
ML Math Derivations (18): Clustering Algorithms
How do you find groups in unlabeled data? This article derives K-means (Lloyd + K-means++), hierarchical, DBSCAN, spectral, and GMM clustering from their mathematical foundations, with seven figures that show why each algorithm encodes a different prior.
What You Will Learn#
A million customer records arrive with no labels. Can you discover meaningful groups automatically? That is clustering, the most fundamental unsupervised learning task. Unlike classification, clustering forces you to first answer a slippery question: what does “similar” even mean? Every clustering algorithm is, at heart, a different answer to that question — a different geometric, probabilistic, or graph-theoretic prior on what a “group” is.
What you will learn:
- K-means as coordinate descent on a discrete-continuous objective, why Lloyd’s algorithm always converges, and how K-means++ tames the initialization problem.
- Hierarchical clustering as a greedy merge process, and how the choice of linkage controls cluster shape.
- DBSCAN as density-based connectivity, and why it can find non-convex clusters and label noise.
- Spectral clustering as a continuous relaxation of NCut, with the graph Laplacian doing the heavy lifting.
- Gaussian Mixture Models as the probabilistic generalization of K-means — what you gain (soft assignments, ellipsoidal covariance) and what you pay for it.
- How to evaluate clustering quality without labels (silhouette, elbow) and how to choose $K$ .
Prerequisites: linear algebra (eigendecomposition), basic probability, and the EM algorithm from Part 13 .
Formalizing the Clustering Problem#
What Makes a Good Clustering?#
Input: dataset $\mathbf{X} = \{\mathbf{x}_1, \dots, \mathbf{x}_N\}$ with $\mathbf{x}_i \in \mathbb{R}^d$ .
Output: assignments $\{c_1, \dots, c_N\}$ with $c_i \in \{1, \dots, K\}$ .
Two competing principles:
- High cohesion: points in the same cluster should be similar.
- Low coupling: points in different clusters should be dissimilar.
Every objective we will write down is just one way to balance these two. The disagreements between K-means, DBSCAN, and spectral clustering all trace back to how they measure “similar”.
Distance and Similarity Measures#
| Measure | Formula | Best for |
|---|---|---|
| Euclidean | $d(\mathbf{x}, \mathbf{y}) = \\mid\mathbf{x} - \mathbf{y}\\mid_2$ | dense, low-dimensional data |
| Manhattan | $d(\mathbf{x}, \mathbf{y}) = \sum_j \\mid x_j - y_j\\mid$ | sparse features, anomaly-resistant |
| Cosine | $\text{sim}(\mathbf{x}, \mathbf{y}) = \frac{\mathbf{x}^T\mathbf{y}}{\\mid\mathbf{x}\\mid\\mid\mathbf{y}\\mid}$ | text, high-dimensional sparse vectors |
| Mahalanobis | $d(\mathbf{x}, \mathbf{y}) = \sqrt{(\mathbf{x}-\mathbf{y})^T \boldsymbol{\Sigma}^{-1}(\mathbf{x}-\mathbf{y})}$ | correlated features |
Practical note: because every distance is sensitive to feature scale, standardize before clustering unless your features are already on comparable scales.
Evaluating Clusters Without Labels#
$$s(i) = \frac{b(i) - a(i)}{\max\{a(i), b(i)\}} \in [-1, 1]$$where $a(i)$ is the mean intra-cluster distance and $b(i)$ is the mean distance to the nearest other cluster. A value near $1$ means $i$ is comfortably inside its cluster; near $0$ it lies on a boundary; negative means it was probably misassigned. Averaging $s(i)$ over the dataset gives a label-free quality score.
K-means: Centroid-Driven Clustering#
The Objective#
$$J(\{c_i\}, \{\boldsymbol{\mu}_k\}) = \sum_{k=1}^{K} \sum_{i: c_i = k} \\|\mathbf{x}_i - \boldsymbol{\mu}_k\\|^2 \tag{1}$$This is a joint optimization over discrete assignments $\{c_i\}$ and continuous centroids $\{\boldsymbol{\mu}_k\}$ — and it is NP-hard in general. Lloyd’s algorithm tackles it with coordinate descent: alternately optimize one set of variables while fixing the other.
Lloyd’s Algorithm#
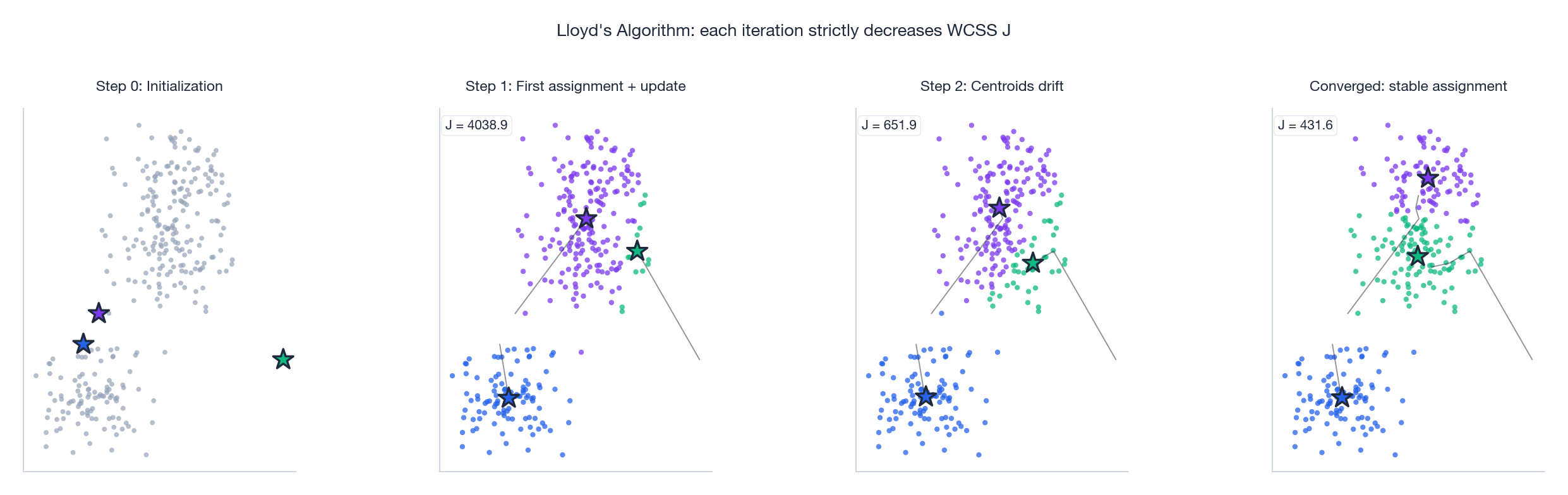
Why it converges. Each step can only decrease $J$ . Step 1 is optimal because every point ends up at its closest centroid. Step 2 is optimal because the within-cluster mean is the unique minimizer of squared error (the second derivative $2|C_k|\mathbf{I}$ is positive definite). Since $J \geq 0$ and decreases monotonically, the iteration must terminate at a local minimum — usually within tens of iterations on real data, as the four-panel figure above shows: centroids drift from a poor initialization, decision regions reshape, and $J$ collapses to a steady value.
