
PDE and ML (8): Reaction-Diffusion Systems and Graph Neural Networks
Deep GNNs collapse because they are diffusion equations on graphs. Turing's reaction-diffusion theory tells us how to fix it -- and closes the eight-chapter PDE+ML series.

Anyone who has trained a deep GNN has seen it collapse — past a dozen or so layers, every node’s embedding becomes nearly identical and the model goes mush. There is a name for this — over-smoothing — and the underlying math is surprisingly clean: GNN message passing is essentially a diffusion equation on the graph, and diffusion’s long-time behavior is to flatten everything to a constant.
In 1952 Turing studied how zebra stripes form and proposed the reaction-diffusion equation: diffusion homogenizes, reaction fights back to maintain inhomogeneity. The interplay can produce stable patterns — this is Turing instability. Drag this idea onto graphs and you get a class of architectures designed to resist over-smoothing: diffusion smears information out, reaction insists on keeping differences alive.
This is the eighth and final article in the series. We’ll dissect over-smoothing, port Turing’s 1952 reaction-diffusion theory to graphs, give a concrete reaction-GNN architecture, and step back over the whole eight-chapter PDE+ML arc — every chapter, in retrospect, has been doing the same thing: using mathematical structure as the inductive bias for a neural network.
What You Will Learn#
Stack 32 layers of GCN on a citation graph and accuracy collapses from 81 % to 20 %. Every node converges to the same vector. This is over-smoothing, the GNN equivalent of heat death — and the diagnosis comes straight from PDE theory. A GCN layer is one explicit-Euler step of the heat equation on a graph, and the heat equation has exactly one fixed point: the constant. The cure was published in 1952. Alan Turing showed that adding a reaction term to a diffusion equation can make a uniform state spontaneously break apart into stripes, spots, or labyrinths. The same trick — a learned reaction term — keeps deep GNNs alive.
This closes the PDE + Machine Learning series. We have spent seven chapters arguing that essentially every modern neural architecture is, secretly, the discretisation of a PDE. Reaction-diffusion + GNNs closes the loop: it is the most explicitly PDE-shaped architecture of all, and it lets us re-examine every preceding chapter through one final lens.
What you will learn
- Reaction-diffusion equations on continuous space — Gray-Scott, FitzHugh-Nagumo, the morphologies they produce.
- Turing instability — the linear-stability argument that explains how diffusion can create structure.
- Graph Laplacians — the discrete analogue of $\nabla^2$ , and why its spectrum dictates GNN behaviour.
- GCN $=$ discretised graph diffusion — and the spectral proof of over-smoothing.
- Reaction-diffusion GNNs (GRAND, GRAND++, RDGNN) — adding a reaction term that keeps node features distinct.
- A retrospective on the entire PDE + ML series.
Prerequisites: linear algebra (eigen-decomposition), basic PDE concepts (the diffusion equation), and familiarity with message-passing GNNs.
Reaction-Diffusion in Continuous Space#

The General Form#
$$\frac{\partial \mathbf{u}}{\partial t} = \mathbf{D}\,\nabla^2\mathbf{u} + \mathbf{R}(\mathbf{u}). \tag{1}$$- The diffusion term $\mathbf{D}\nabla^2\mathbf{u}$ is linear and smoothing — it always reduces gradients.
- The reaction term $\mathbf{R}(\mathbf{u})$ is local (no spatial derivatives) and nonlinear — it can either reinforce or oppose the smoothing.
Two perspectives are useful. Physically, $\mathbf{u}$ is a vector of concentrations; diffusion is Fick’s law, reaction is the local rate equation. Mathematically, (1) is a semilinear parabolic PDE — the heat equation we met in Chapter 7 , plus a pointwise nonlinear forcing.
The remarkable thing — and Turing’s insight — is that the competition between these two terms can produce stable, non-trivial spatial patterns from a uniform initial state. Call this diffusion-driven instability.
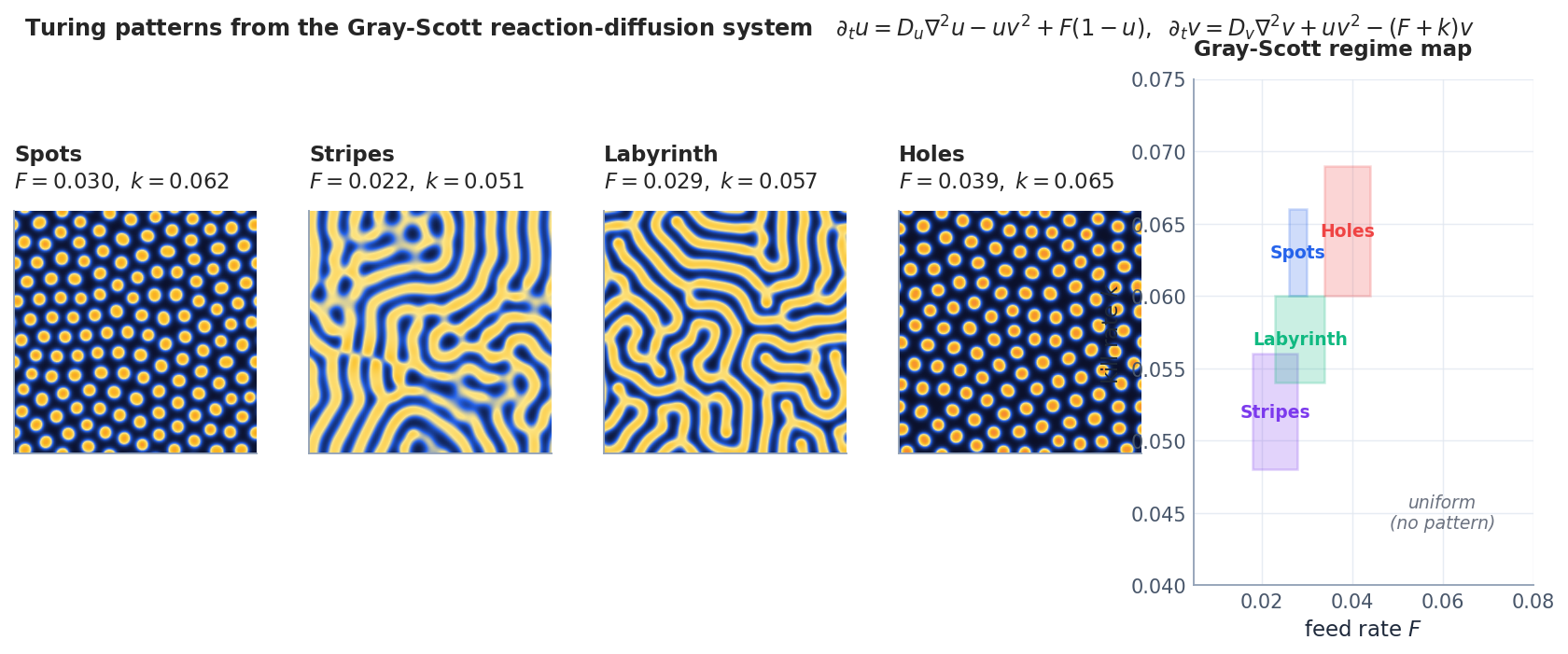
Gray-Scott#

- $u$ is a substrate fed in at rate $F$ ; $v$ is an autocatalyst that consumes $u$ via $u + 2v \to 3v$ and decays at rate $k$ .
