | yes | classifier preprocessing |
| ICA | statistical independence | $K$ sources | yes | no | source separation |
| t-SNE | local neighbourhood KL | $2$ or $3$ | no | no | visualisation only |
| UMAP | fuzzy topological structure | low (any) | no | optional | faster, more global-aware visualisation |
A reasonable default workflow when you meet new tabular data: standardise -> PCA to inspect the spectrum -> drop noise components -> LDA or t-SNE to visualise. Everything else in this article is a tool you reach for when one of the assumptions of that pipeline breaks (nonlinear manifold, independent sources, neighbourhood-only structure, etc.).
Exercises#
Exercise 1 (variance retention). The eigenvalues of $\mathbf{S}$ are $(5, 3, 1, 0.5, 0.5)$ . What fraction of variance do the top two PCs retain?
Solution. Total $= 10$ , top two $= 8$ . Retention $= 80\%$ .
Exercise 2 (centring). Why must the data be centred before PCA?
Solution. The covariance matrix $\mathbf{S} = \tfrac{1}{N}\mathbf{X}^\top \mathbf{X}$ is only the sample covariance when $\bar{\mathbf{x}} = \mathbf{0}$ . Without centring, the largest eigenvector of $\mathbf{X}^\top \mathbf{X}$ points roughly toward the data’s mean direction, not toward the direction of maximum variance.
Exercise 3 (PCA vs t-SNE on $10$ well-separated clusters). What do you expect to see?
Solution. PCA: a linear projection that may fold partly overlapping clusters on top of each other. t-SNE: ten clearly separated blobs, but the distances between blobs and the sizes of the blobs carry no information.
Exercise 4 (kernel PCA on the Swiss roll). Why does it work?
Solution. The RBF feature map sends nearby points (small $\|\mathbf{x}_i - \mathbf{x}_j\|$ ) to similar feature vectors regardless of their global position in the original space. PCA in that feature space therefore captures the manifold’s intrinsic coordinates rather than the ambient $3$ D ones.
Exercise 5 (PCA whitening). What is it and what is it good for?
Solution. The whitening transform $\mathbf{z} = \boldsymbol{\Lambda}^{-1/2} \mathbf{W}^\top \mathbf{x}$ produces components with identity covariance — decorrelated and unit-variance. It is a common preprocessing step before ICA (which assumes whitened input) and a useful normalisation before training models that are sensitive to feature scale.
What’s next#
Dimensionality reduction compresses data onto a low-dimensional manifold, but it does not yet answer the key question: does the data, in that low-dimensional space, naturally fall into groups? That is what clustering does. The next chapter switches from “projection” to “grouping”.
Clustering has no labels, so each algorithm corresponds to an assumption about “what counts as the same group”: K-means assumes spherical clusters with equal variance, GMM assumes ellipsoidal clusters as a probabilistic mixture, DBSCAN assumes density connectivity, spectral clustering assumes a low-conductance cut on a graph, hierarchical clustering assumes a nested structure. I walk through all five — not to memorize algorithms, but so that when you face a new dataset, you can reason from “what do my clusters look like” back to “which algorithm fits”. That meta-skill is the most-overlooked piece of unsupervised learning.
References#
[1] Pearson, K. (1901). On lines and planes of closest fit to systems of points in space. Philosophical Magazine, 2(11), 559-572.
[2] Hotelling, H. (1933). Analysis of a complex of statistical variables into principal components. Journal of Educational Psychology, 24(6), 417-441.
[3] Fisher, R. A. (1936). The use of multiple measurements in taxonomic problems. Annals of Eugenics, 7(2), 179-188.
[4] Scholkopf, B., Smola, A., & Muller, K.-R. (1998). Nonlinear component analysis as a kernel eigenvalue problem. Neural Computation, 10(5), 1299-1319.
[5] Hyvarinen, A., & Oja, E. (2000). Independent component analysis: algorithms and applications. Neural Networks, 13(4-5), 411-430.
[6] Van der Maaten, L., & Hinton, G. (2008). Visualizing data using t-SNE. JMLR, 9(11), 2579-2605.
[7] McInnes, L., Healy, J., & Melville, J. (2018). UMAP: Uniform manifold approximation and projection for dimension reduction. arXiv:1802.03426 .
[8] Jolliffe, I. T. (2002). Principal Component Analysis (2nd ed.). Springer.
This is Part 17 of the ML Mathematical Derivations series. Next: Part 18 — Clustering Algorithms . Previous: Part 16 — Conditional Random Fields .
ML Math Derivations 20 parts
- 01 ML Math Derivations (1): Introduction and Mathematical Foundations
- 02 ML Math Derivations (2): Linear Algebra and Matrix Theory
- 03 ML Math Derivations (3): Probability Theory and Statistical Inference
- 04 ML Math Derivations (4): Convex Optimization Theory
- 05 ML Math Derivations (5): Linear Regression
- 06 ML Math Derivations (6): Logistic Regression and Classification
- 07 ML Math Derivations (7): Decision Trees
- 08 ML Math Derivations (8): Support Vector Machines
- 09 ML Math Derivations (9): Naive Bayes
- 10 ML Math Derivations (10): Semi-Naive Bayes and Bayesian Networks
- 11 ML Math Derivations (11): Ensemble Learning
- 12 ML Math Derivations (12): XGBoost and LightGBM
- 13 ML Math Derivations (13): EM Algorithm and GMM
- 14 ML Math Derivations (14): Variational Inference and Variational EM
- 15 ML Math Derivations (15): Hidden Markov Models
- 16 ML Math Derivations (16): Conditional Random Fields
- 17 ML Math Derivations (17): Dimensionality Reduction and PCA you are here
- 18 ML Math Derivations (18): Clustering Algorithms
- 19 ML Math Derivations (19): Neural Networks and Backpropagation
- 20 ML Math Derivations (20): Regularization and Model Selection