Caveat: local optima. Lloyd’s algorithm finds a local minimum, not the global one. Bad initialization can trap it on a plateau where two centroids share one true cluster while a third tries to cover two. The standard remedy is to restart Lloyd’s algorithm from many random initializations and keep the lowest-$J$ run; a smarter remedy is K-means++.
| |
K-means++ Initialization#
The idea is simple: spread initial centroids far apart by picking each one with probability proportional to its squared distance from the existing seeds.
- Pick $\boldsymbol{\mu}_1$ uniformly at random from $\mathbf{X}$ .
- For $k = 2, \dots, K$ : compute $D(\mathbf{x}_i)^2 = \min_{j < k} \\|\mathbf{x}_i - \boldsymbol{\mu}_j\\|^2$ , then sample $\boldsymbol{\mu}_k = \mathbf{x}_i$ with probability $D(\mathbf{x}_i)^2 / \sum_l D(\mathbf{x}_l)^2$ .
Theoretical guarantee (Arthur & Vassilvitskii, 2007): the expected K-means++ objective is at most $8(\ln K + 2) \cdot J_{\text{opt}}$ — a logarithmic-in-$K$ approximation, without even running Lloyd’s iterations afterward.
Choosing $K$ : Silhouette and Elbow#
K-means makes you specify $K$ . The two most common ways to pick it are visualized below.
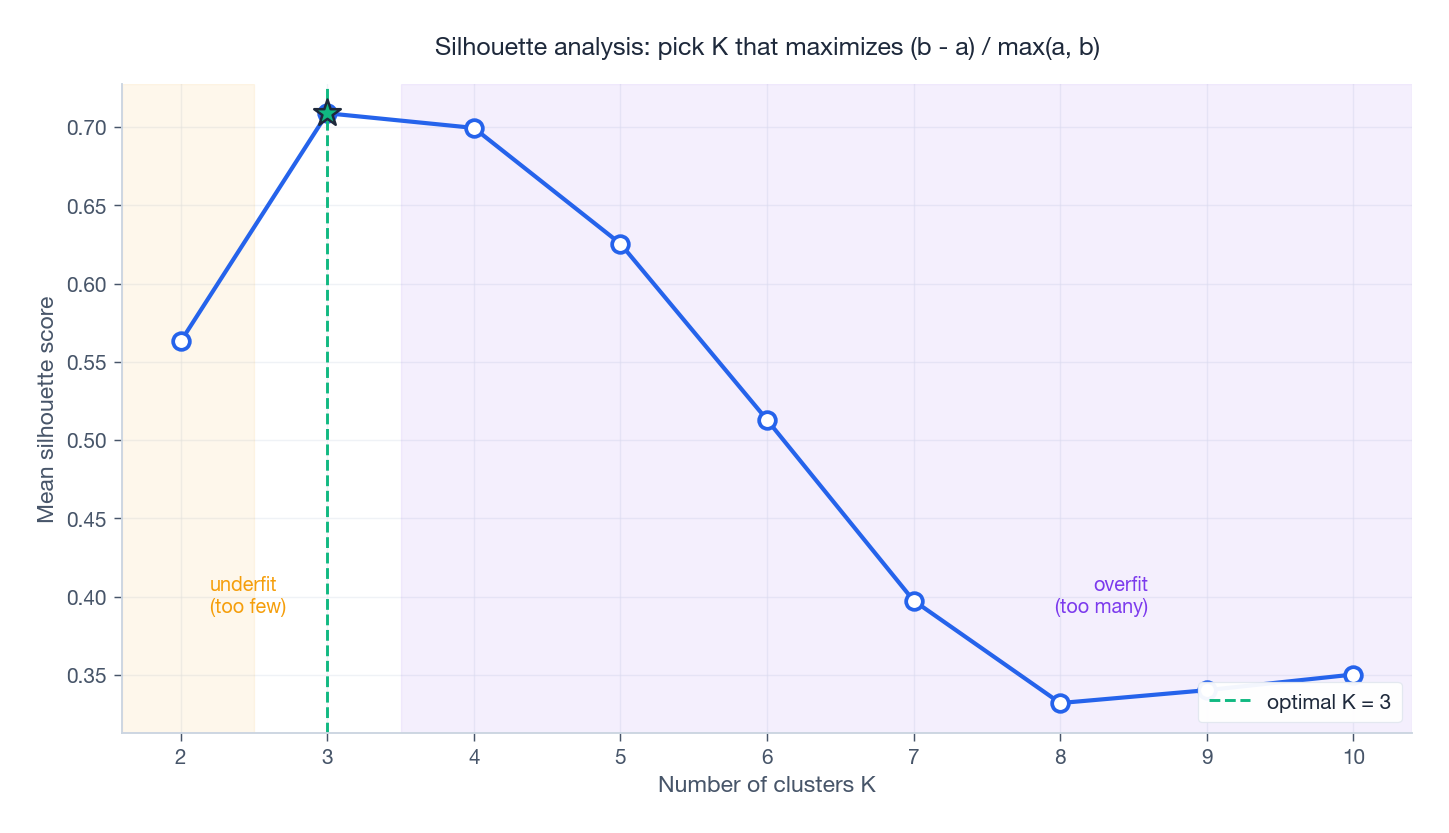