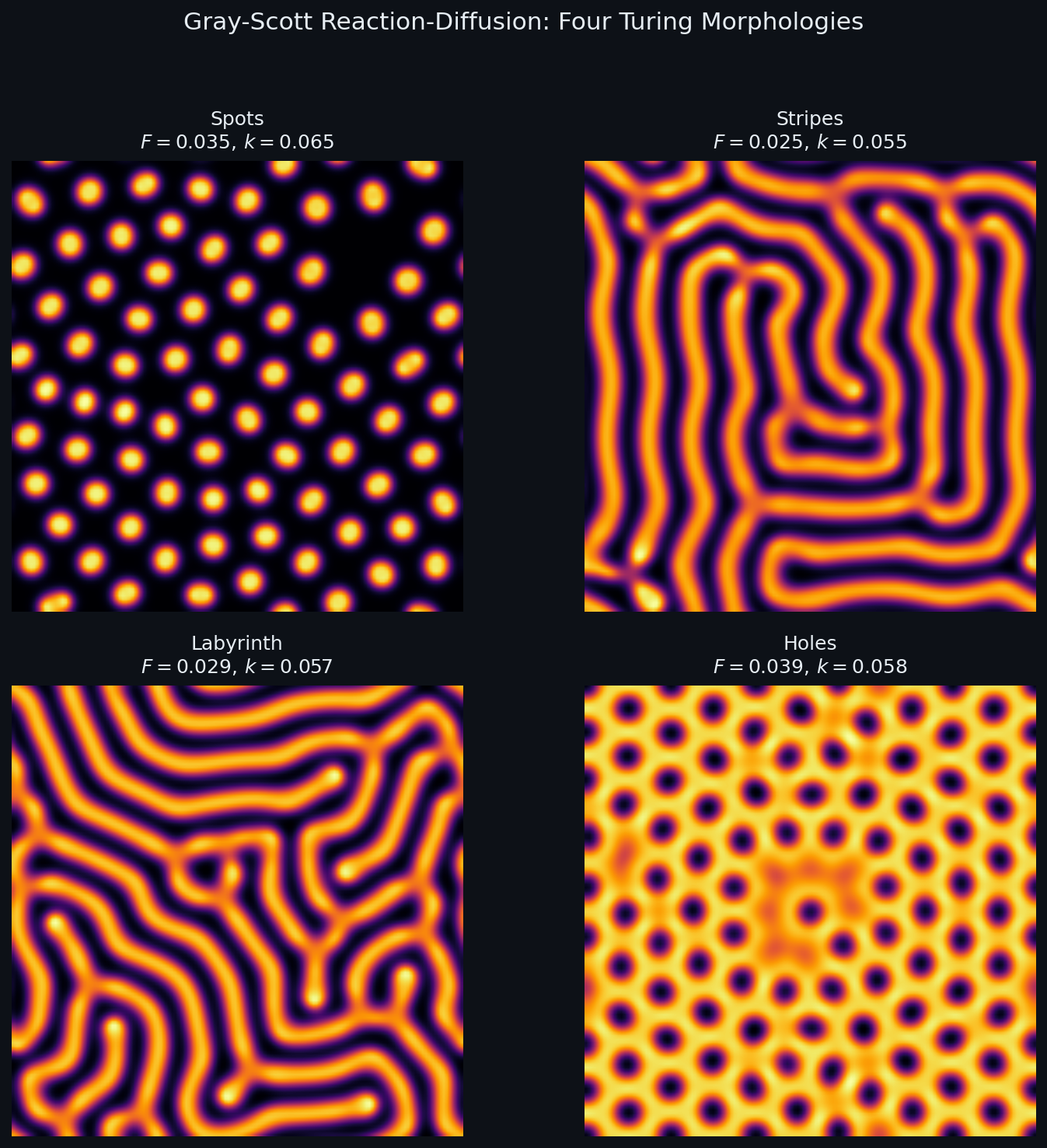
- With $D_u > D_v$ (substrate diffuses faster than the catalyst), small patches of $v$ stabilise into the morphologies of Fig. 1.
The same equation, with different $(F, k)$ , gives spots, stripes, labyrinths, holes, moving spots, even self-replicating spots — Pearson (1993) catalogued a dozen distinct regimes.
FitzHugh-Nagumo#
$$ \partial_t v = D \nabla^2 v + v - \tfrac{v^3}{3} - w + I,\qquad \partial_t w = \varepsilon\,(v + \beta - \gamma w),\quad \varepsilon \ll 1. $$- $v$ is the fast membrane potential; $w$ is a slow recovery variable.
- The cubic nonlinearity makes $v$ excitable: a super-threshold push sets off a stereotyped pulse that the slow $w$ then resets.
In 2D you get spiral waves and target patterns — exactly the patterns observed in cardiac tissue during arrhythmia and in the developing chick retina (see Fig. 6 below).
Implementation: Gray-Scott pattern simulation. The Gray-Scott system is the textbook example of a reaction-diffusion system that produces Turing patterns. Here is a complete simulation:
| |
Each $(F, k)$ pair produces a qualitatively different pattern. The mechanism is exactly Turing’s insight: the activator $V$ diffuses slowly and amplifies locally, while the inhibitor $U$ diffuses fast and suppresses at long range. Their competition creates spatial structure from an almost-uniform initial state.
Turing Instability: Patterns from Uniformity#
The Question#
Pick a uniform steady state $\bar{\mathbf{u}}$ with $\mathbf{R}(\bar{\mathbf{u}}) = \mathbf{0}$ that is stable in the well-mixed (no diffusion) system. Can adding diffusion ever destabilise it?
The naive answer is no — diffusion only smooths, surely it can only help stability. Turing (1952) proved that intuition wrong.