The silhouette curve typically peaks at the right $K$ and decays on both sides. The elbow plot tells the same story from a different angle:
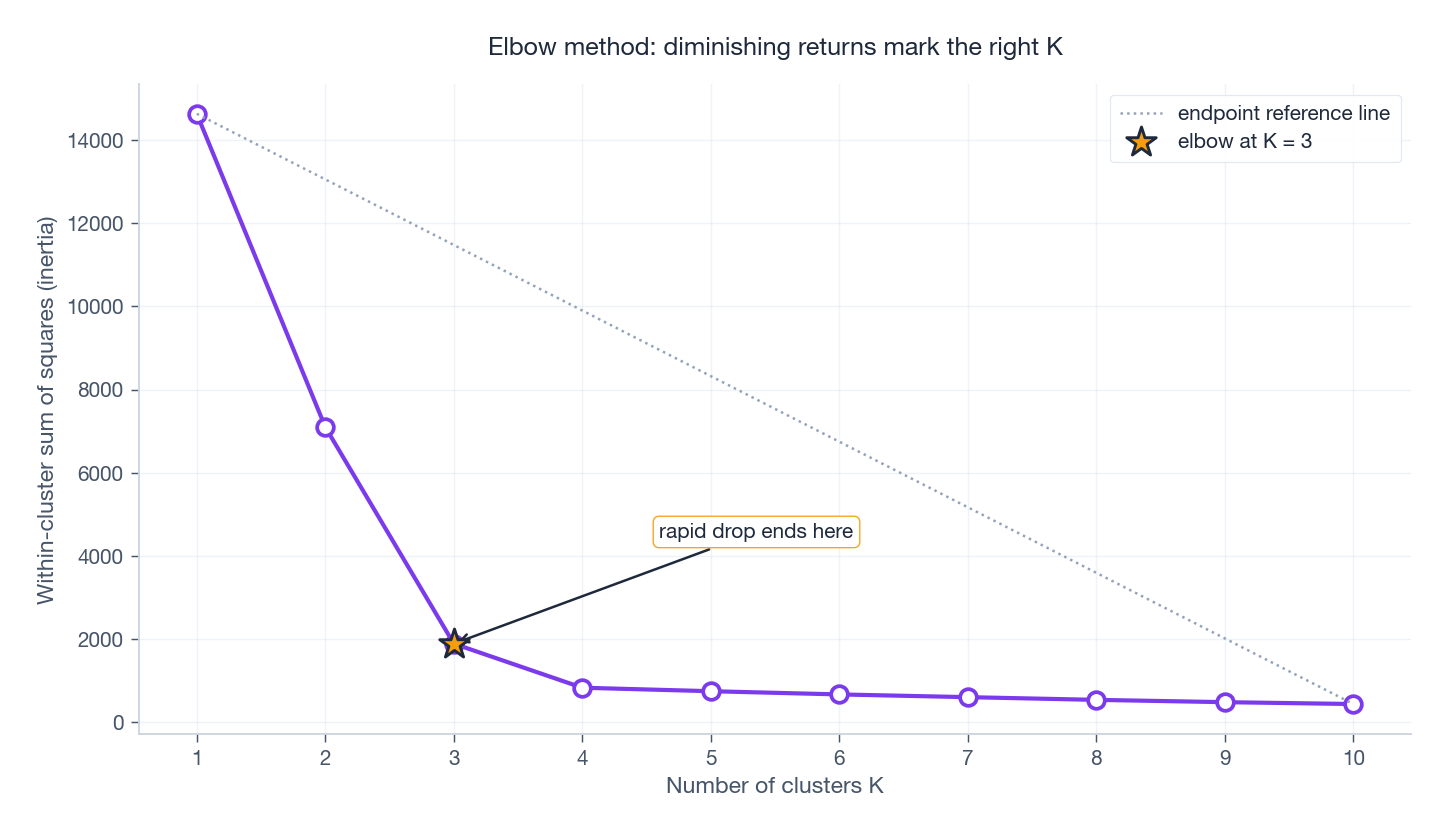
WCSS strictly decreases as $K$ grows (more centroids can always fit better), so we look for the kink where extra clusters stop helping much. The point of maximum perpendicular distance from the line connecting the two endpoints is a robust elbow heuristic.
Limitations#
| Limitation | Cause | Remedy |
|---|---|---|
| Need to specify $K$ | no built-in model selection | silhouette / elbow / BIC |
| Spherical clusters only | uses Euclidean distance, equal radii | GMM, spectral, kernel K-means |
| Sensitive to outliers | mean is not robust | K-medoids |
| Scale-sensitive | distance is dominated by largest features | standardize first |
Hierarchical Clustering#
Agglomerative Approach#
Build a dendrogram from the bottom up:
- Start with $N$ singleton clusters.
- Merge the two closest clusters.
- Repeat until one cluster (or until you have $K$ ).
The result is a binary tree where the height of each merge encodes the distance at which it happened. To get a flat partition, draw a horizontal line through the tree — every branch it cuts is one cluster.
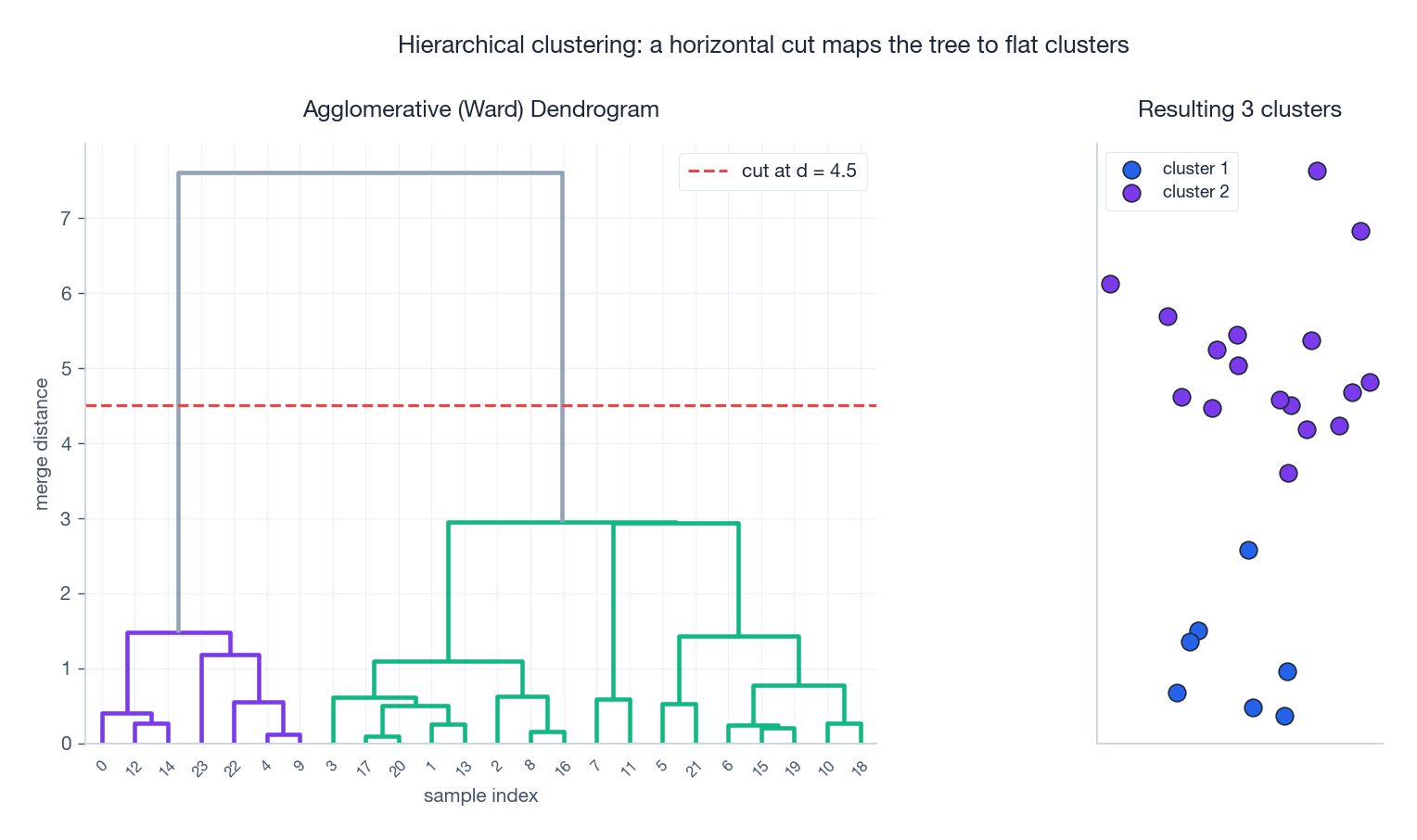
Linkage Criteria#
The merge rule depends on linkage, which is just a choice of how to define distance between two sets of points:
| Linkage | $d(C_i, C_j)$ | Character |
|---|---|---|
| Single | $\min_{a \in C_i, b \in C_j} d(a, b)$ | finds elongated/chained clusters; fragile to noise bridges |
| Complete | $\max_{a \in C_i, b \in C_j} d(a, b)$ | compact, roughly spherical clusters; noise-robust |
| Average | $\frac{1}{\lvert C_i\rvert\lvert C_j\rvert}\sum_{a,b} d(a,b)$ | balanced compromise |
| Ward | minimize variance increase $\Delta\!J$ after merge | most consistent with the K-means objective; produces balanced sizes |
Ward is the default choice for most numerical data because its objective coincides with WCSS reduction.
When to Use It#
- You don’t know $K$ and want to see the data’s natural granularity.
- You want a hierarchy (e.g. taxonomy of products, gene families).
- $N$ is small-to-medium (naive complexity is $O(N^3)$ ; $O(N^2 \log N)$ with priority queues).
DBSCAN: Density-Based Clustering#

Core Concepts#
DBSCAN (“Density-Based Spatial Clustering of Applications with Noise”) replaces the “every point belongs to some cluster” assumption with a density rule. It uses two parameters: neighborhood radius $\epsilon$ and minimum points $\text{MinPts}$ .
- $\epsilon$ -neighborhood: $N_\epsilon(\mathbf{x}) = \{\mathbf{x}' \in \mathbf{X} : \\|\mathbf{x} - \mathbf{x}'\\| \leq \epsilon\}$ .
- Core point: $|N_\epsilon(\mathbf{x})| \geq \text{MinPts}$ — it sits in a dense neighborhood.
- Border point: not core, but inside some core point’s neighborhood.
- Noise point: neither — DBSCAN explicitly labels it as an outlier.
Density Reachability#
DBSCAN grows clusters by chaining core points:
- Directly density-reachable: $\mathbf{x}'$ lies in $\mathbf{x}$ ’s neighborhood, and $\mathbf{x}$ is a core point.
- Density-reachable: there is a chain of directly density-reachable links from $\mathbf{x}$ to $\mathbf{x}'$ .
- Density-connected: both points are density-reachable from a common third point.
A cluster is a maximal set of density-connected points. This is why DBSCAN handles arbitrary cluster shapes — no assumption of convexity, no assumption of equal radii, no fixed $K$ .