The Argument#
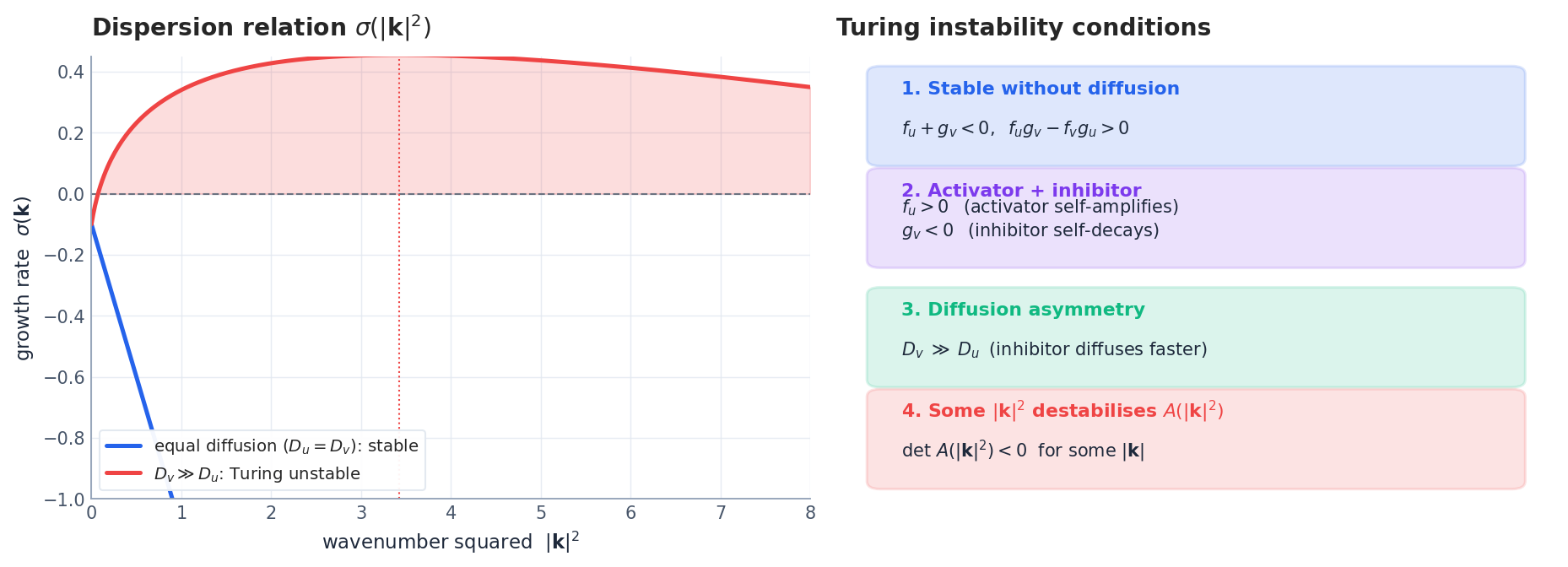
$$ \sigma\,\mathbf{q} \;=\; \underbrace{\bigl(\mathbf{J} - |\mathbf{k}|^2\,\mathbf{D}\bigr)}_{\mathbf{A}(|\mathbf{k}|^2)}\,\mathbf{q},\qquad \mathbf{J} = \nabla_{\mathbf{u}}\mathbf{R}(\bar{\mathbf{u}}). \tag{2} $$The mode $\mathbf{q}\,e^{i\mathbf{k}\cdot\mathbf{x}}$ grows when $\mathbf{A}(|\mathbf{k}|^2)$ has an eigenvalue with positive real part. The full Turing condition is then a list of four inequalities (Fig. 2, right):
- $\mathrm{tr}\,\mathbf{J} < 0$ and $\det\,\mathbf{J} > 0$ — the well-mixed system is stable.
- The Jacobian has activator-inhibitor structure: $f_u > 0$ , $g_v < 0$ , with $f_v\,g_u < 0$ .
- Diffusion asymmetry: $D_v \gg D_u$ — the inhibitor diffuses much faster than the activator.
- There exists some $|\mathbf{k}|^2$ with $\det\,\mathbf{A}(|\mathbf{k}|^2) < 0$ , i.e. an unstable wavenumber.
Conditions 1-3 are algebraic facts about the kinetics. Condition 4 follows once the first three hold and is the actual mechanism: a band of wavenumbers becomes unstable, and the most unstable mode $|\mathbf{k}_*|$ sets the characteristic length scale of the resulting pattern, $\ell \sim 2\pi/|\mathbf{k}_*|$ .
The Intuition Behind Diffusion-Driven Instability#
Why does asymmetric diffusion destabilise an otherwise-stable steady state? Imagine a tiny local bump of activator. Locally, the activator self-amplifies (positive feedback). It also produces inhibitor — but the inhibitor diffuses away quickly, so its concentration near the bump stays low, while far away the inhibitor builds up and suppresses other potential bumps. This is short-range activation, long-range inhibition — the universal recipe behind animal coat patterns, vegetation stripes, sand ripples, and (we’ll see in §5 ) the architecture of deep GNNs.
Implementation: Turing instability checker. Given the reaction kinetics and diffusion coefficients, we can check algorithmically whether a Turing instability will occur:
| |
This is the algorithmic version of the four Turing conditions from the theory section. You can use it to predict whether a given reaction-diffusion system will spontaneously form patterns — before running any simulation.
From Grids to Graphs#
Why Graphs?#
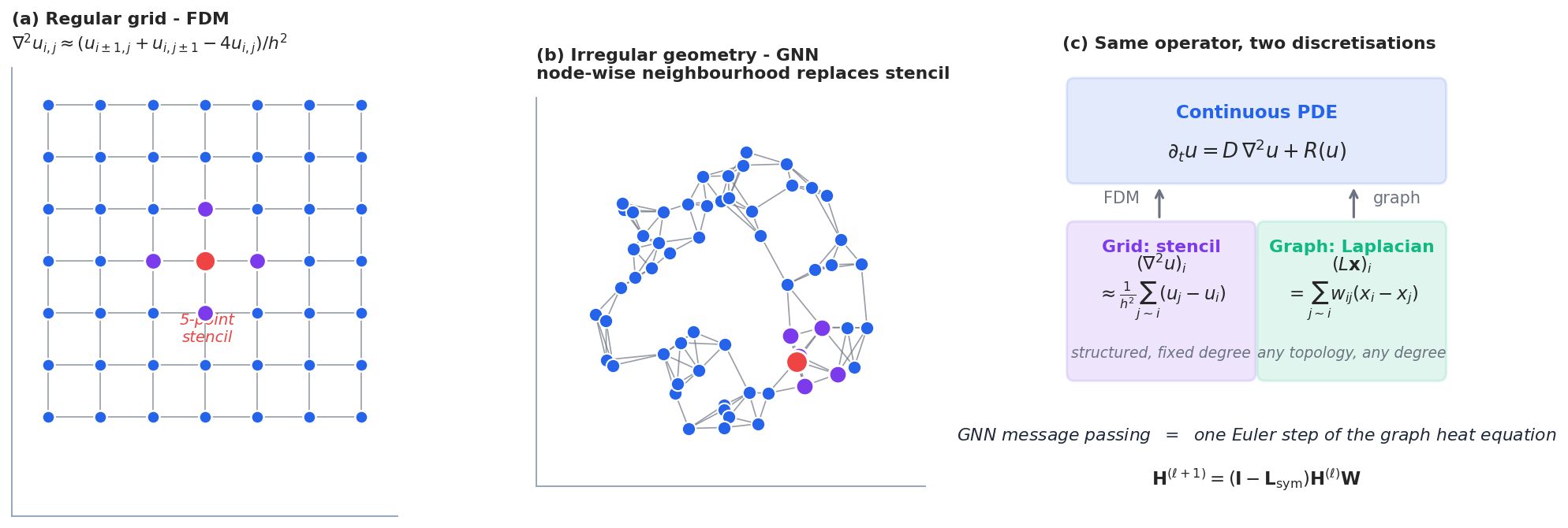
Finite differences (FDM) and finite elements (FEM) discretise PDEs on regular grids or carefully designed meshes. They are extremely powerful when the domain is simple. But for molecular structures, social networks, citation graphs, road networks, brain connectomes, there is no natural notion of a regular grid — the connectivity is the geometry.