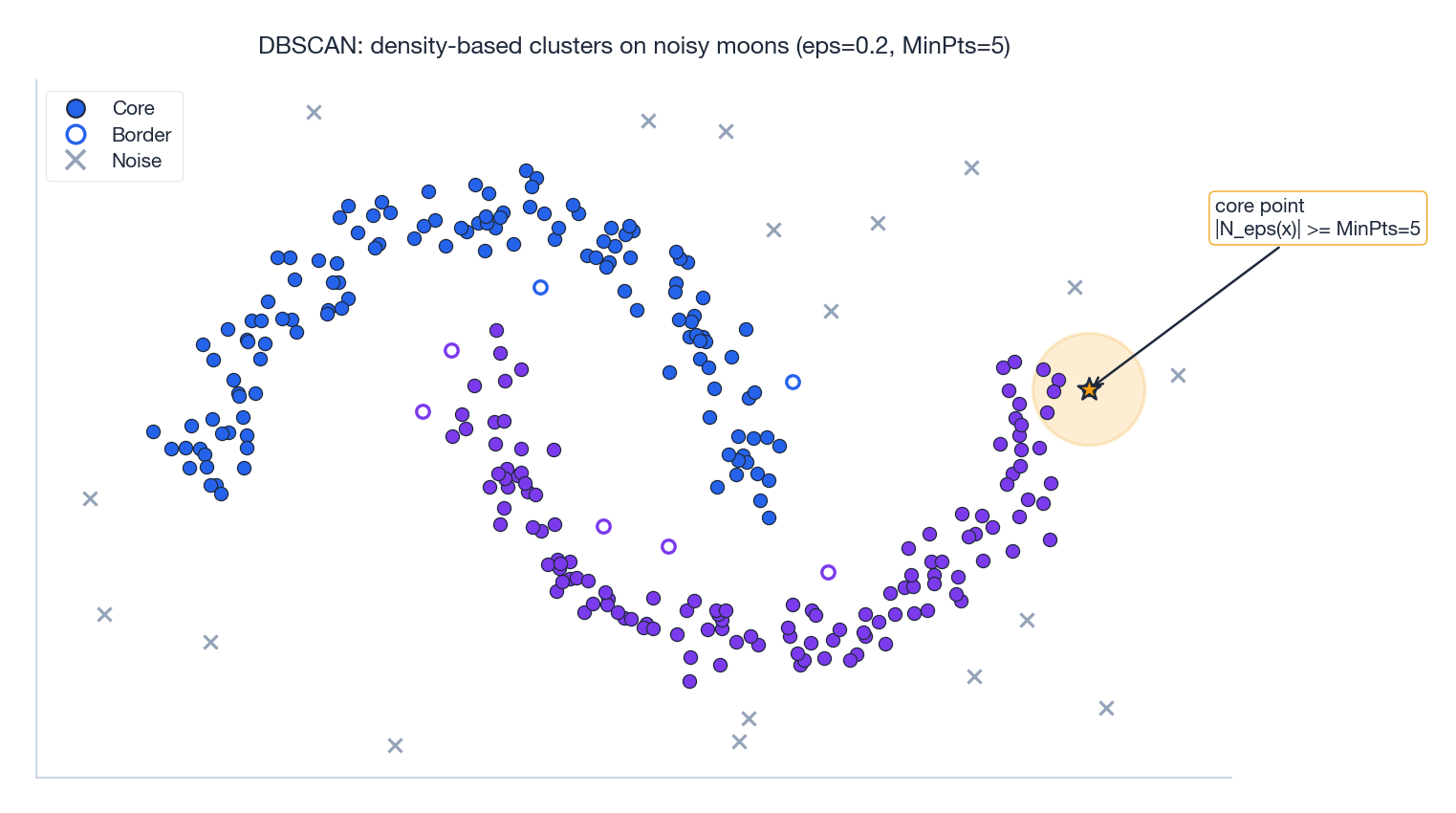
The figure makes the bookkeeping concrete. The amber circle around the highlighted core point contains at least $\text{MinPts}=5$ neighbors, qualifying it as a core point. Any other point that falls inside such a ball joins the same cluster, and from there the density-reachability relation propagates outward through the moon shape. Stray points in low-density regions never accumulate enough neighbors and get labeled noise.
Choosing $\epsilon$ : the K-Distance Plot#
Plot, for every point, the distance to its $k$ -th nearest neighbor (with $k = \text{MinPts}$ ), then sort those values descending. The curve has a knee where it transitions from “dense” to “sparse” distances — pick $\epsilon$ at that knee.
Strengths and Weaknesses#
Strengths. No need to specify $K$ ; finds arbitrary shapes; identifies noise; robust to outliers.
Weaknesses. Two parameters that interact ($\epsilon$ , $\text{MinPts}$ ); struggles when clusters have very different densities (use HDBSCAN for this); suffers in high dimensions where Euclidean distances concentrate.
Spectral Clustering: A Graph Theory Approach#
From Data to Graphs#
$$W_{ij} = \exp\left(-\frac{\\|\mathbf{x}_i - \mathbf{x}_j\\|^2}{2\sigma^2}\right)$$For scalability, sparsify with $k$ -nearest-neighbor or $\epsilon$ -ball graphs.
The Graph Laplacian#
$$\mathbf{L} = \mathbf{D} - \mathbf{W}.$$ $$\mathbf{f}^T \mathbf{L} \mathbf{f} = \tfrac{1}{2}\sum_{i,j} W_{ij}(f_i - f_j)^2 \geq 0.$$That is, $\mathbf{L}$ measures how much a function $\mathbf{f}$ varies across edges. Smooth functions on the graph — those that change slowly between strongly connected nodes — have small Laplacian quadratic form, and they are exactly the eigenvectors of $\mathbf{L}$ with the smallest eigenvalues.
$$\mathbf{L}_{\text{sym}} = \mathbf{I} - \mathbf{D}^{-1/2}\mathbf{W}\mathbf{D}^{-1/2}.$$Normalized Cut Objective#
$$\text{NCut}(A, B) = \frac{\text{cut}(A, B)}{\text{vol}(A)} + \frac{\text{cut}(A, B)}{\text{vol}(B)} \tag{3}$$where $\text{cut}(A,B) = \sum_{i \in A, j \in B} W_{ij}$ and $\text{vol}(A) = \sum_{i \in A} D_{ii}$ . Dividing by volume prevents the trivial “split off one point” solution. Combinatorially this is NP-hard, but a continuous relaxation — letting cluster indicators take real values — has an exact closed-form solution: it is the eigenvalue problem for $\mathbf{L}_{\text{sym}}$ .
The Algorithm#
- Build the similarity matrix $\mathbf{W}$ .
- Compute the Laplacian $\mathbf{L}_{\text{sym}}$ .
- Compute the $K$ eigenvectors corresponding to the smallest eigenvalues.
- Stack them as columns of $\mathbf{U} \in \mathbb{R}^{N \times K}$ .
- Normalize each row of $\mathbf{U}$ to unit length.
- Run K-means on the rows to recover discrete cluster labels.
Why this works. Step 3 maps the data into a low-dimensional embedding where graph-connected clusters become linearly separable point clouds. K-means, which fails on the original non-convex shape, succeeds in this embedding.
Complexity. $O(N^2)$ to build $\mathbf{W}$ , $O(N^3)$ for the eigendecomposition. For large $N$ , use sparse $k$ -NN graphs and Lanczos / Nyström approximation.
Gaussian Mixture Models: K-means with Probabilities#
The Generative Model#
$$p(\mathbf{x}) = \sum_{k=1}^{K} \pi_k \, \mathcal{N}(\mathbf{x} \mid \boldsymbol{\mu}_k, \boldsymbol{\Sigma}_k).$$Fit by EM (see Part 13 ):
- E-step: posterior responsibility $\gamma_{ik} = \pi_k \mathcal{N}(\mathbf{x}_i \mid \boldsymbol{\mu}_k, \boldsymbol{\Sigma}_k) / \sum_j \pi_j \mathcal{N}(\mathbf{x}_i \mid \boldsymbol{\mu}_j, \boldsymbol{\Sigma}_j)$ .
- M-step: weighted updates $\boldsymbol{\mu}_k = \sum_i \gamma_{ik} \mathbf{x}_i / \sum_i \gamma_{ik}$ , similarly for $\boldsymbol{\Sigma}_k$ and $\pi_k$ .
Why GMM Generalizes K-means#
K-means is the limit of GMM with $\boldsymbol{\Sigma}_k = \sigma^2 \mathbf{I}$ and $\sigma \to 0$ : as the variance shrinks, the soft posterior $\gamma_{ik}$ collapses to a one-hot vector, and the M-step becomes the centroid mean. So K-means is GMM with two strong assumptions hard-baked: spherical equal-radius clusters, hard assignments.
GMM relaxes both — it allows ellipsoidal covariance (rotated, stretched clusters) and soft membership (a point can be 70% cluster A, 30% cluster B). The price is more parameters and slower convergence.
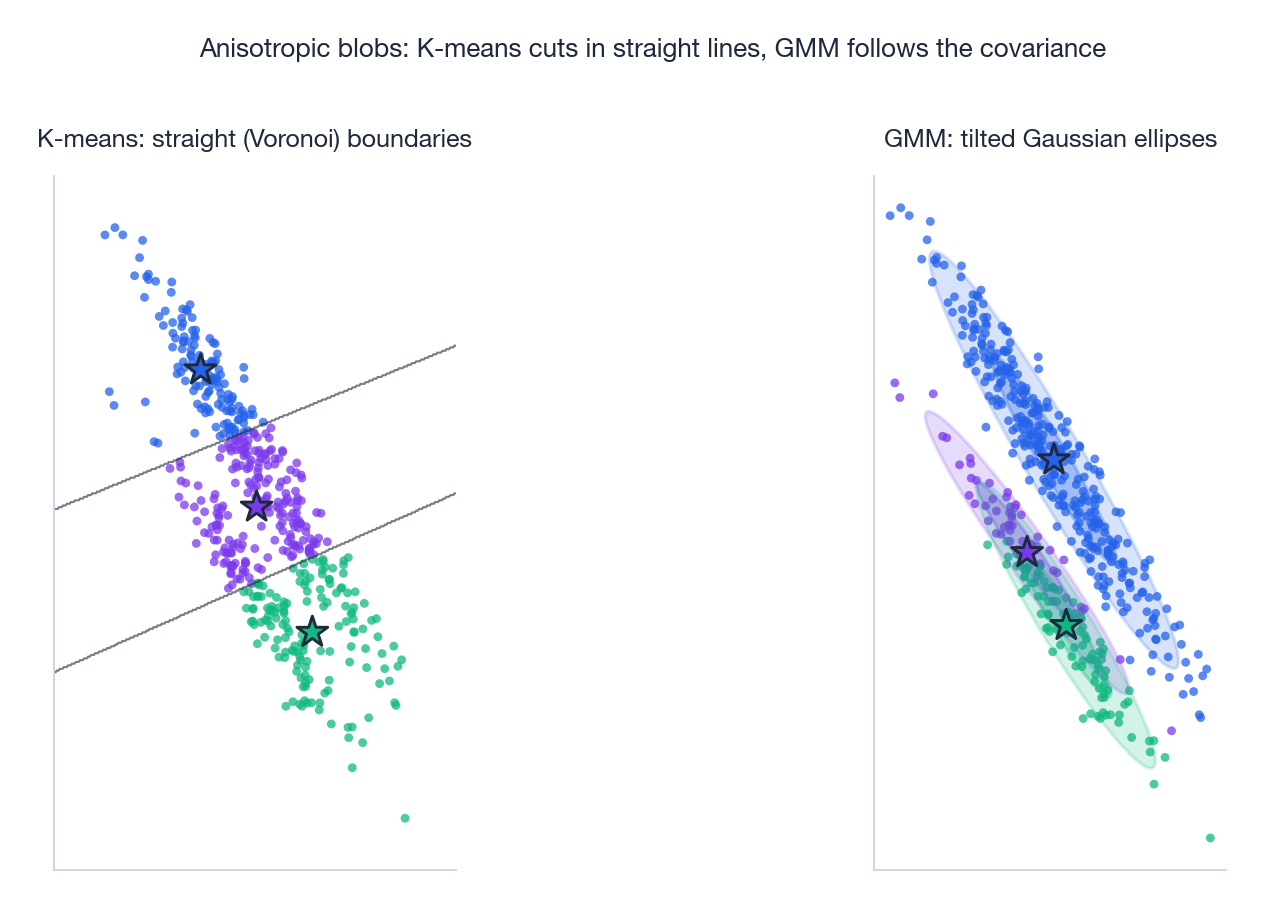
The figure makes the difference vivid. On stretched, tilted clusters, K-means’ Voronoi cells slice across them at the wrong angles, splitting one true cluster between two centroids. GMM’s ellipses align with the data’s covariance and recover the true partition.