A graph $G = (V, E)$ generalises both: it is just a set of nodes plus a relation between them. A GNN solves a “PDE” on this graph. The job of this section is to write down what that PDE looks like.
The Graph Laplacian#
For a weighted, undirected graph with adjacency matrix $\mathbf{A}$ and degree matrix $\mathbf{D} = \mathrm{diag}(d_i)$ :
| Variant | Formula | Spectrum lives in |
|---|---|---|
| Combinatorial | $\mathbf{L} = \mathbf{D} - \mathbf{A}$ | $[0, 2 d_{\max}]$ |
| Symmetric normalised | $\mathbf{L}_{\text{sym}} = \mathbf{I} - \mathbf{D}^{-1/2}\mathbf{A}\mathbf{D}^{-1/2}$ | $[0, 2]$ |
| Random-walk | $\mathbf{L}_{\text{rw}} = \mathbf{I} - \mathbf{D}^{-1}\mathbf{A}$ | $[0, 2]$ |
The Laplacian is the discrete analogue of $-\nabla^2$ in the sense that it integrates the squared gradient. It is symmetric positive semi-definite, with spectral decomposition $\mathbf{L} = \mathbf{U}\boldsymbol{\Lambda}\mathbf{U}^{\!\top}$ , eigenvalues $0 = \lambda_1 \leq \lambda_2 \leq \cdots \leq \lambda_n$ .
The smallest eigenvalue is always zero, with eigenvector proportional to $\mathbf{1}$ (the constant). The second smallest, $\lambda_2$ — the algebraic connectivity — measures how well-connected the graph is.
The Graph Heat Equation#
$$\frac{d\mathbf{X}}{dt} = -\mathbf{L}\mathbf{X}. \tag{4}$$ $$\hat x_k(t) = e^{-\lambda_k t}\,\hat x_k(0),\qquad \hat x_k = \mathbf{u}_k^{\!\top}\mathbf{X}(0).$$Every mode decays exponentially at its own rate $\lambda_k$ — except $\lambda_1 = 0$ , which is preserved forever. As $t \to \infty$ only the constant survives.
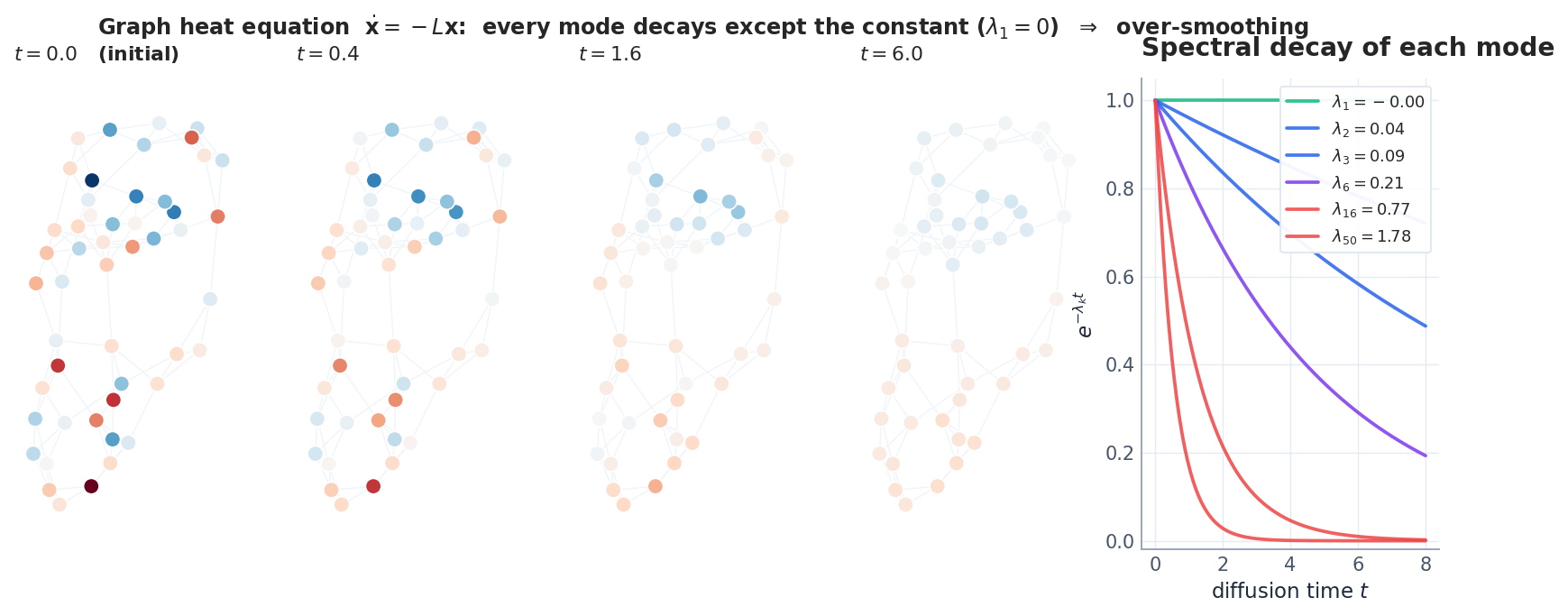
This is over-smoothing in its purest form, and we have not even mentioned neural networks yet.