Putting It All Together: When to Use What#
Different priors, different victories. The grid below runs K-means, DBSCAN, and spectral clustering on three classic dataset shapes:
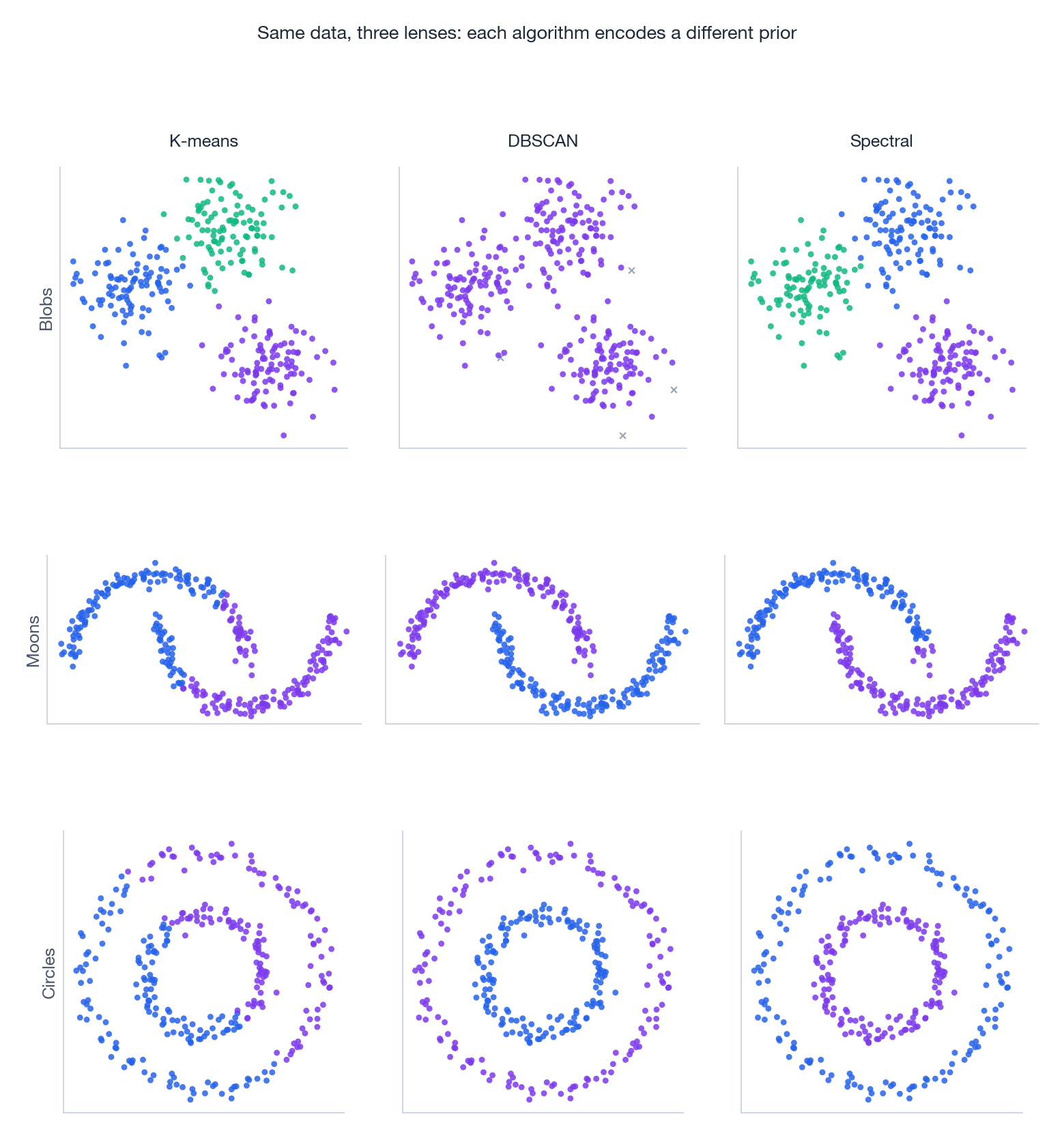
Reading the grid:
- Blobs (top row). All three succeed — this is the easy case. K-means is fastest.
- Moons (middle). K-means slices each moon in half (its prior is convex, the data isn’t). DBSCAN and spectral both follow the curves correctly.
- Circles (bottom). K-means again fails for the same reason. DBSCAN traces the inner and outer rings via density chains; spectral exploits the graph structure.
Rule of thumb:
| If your data… | Try first |
|---|---|
| has roughly spherical, well-separated clusters | K-means |
| has elongated or non-convex clusters with noise | DBSCAN |
| has non-convex clusters in a smooth manifold | Spectral |
| needs soft assignments or covariance modelling | GMM |
| needs a hierarchy / unknown $K$ | Agglomerative (Ward) |
Cost and complexity comparison#
The five algorithms above span four orders of magnitude in cost. Choosing the wrong one for your scale is the most common mistake I see — people reach for spectral clustering on a million points, then wonder why their machine froze.
| Algorithm | Time | Space | Notes |
|---|---|---|---|
| K-means (Lloyd) | $O(NKDI)$ | $O(NK)$ | $I$ = iterations, usually 10-50 |
| Mini-batch K-means | $O(BKDI)$ | $O(NK)$ | Batch size $B \ll N$ , near-identical quality |
| Hierarchical (agglomerative) | $O(N^2 \log N)$ time, $O(N^2)$ space | $O(N^2)$ | Distance matrix is the bottleneck |
| DBSCAN | $O(N \log N)$ with spatial index, $O(N^2)$ without | $O(N)$ | Index requires $D \lesssim 20$ |
| Spectral | $O(N^3)$ for full eigendecomp | $O(N^2)$ | Use Nyström or Lanczos for $N > 10^4$ |
| GMM (EM) | $O(NKD^2 I)$ for full covariance | $O(NK + KD^2)$ | $O(NKDI)$ for diagonal covariance |
The numbers that matter in practice. K-means on 1M points × 128 dimensions × 10 clusters × 30 iterations is $\sim 4 \times 10^{10}$ operations — about 30 seconds on a modern CPU. The same problem on hierarchical clustering needs the $10^{12}$ -entry distance matrix, which by itself is 4 TB at float32. DBSCAN with a KD-tree handles it in a few minutes; without the tree it dies. Spectral clustering on 100k points needs $10^{10}$ memory for the affinity matrix — already too large.
This is why mini-batch K-means and Nyström-approximated spectral clustering exist: not because the original algorithms are wrong, but because their constants are too large for modern data sizes.
Production gotchas I keep hitting#
K-means on cosine-similar embeddings. If your features are L2-normalised text or image embeddings, Euclidean distance reduces to angular distance, but K-means computes cluster means by Euclidean averaging, which produces non-unit vectors. The next assignment step then mixes “distance to a unit vector” with “distance to a sub-unit vector” — silently breaking the geometry. Fix: re-normalise centroids after every M-step (this is spherical K-means), or use K-means on the original unit-sphere data and accept the bias.
The K-means “elbow” is rarely an elbow. The sum of within-cluster squared distances $J(K)$ is monotonically decreasing in $K$ , so any plot looks vaguely elbow-shaped if you squint. On real data the curve is usually smooth with no sharp bend. Use the silhouette score or gap statistic instead — both have a maximum at the right $K$ , not just an inflection. Even better: ask the downstream task what $K$ it needs, instead of letting the data dictate.
DBSCAN’s eps is data-dependent. The single hardest hyperparameter in clustering. Too small: everything is noise. Too large: everything is one cluster. The standard heuristic is the k-distance plot: sort all points by distance to their $k$
-th nearest neighbour ($k = $
min_samples), look for the knee. The knee value is your eps. This works well in 2-3 dimensions and degrades fast in higher dimensions because nearest-neighbour distances concentrate.