GCN Is the Heat Equation#

The Identification#
$$ \mathbf{H}^{(\ell+1)} = \sigma\bigl(\tilde{\mathbf{A}}\,\mathbf{H}^{(\ell)}\,\mathbf{W}^{(\ell)}\bigr), \qquad \tilde{\mathbf{A}} = \tilde{\mathbf{D}}^{-1/2}(\mathbf{A} + \mathbf{I})\tilde{\mathbf{D}}^{-1/2}. $$ $$ \mathbf{H}^{(\ell+1)} = \tilde{\mathbf{A}}\,\mathbf{H}^{(\ell)} \;=\; \bigl(\mathbf{I} - \tilde{\mathbf{L}}_{\text{sym}}\bigr)\mathbf{H}^{(\ell)}. \tag{5} $$This is exactly the explicit Euler step of the graph heat equation $\dot{\mathbf{H}} = -\tilde{\mathbf{L}}_{\text{sym}}\mathbf{H}$ with step size $h = 1$ . The “self-loops” trick $\mathbf{A} + \mathbf{I}$ is just the standard FDM stabilisation that pushes the spectrum of $\tilde{\mathbf{L}}_{\text{sym}}$ into $[0, 2)$ so the explicit scheme remains stable.
The Spectral Proof of Over-Smoothing#
$$\mathbf{H}^{(L)} = \tilde{\mathbf{A}}^L\,\mathbf{H}^{(0)}.$$ $$\tilde{\mathbf{A}}^L \xrightarrow[L \to \infty]{} \pi_{\text{const}}.$$Every node feature collapses onto the same vector. This is not a quirk of GCN — it is a theorem about iterating any low-pass filter. Adding ReLU and learning the weight matrices delays the collapse but does not prevent it: Oono & Suzuki (2020) proved that for an arbitrary weight matrix sequence with bounded singular values, GCN features still converge to a low-dimensional subspace.
Continuous-Depth GNNs#
$$\frac{d\mathbf{X}}{dt} = -\mathcal{L}_\theta(\mathbf{X})\,\mathbf{X},\qquad \mathbf{X}(T) = \text{output.}$$$\mathcal{L}_\theta$ is a learned attention-weighted Laplacian, and the integration is done with an off-the-shelf ODE solver (Dormand-Prince, etc.). This does not fix over-smoothing — solving a heat equation more accurately is still solving a heat equation. GRAND++ (Thorpe et al., 2022) adds a source term, and RDGNN (Eliasof et al., 2024 and predecessors) adds a full reaction term. We now build the latter.
Over-smoothing quantified: the Dirichlet energy metric. To measure how much a GNN over-smooths, we track the Dirichlet energy — a scalar that measures how different neighbouring node features are:
| |
The exponential decay is not a bug — it’s a theorem: for the normalised adjacency $\tilde{A}$ , $E(\tilde{A}^L X_0) \leq \lambda_2^{2L} \cdot E(X_0)$ where $\lambda_2 < 1$ is the second-largest eigenvalue of $\tilde{A}$ . On Cora, $\lambda_2 \approx 0.98$ , so by layer 64, the energy has decayed by a factor of $0.98^{128} \approx 0.08$ — nearly zero.
Comparison: depth-handling techniques.
| Method | Mechanism | Max useful depth | Extra cost | Preserves structure? |
|---|---|---|---|---|
| Vanilla GCN | $\tilde{A}X$ propagation | ~4-6 layers | — | Over-smooths |
| DropEdge | Randomly drop edges | ~16 layers | Negligible | Partially |
| PairNorm | Normalise features | ~16 layers | $O(N \cdot d)$ | No |
| Residual connections | $X + \tilde{A}X$ | ~32 layers | Negligible | Partially |
| GRAND | Neural ODE on graph | ~64 layers | ODE solver | Adaptive |
| RDGNN | Diffusion + reaction | ~64+ layers | Reaction MLP | Yes (Turing) |
RDGNN is the only method with a principled mechanism (Turing instability) that explains why it works at depth. The others are heuristics that slow down the smoothing without preventing it.
RDGNN: Reaction-Diffusion Graph Neural Networks#
The Architecture#
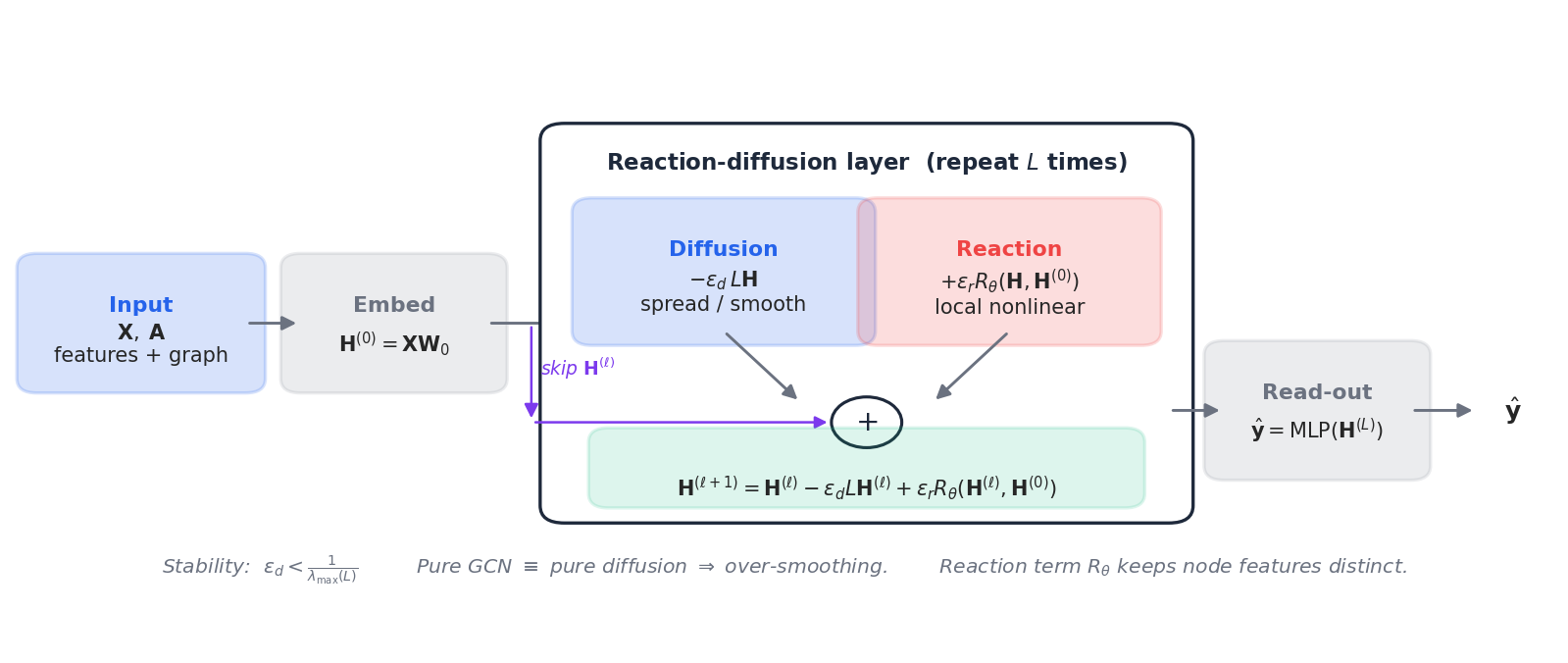
$$ \frac{d\mathbf{H}}{dt} = -\epsilon_d\,\mathbf{L}\,\mathbf{H} \;+\; \epsilon_r\,R_\theta(\mathbf{H}, \mathbf{H}^{(0)}). \tag{6} $$ $$\boxed{\;\mathbf{H}^{(\ell+1)} = \mathbf{H}^{(\ell)} \;-\; \epsilon_d\,\mathbf{L}\,\mathbf{H}^{(\ell)} \;+\; \epsilon_r\,R_\theta\bigl(\mathbf{H}^{(\ell)},\,\mathbf{H}^{(0)}\bigr).\;} \tag{7}$$Three blocks (Fig. 5):
- Diffusion $-\epsilon_d \mathbf{L}\mathbf{H}^{(\ell)}$ : standard graph smoothing. Step size constraint: $\epsilon_d < 1/\lambda_{\max}(\mathbf{L})$ for explicit-Euler stability.
- Reaction $\epsilon_r R_\theta(\mathbf{H}^{(\ell)}, \mathbf{H}^{(0)})$ : a learned, purely local transform — typically a small MLP applied node-wise. Conditioning on $\mathbf{H}^{(0)}$ acts like the input-skip in a ResNet and prevents drift.
- Skip $\mathbf{H}^{(\ell)}$ in the update: keeps the dynamics close to the identity, which is what makes large $L$ stable in practice.
Why It Works#
There are two ways to see why a reaction term cures over-smoothing.
Spectral view. Pure diffusion attenuates every non-constant mode at rate $e^{-\lambda_k t}$ . The reaction term is not a function of $\mathbf{L}$ and therefore can have arbitrary spectral content; in particular it can pump energy back into the high-frequency modes that diffusion is killing. The net effect is a non-trivial steady-state distribution of energy across $k$ .
Turing view. If $R_\theta$ has activator-inhibitor structure (which a sufficiently expressive MLP can learn), and the diffusion strength $\epsilon_d$ is chosen so that the eigenvalues of $\mathbf{J} - \epsilon_d\,\lambda_k$ are unstable for some $k$ , the network exhibits node-level Turing patterns — different nodes converge to different feature values, organised by the graph spectrum. This is the GNN equivalent of stripes on a fish.
PyTorch Implementation#
| |
Notice how lean the architecture is: one shared GCN for diffusion, $L$ small MLPs for reaction, two linear projections at the ends. The accuracy curves in Fig. 6c show that this is enough to maintain Cora performance up to 64 layers — eight times deeper than the regime in which GCN collapses.
Where Reaction-Diffusion Already Wins#
The same equation, three application stories.
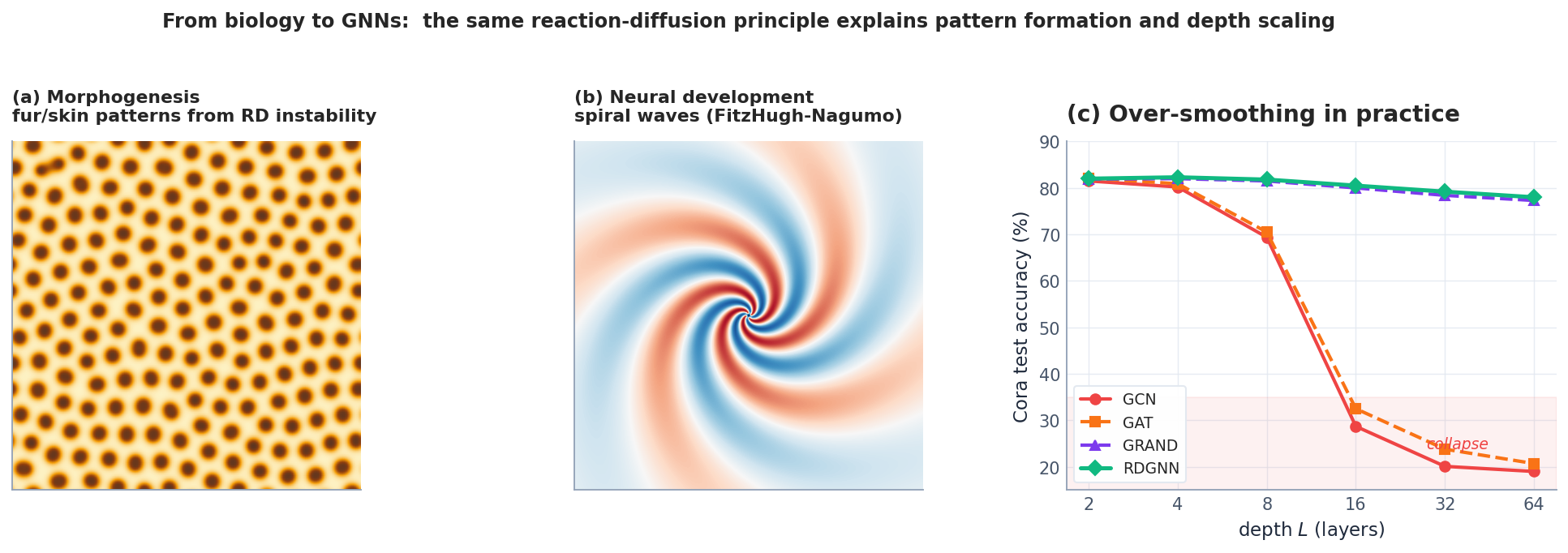
Morphogenesis. Murray’s Mathematical Biology uses Turing’s mechanism to fit the spot-and-stripe transitions on big-cat coats: large cats (jaguar) get spots, intermediate cats (leopard) get rosettes, the tail (where the local geometry constrains $|\mathbf{k}|$ ) gets stripes — all from a single set of reaction parameters and the geometry of the embryo. The same maths predicts vegetation stripes in semi-arid landscapes (water as inhibitor, biomass as activator) and the spiral arms of Belousov-Zhabotinsky chemistry experiments.
Neural development. During retinal mosaic formation, neighbouring photoreceptors inhibit each other from differentiating into the same subtype while activating the same subtype across longer ranges via diffusing morphogens. This is mathematically a Turing system, and the resulting cone arrangements have measurable wavelength $\ell \sim 2\pi/|\mathbf{k}_*|$ . Spiral waves in cortical electrical activity — pathological during epilepsy, important during developmental wiring — are FitzHugh-Nagumo solutions on a 2D excitable medium.