GMM with covariance_type='full' on high-dimensional data. Each component has $D(D+1)/2$
covariance parameters. With $D = 768$
(BERT embeddings) and $K = 50$
components, that is 14M parameters before counting the means — almost certainly more than your sample count. Use covariance_type='diag' or run PCA to $D = 50$
first. The same warning applies to LDA, QDA, and any model that estimates a full covariance per class.
Cluster labels are arbitrary across runs. K-means run twice produces the same partition but different label IDs. If you compare cluster assignments across versions, use the Hungarian algorithm to align labels first, or use a label-invariant metric like adjusted Rand index. I have seen entire dashboards “break” on retraining for this reason alone.
Exercises#
Exercise 1: K-means convergence. Prove that the update step minimizes WCSS for fixed assignments.
Solution. Take $\partial / \partial \boldsymbol{\mu}_k$ of $\sum_{i \in C_k}\\|\mathbf{x}_i - \boldsymbol{\mu}_k\\|^2$ and set to zero: $\boldsymbol{\mu}_k = \tfrac{1}{|C_k|}\sum_{i \in C_k}\mathbf{x}_i$ . The Hessian is $2|C_k|\mathbf{I} \succ 0$ , so this is a global minimum.
Exercise 2: DBSCAN. With $\text{MinPts} = 3$ , $\epsilon = 0.5$ , point $\mathbf{x}$ has 2 neighbors within radius 0.5. Is $\mathbf{x}$ a core point?
Solution. The $\epsilon$ -neighborhood includes $\mathbf{x}$ itself plus 2 neighbors = 3 points. Since $3 \geq 3$ , yes, $\mathbf{x}$ is a core point. (If $\text{MinPts}$ were $4$ it would be a border or noise point.)
Exercise 3: GMM vs K-means. When should you reach for GMM instead of K-means?
Solution. When clusters are elliptical (different variances along different axes), when you need soft assignments (e.g. for downstream Bayesian reasoning), or when you want a generative model you can sample from. Stick with K-means when clusters are spherical, $N$ is huge, or you need fast iteration.
Exercise 4: Linkage choice. Why does single-linkage often fail on noisy data?
Solution. Single-linkage merges based on the minimum pairwise distance, so a single noise point bridging two clusters is enough to chain them together (the “chaining effect”). Complete- and Ward-linkage use aggregate distances and are robust to such bridges.
Exercise 5: Silhouette. $a(i) = 2$ , $b(i) = 5$ . Compute $s(i)$ .
Solution. $s = (5 - 2)/\max(2, 5) = 3/5 = 0.6$ . A respectable score: point $i$ is roughly twice as far from the next cluster as from its own.
What’s next#
The core algorithms of classical machine learning are now covered — linear, tree, kernel, Bayesian, latent-variable, sequence, dimensionality reduction, clustering. They share one limitation: their expressivity is bounded by hand-crafted features or kernels. The next chapter breaks that limitation outright — neural networks and backpropagation.
Neural networks turn “features” into things learned jointly with parameters: each layer is a nonlinear transform of the previous, and training optimizes all the layer weights simultaneously by gradient descent. Backpropagation is the chain rule run efficiently on a computation graph — nothing mysterious about it on its own, but it is what makes end-to-end learning possible. I derive the universal approximation theorem, the Jacobian form of backprop, the exponential expressivity advantage of depth, and why the loss landscape “looks non-convex but is in practice optimizable”. Modern deep learning is, almost entirely, the engineering extrapolation of this chapter.
References#
[1] Lloyd, S. (1982). Least squares quantization in PCM. IEEE Trans. Info. Theory, 28(2), 129-137.
[2] Arthur, D., & Vassilvitskii, S. (2007). k-means++: The advantages of careful seeding. SODA, 1027-1035.
[3] Ester, M., Kriegel, H. P., Sander, J., & Xu, X. (1996). A density-based algorithm for discovering clusters in large spatial databases with noise. KDD, 226-231.
[4] Ng, A. Y., Jordan, M. I., & Weiss, Y. (2002). On spectral clustering: Analysis and an algorithm. NIPS, 849-856.
[5] Von Luxburg, U. (2007). A tutorial on spectral clustering. Statistics and Computing, 17(4), 395-416.
[6] Shi, J., & Malik, J. (2000). Normalized cuts and image segmentation. IEEE Trans. PAMI, 22(8), 888-905.
[7] Ward, J. H. (1963). Hierarchical grouping to optimize an objective function. JASA, 58(301), 236-244.
[8] Campello, R. J., Moulavi, D., & Sander, J. (2013). Density-based clustering based on hierarchical density estimates (HDBSCAN). PAKDD, 160-172.
This is Part 18 of the ML Mathematical Derivations series. Next: Part 19 — Neural Networks and Backpropagation . Previous: Part 17 — Dimensionality Reduction and PCA .
ML Math Derivations 20 parts
- 01 ML Math Derivations (1): Introduction and Mathematical Foundations
- 02 ML Math Derivations (2): Linear Algebra and Matrix Theory
- 03 ML Math Derivations (3): Probability Theory and Statistical Inference
- 04 ML Math Derivations (4): Convex Optimization Theory
- 05 ML Math Derivations (5): Linear Regression
- 06 ML Math Derivations (6): Logistic Regression and Classification
- 07 ML Math Derivations (7): Decision Trees
- 08 ML Math Derivations (8): Support Vector Machines
- 09 ML Math Derivations (9): Naive Bayes
- 10 ML Math Derivations (10): Semi-Naive Bayes and Bayesian Networks
- 11 ML Math Derivations (11): Ensemble Learning
- 12 ML Math Derivations (12): XGBoost and LightGBM
- 13 ML Math Derivations (13): EM Algorithm and GMM
- 14 ML Math Derivations (14): Variational Inference and Variational EM
- 15 ML Math Derivations (15): Hidden Markov Models
- 16 ML Math Derivations (16): Conditional Random Fields
- 17 ML Math Derivations (17): Dimensionality Reduction and PCA
- 18 ML Math Derivations (18): Clustering Algorithms you are here
- 19 ML Math Derivations (19): Neural Networks and Backpropagation
- 20 ML Math Derivations (20): Regularization and Model Selection