Deep GNNs. On standard benchmarks, the depth-vs-accuracy story is dramatic (Fig. 6c, replicated from Eliasof et al. (2024) on Cora). GCN and GAT collapse beyond 8 layers as the spectral proof predicts; pure-diffusion GRAND merely delays the collapse by being more accurate; only RDGNN — explicit reaction term — maintains accuracy at $L = 64$ . The same effect is even larger on heterophilic graphs (where neighbours tend to have different labels), because pure smoothing is actively harmful there: the reaction term can learn to amplify differences between connected nodes.
Series Finale: Eight Chapters, One Idea#
We are at the end of the PDE + Machine Learning series. Step back.
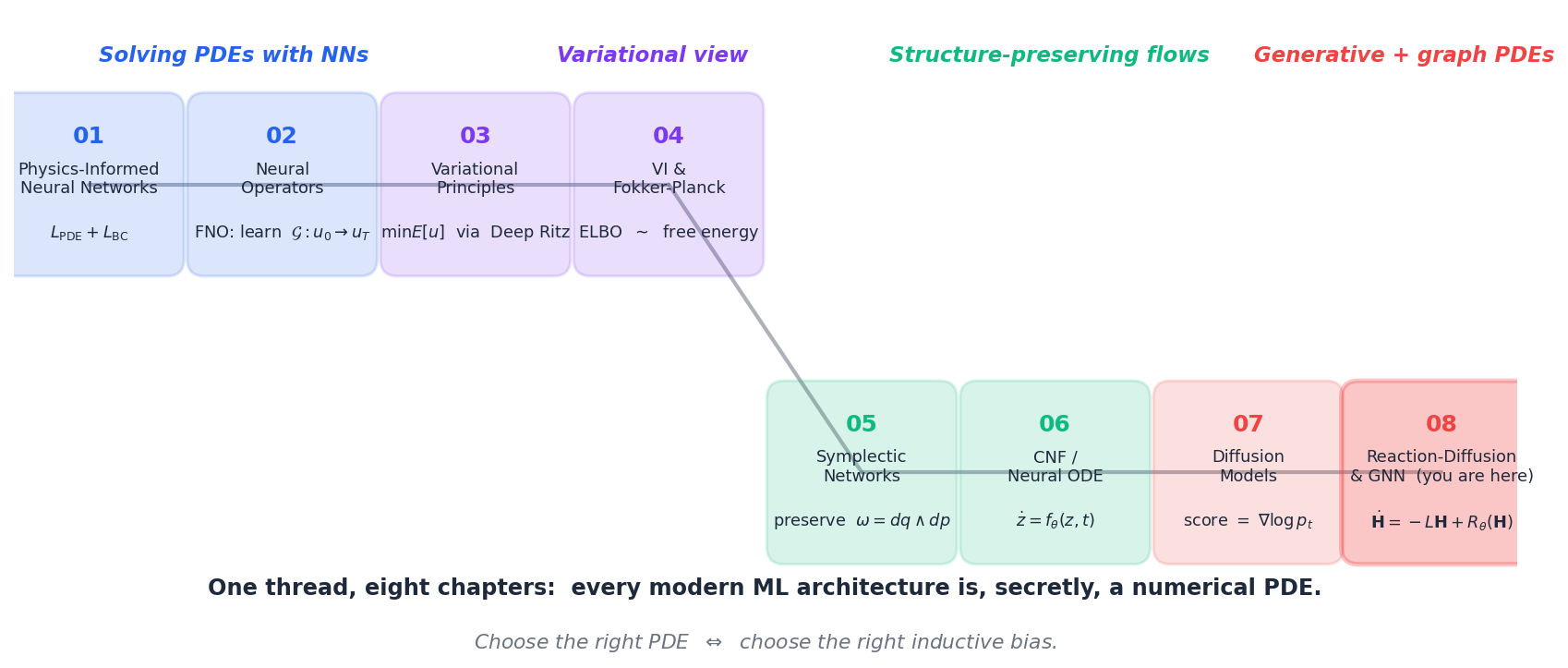
The eight chapters fall into four pairs.
| Pair | Chapters | The PDE in the picture |
|---|---|---|
| Solving PDEs with NNs | 1 PINN, 2 Neural Operators | The target equation itself becomes the loss. |
| Variational view | 3 Deep Ritz, 4 VI / Fokker-Planck | Loss $=$ free energy; gradient flow $=$ continuity equation. |
| Structure-preserving flows | 5 Symplectic nets, 6 Neural ODE / CNF | Network respects the symplectic / volume / divergence structure of the flow. |
| Generative + graph PDEs | 7 Diffusion models, 8 RD + GNN | Forward / reverse Fokker-Planck; reaction-diffusion on graphs. |
Underneath every chapter sits one slogan:
A neural architecture is a discretised PDE. Choose the architecture by choosing the right PDE.
Concretely:
- Want extrapolation off the training grid? Choose an operator-learning PDE (Chapter 2 ).
- Want a network that reproduces conserved quantities? Choose a symplectic integrator (Chapter 5 ).
- Want a tractable likelihood? Choose a continuity equation and learn its drift (Chapter 6 ).
- Want to sample a complicated distribution? Choose a Fokker-Planck and learn its score (Chapter 7 ).
- Want a deep GNN that does not collapse? Choose a reaction-diffusion equation, not just diffusion (this chapter).
The PDE perspective is not the only useful lens on deep learning, but it is uncommonly generative: every time we have asked “what would the corresponding numerical analysis say?” we have learned something concrete — a stability bound, a step-size constraint, a structural fix. This is what physics-style thinking buys you in machine learning, and it is why the conversation between the two fields is far from over.
Exercises#
Exercise 1. Show that for a connected graph the only solution of $\mathbf{L}\mathbf{x} = \mathbf{0}$ is the constant. Conclude that the heat equation on a connected graph drives every initial condition to its mean.
Solution. From (3), $\mathbf{x}^\top\!\mathbf{L}\mathbf{x} = \tfrac{1}{2}\sum w_{ij}(x_i - x_j)^2 = 0$ forces $x_i = x_j$ across every edge; on a connected graph this means $\mathbf{x}$ is constant. The kernel of $\mathbf{L}$ is therefore one-dimensional. Every other eigenvalue is strictly positive, so $e^{-\mathbf{L}t}$ kills all non-constant components and preserves the mean. $\blacksquare$
Exercise 2. Derive the explicit-Euler stability bound $\epsilon_d < 1/\lambda_{\max}(\mathbf{L})$ for the diffusion step.
Solution. In spectral coordinates the Euler update is $\hat{x}_k^{(\ell+1)} = (1 - \epsilon_d \lambda_k)\,\hat{x}_k^{(\ell)}$ . For non-growth we need $|1 - \epsilon_d \lambda_k| \leq 1$ for every $k$ , which gives $0 \leq \epsilon_d \lambda_k \leq 2$ , i.e. $\epsilon_d \leq 2/\lambda_{\max}$ . Asymptotic monotone decay (no oscillation) requires the stricter $\epsilon_d < 1/\lambda_{\max}$ . $\blacksquare$
Exercise 3. Why does RDGNN help especially on heterophilic graphs?
Solution. On heterophilic graphs neighbours tend to carry opposite labels. The pure diffusion step averages neighbouring features and therefore actively destroys the discriminative signal — the more layers, the worse. The reaction term operates node-wise and is conditioned on $\mathbf{H}^{(0)}$ , so it can produce node-specific updates that increase the difference between neighbours, restoring class separation. $\blacksquare$
Exercise 4. Show that the discrete RDGNN update (7) is the first-order operator-splitting (Lie-Trotter) discretisation of the continuous RD-GNN (6).
Solution. Operator splitting writes $\dot{\mathbf{H}} = (\mathcal{D} + \mathcal{R})\,\mathbf{H}$ with $\mathcal{D}\mathbf{H} = -\mathbf{L}\mathbf{H}$ and $\mathcal{R}\mathbf{H} = R_\theta(\mathbf{H})/\epsilon_r$ , then alternates one Euler step of each: $\mathbf{H}^{1/2} = \mathbf{H} + h\mathcal{D}\mathbf{H}$ , $\mathbf{H}^{(\ell+1)} = \mathbf{H}^{1/2} + h\mathcal{R}\mathbf{H}^{1/2}$ . Because both operators are evaluated at the same $\mathbf{H}^{(\ell)}$ in the standard implementation, the result is exactly (7). The local truncation error is $\mathcal{O}(h^2[\mathcal{D}, \mathcal{R}])$ , i.e. first order. $\blacksquare$
Exercise 5. A single Turing instability condition can be checked numerically: pick parameters $(D_u, D_v, F, k)$ for Gray-Scott, linearise around the homogeneous steady state, sweep $|\mathbf{k}|^2$ and look for sign changes of $\det\,\mathbf{A}(|\mathbf{k}|^2)$ . Implement this and reproduce a row of Fig. 1.
Solution sketch. The homogeneous steady state of Gray-Scott solves $u v^2 = F(1 - u)$ and $u v^2 = (F + k) v$ . Compute the $2\times 2$ Jacobian of the kinetics at this state, build $\mathbf{A}(|\mathbf{k}|^2) = \mathbf{J} - |\mathbf{k}|^2 \mathrm{diag}(D_u, D_v)$ , and plot $\det\,\mathbf{A}$ vs $|\mathbf{k}|^2$ . A negative dip indicates an unstable band; the corresponding wavelength $2\pi/|\mathbf{k}_*|$ matches the visual scale of the simulated patterns. $\blacksquare$
Where to go from here#
That closes the eight-chapter PDE-and-ML arc. Looking back, the thread is cleaner than it looked along the way:
- PINNs (Ch. 1) wrote the PDE residual into the loss;
- Neural operators (Ch. 2) upgraded “learn one solution” to “learn the entire solution operator”;
- Variational principles (Ch. 3) revealed the continuous-time geometry behind training;
- Variational inference & Fokker-Planck (Ch. 4) unified optimization and sampling under one PDE;
- Symplectic geometry (Ch. 5) baked conservation laws into network architecture;
- Continuous normalizing flows (Ch. 6) turned generation into an invertible ODE;
- Diffusion models (Ch. 7) translated generation into a reverse heat equation;
- Reaction-diffusion & GNNs (Ch. 8) used Turing’s 1952 theory to repair deep GNNs.
Lined up together, the whole series has been doing exactly one thing: using mathematical structure as the inductive bias of a neural network. Each chapter is the same idea with a different constraint plugged in.
Where to next? Three threads are worth chasing further: (1) operator learning meeting foundation models (PDE foundation models); (2) structure-preserving networks landing in molecular dynamics and climate modeling; (3) score-based methods crossing into reinforcement learning and planning. I hope these eight chapters left you with the key needed to walk down any one of them.
References#
[1] Turing, A. M. (1952). The chemical basis of morphogenesis. Phil. Trans. R. Soc. B, 237(641), 37-72.
[2] Pearson, J. E. (1993). Complex patterns in a simple system. Science, 261(5118), 189-192.
[3] Murray, J. D. (2003). Mathematical biology II: Spatial models and biomedical applications (3rd ed.). Springer.
[4] Kipf, T. N., & Welling, M. (2017). Semi-supervised classification with graph convolutional networks. ICLR. arXiv:1609.02907 .
[5] Oono, K., & Suzuki, T. (2020). Graph neural networks exponentially lose expressive power for node classification. ICLR. arXiv:1905.10947 .
[6] Chamberlain, B., Rowbottom, J., Gorinova, M., Webb, S., Rossi, E., & Bronstein, M. (2021). GRAND: Graph neural diffusion. ICML. arXiv:2106.10934 .
[7] Thorpe, M., Nguyen, T. M., Xia, H., Strohmer, T., Bertozzi, A., Osher, S., & Wang, B. (2022). GRAND++: Graph neural diffusion with a source term. ICLR.
[8] Eliasof, M., Haber, E., & Treister, E. (2021). PDE-GCN: Novel architectures for graph neural networks motivated by partial differential equations. NeurIPS. arXiv:2108.01938 .
[9] Di Giovanni, F., Rowbottom, J., Chamberlain, B., Markovich, T., & Bronstein, M. (2022). Graph neural networks as gradient flows. arXiv:2206.10991 .
[10] Choi, J., Hong, S., Park, N., & Cho, S.-B. (2023). GREAD: Graph neural reaction-diffusion networks. ICML. arXiv:2211.14208 .
[11] Eliasof, M., Haber, E., & Treister, E. (2024). Graph neural reaction-diffusion models. SIAM J. Sci. Comput. arXiv:2406.10871 .
This is Part 8 — the final part — of the PDE and Machine Learning series. Previous: Part 7 — Diffusion Models and Score Matching . Start from the beginning: Part 1 — Physics-Informed Neural Networks . Thanks for reading.
PDE and Machine Learning 8 parts
- 01 PDE and ML (1): Physics-Informed Neural Networks
- 02 PDE and ML (2): Neural Operator Theory
- 03 PDE and ML (3): Variational Principles and Optimization
- 04 PDE and ML (4): Variational Inference and the Fokker-Planck Equation
- 05 PDE and ML (5): Symplectic Geometry and Structure-Preserving Networks
- 06 PDE and ML (6): Continuous Normalizing Flows and Neural ODE
- 07 PDE and ML (7): Diffusion Models and Score Matching
- 08 PDE and ML (8): Reaction-Diffusion Systems and Graph Neural Networks you are here